Abstract
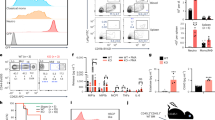
The deaminase ADAR1 edits adenosines in nuclear transcripts of nervous tissue and is required in the fetal liver of the developing mouse embryo. Here we show by inducible gene disruption in mice that ADAR1 is essential for maintenance of both fetal and adult hematopoietic stem cells. Loss of ADAR1 in hematopoietic stem cells led to global upregulation of type I and II interferon–inducible transcripts and rapid apoptosis. Our findings identify ADAR1 as an essential regulator of hematopoietic stem cell maintenance and suppressor of interferon signaling that may protect organisms from the deleterious effects of interferon activation associated with many pathological processes, including chronic inflammation, autoimmune disorders and cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
09 April 2009
In the version of this article initially published, the Cre-transgenic mouse is identified incorrectly as Tg(SV40-cre)1Jrg. The correct mouse strain should be Tg(SCL6E5-Cre)1Jrg, and the citation describing this mouse (ref. 29) should be as follows: Gothert, J.R. et al. In vivo fate-tracing studies using the Scl stem cell enhancer: embryonic hematopoietic stem cells significantly contribute to adult hematopoiesis. Blood 105, 2724–2732 (2005). The error has been corrected in the HTML and PDF versions of the article.
References
Bass, B.L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71, 817–846 (2002).
Athanasiadis, A., Rich, A. & Maas, S. Widespread A-to-I RNA editing of Alu-containing mRNAs in the human transcriptome. PLoS Biol. 2, e391 (2004).
Kim, D.D. et al. Widespread RNA editing of embedded alu elements in the human transcriptome. Genome Res. 14, 1719–1725 (2004).
Levanon, E.Y. et al. Systematic identification of abundant A-to-I editing sites in the human transcriptome. Nat. Biotechnol. 22, 1001–1005 (2004).
Kawahara, Y. et al. Redirection of silencing targets by adenosine-to-inosine editing of miRNAs. Science 315, 1137–1140 (2007).
Luciano, D.J., Mirsky, H., Vendetti, N.J. & Maas, S. RNA editing of a miRNA precursor. RNA 10, 1174–1177 (2004).
Yang, W. et al. Modulation of microRNA processing and expression through RNA editing by ADAR deaminases. Nat. Struct. Mol. Biol. 13, 13–21 (2006).
Hartner, J.C. et al. Liver disintegration in the mouse embryo caused by deficiency in the RNA-editing enzyme ADAR1. J. Biol. Chem. 279, 4894–4902 (2004).
Wang, Q. et al. Stress-induced apoptosis associated with null mutation of ADAR1 RNA editing deaminase gene. J. Biol. Chem. 279, 4952–4961 (2004).
Higuchi, M. et al. Point mutation in an AMPA receptor gene rescues lethality in mice deficient in the RNA-editing enzyme ADAR2. Nature 406, 78–81 (2000).
Melcher, T. et al. RED2, a brain-specific member of the RNA-specific adenosine deaminase family. J. Biol. Chem. 271, 31795–31798 (1996).
Herbert, A. et al. A Z-DNA binding domain present in the human editing enzyme, double-stranded RNA adenosine deaminase. Proc. Natl. Acad. Sci. USA 94, 8421–8426 (1997).
Kim, U., Wang, Y., Sanford, T., Zeng, Y. & Nishikura, K. Molecular cloning of cDNA for double-stranded RNA adenosine deaminase, a candidate enzyme for nuclear RNA editing. Proc. Natl. Acad. Sci. USA 91, 11457–11461 (1994).
O'Connell, M.A. et al. Cloning of cDNAs encoding mammalian double-stranded RNA-specific adenosine deaminase. Mol. Cell. Biol. 15, 1389–1397 (1995).
George, C.X., Das, S. & Samuel, C.E. Organization of the mouse RNA-specific adenosine deaminase Adar1 gene 5′-region and demonstration of STAT1-independent, STAT2-dependent transcriptional activation by interferon. Virology 380, 338–343 (2008).
