Abstract
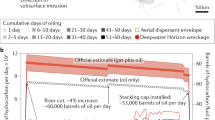
The blowout of the Macondo oil well in the Gulf of Mexico in April 2010 injected up to 500,000 tonnes of natural gas, mainly methane, into the deep sea1. Most of the methane released was thought to have been consumed by marine microbes