George, C.X. & Samuel, C.E. Human RNA-specific adenosine deaminase ADAR1 transcripts possess alternative exon 1 structures that initiate from different promoters, one constitutively active and the other interferon inducible. Proc. Natl. Acad. Sci. USA 96, 4621–4626 (1999).
George, C.X., Wagner, M.V. & Samuel, C.E. Expression of interferon-inducible RNA adenosine deaminase ADAR1 during pathogen infection and mouse embryo development involves tissue-selective promoter utilization and alternative splicing. J. Biol. Chem. 280, 15020–15028 (2005).
Eckmann, C.R., Neunteufl, A., Pfaffstetter, L. & Jantsch, M.F. The human but not the Xenopus RNA-editing enzyme ADAR1 has an atypical nuclear localization signal and displays the characteristics of a shuttling protein. Mol. Biol. Cell 12, 1911–1924 (2001).
Samuel, C.E. Antiviral actions of interferons. Clin. Microbiol. Rev. 14, 778–809 (2001).
Wang, Q., Khillan, J., Gadue, P. & Nishikura, K. Requirement of the RNA editing deaminase ADAR1 gene for embryonic erythropoiesis. Science 290, 1765–1768 (2000).
Orkin, S.H. & Zon, L.I. Hematopoiesis: an evolving paradigm for stem cell biology. Cell 132, 631–644 (2008).
Jordan, C.T., McKearn, J.P. & Lemischka, I.R. Cellular and developmental properties of fetal hematopoietic stem cells. Cell 61, 953–963 (1990).
Okada, S. et al. In vivo and in vitro stem cell function of c-kit- and Sca-1-positive murine hematopoietic cells. Blood 80, 3044–3050 (1992).
Kiel, M.J. et al. SLAM family receptors distinguish hematopoietic stem and progenitor cells and reveal endothelial niches for stem cells. Cell 121, 1109–1121 (2005).
Kim, I., He, S., Yilmaz, O.H., Kiel, M.J. & Morrison, S.J. Enhanced purification of fetal liver hematopoietic stem cells using SLAM family receptors. Blood 108, 737–744 (2006).
Kuhn, R., Schwenk, F., Aguet, M. & Rajewsky, K. Inducible gene targeting in mice. Science 269, 1427–1429 (1995).
Hock, H. et al. Gfi-1 restricts proliferation and preserves functional integrity of haematopoietic stem cells. Nature 431, 1002–1007 (2004).
Mikkola, H.K. et al. Haematopoietic stem cells retain long-term repopulating activity and multipotency in the absence of stem-cell leukaemia SCL/tal-1 gene. Nature 421, 547–551 (2003).
Gothert, J.R. et al. In vivo fate-tracing studies using the Scl stem cell enhancer: embryonic hematopoietic stem cells significantly contribute to adult hematopoiesis. Blood 105, 2724–2732 (2005).
Lessard, J., Faubert, A. & Sauvageau, G. Genetic programs regulating HSC specification, maintenance and expansion. Oncogene 23, 7199–7209 (2004).
Randall, T.D. & Weissman, I.L. Phenotypic and functional changes induced at the clonal level in hematopoietic stem cells after 5-fluorouracil treatment. Blood 89, 3596–3606 (1997).
Morrison, S.J. & Weissman, I.L. The long-term repopulating subset of hematopoietic stem cells is deterministic and isolatable by phenotype. Immunity 1, 661–673 (1994).
Sinclair, A., Daly, B. & Dzierzak, E. The Ly-6E.1 (Sca-1) gene requires a 3′ chromatin-dependent region for high-level gamma-interferon-induced hematopoietic cell expression. Blood 87, 2750–2761 (1996).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Geiss, G. et al. A comprehensive view of regulation of gene expression by double-stranded RNA-mediated cell signaling. J. Biol. Chem. 276, 30178–30182 (2001).
Nie, Y., Ding, L., Kao, P.N., Braun, R. & Yang, J.H. ADAR1 interacts with NF90 through double-stranded RNA and regulates NF90-mediated gene expression independently of RNA editing. Mol. Cell. Biol. 25, 6956–6963 (2005).
Krasnoselskaya-Riz, I. et al. Nuclear factor 90 mediates activation of the cellular antiviral expression cascade. AIDS Res. Hum. Retroviruses 18, 591–604 (2002).
Tsai, F.Y. et al. An early haematopoietic defect in mice lacking the transcription factor GATA-2. Nature 371, 221–226 (1994).
Wang, Z. et al. Regulation of innate immune responses by DAI (DLM-1/ZBP1) and other DNA-sensing molecules. Proc. Natl. Acad. Sci. USA 105, 5477–5482 (2008).
Apostolou, E. & Thanos, D. Virus infection induces NF-κB-dependent interchromosomal associations mediating monoallelic IFN-β gene expression. Cell 134, 85–96 (2008).
Walkley, C.R. & Orkin, S.H. Rb is dispensable for self-renewal and multilineage differentiation of adult hematopoietic stem cells. Proc. Natl. Acad. Sci. USA 103, 9057–9062 (2006).
Walkley, C.R., Shea, J.M., Sims, N.A., Purton, L.E. & Orkin, S.H. Rb regulates interactions between hematopoietic stem cells and their bone marrow microenvironment. Cell 129, 1081–1095 (2007).
Gordon, K.M., Duckett, L., Daul, B. & Petrie, H.T. A simple method for detecting up to five immunofluorescent parameters together with DNA staining for cell cycle or viability on a benchtop flow cytometer. J. Immunol. Methods 275, 113–121 (2003).
Acknowledgements
We thank M. Higuchi and P.H. Seeburg (Max-Planck Institute for Medical Research) for Adar mutant mice; K. Rajewsky (Immune Disease Institute, Boston) for Mx1-Cre mice; J. Goethert (University Hospital Essen) for SCL-Cre-ERT mice; the Dana-Farber Cancer Institute and Children's Hospital animal facility staff for care of experimental mice; J. Daley and S. Lazo-Kallanian of the Dana-Farber Cancer Institute flow cytometry facility for assistance with cell sorting; J. Shea for technical assistance; T.R. Golub for support in gene expression analysis; and M. Higuchi, A. Athanasiadis, S. Maas and L. Purton for critical comments on the manuscript. Supported by the Leukemia and Lymphoma Society (C.R.W.) and the Howard Hughes Medical Institute (S.H.O.).
Author information
Authors and Affiliations
Contributions
J.C.H. conceived the study, designed and did experiments, analyzed and interpreted data and wrote the paper; C.R.W. did experiments and analyzed and interpreted data; J.L. analyzed and interpreted data; and S.H.O. analyzed and interpreted data and wrote the paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Tables 1–4 (PDF 1257 kb)
Rights and permissions
About this article
Cite this article
Hartner, J., Walkley, C., Lu, J. et al. ADAR1 is essential for the maintenance of hematopoiesis and suppression of interferon signaling. Nat Immunol 10, 109–115 (2009). https://doi.org/10.1038/ni.1680
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.1680
This article is cited by
-
Biological roles of A-to-I editing: implications in innate immunity, cell death, and cancer immunotherapy
Journal of Experimental & Clinical Cancer Research (2023)
-
Emerging role of the RNA-editing enzyme ADAR1 in stem cell fate and function
Biomarker Research (2023)
-
Novel insights into double-stranded RNA-mediated immunopathology
Nature Reviews Immunology (2023)
-
SARS-CoV-2 and innate immunity: the good, the bad, and the “goldilocks”
Cellular & Molecular Immunology (2023)
-
A novel computational method enables RNA editome profiling during human hematopoiesis from scRNA-seq data
Scientific Reports (2023)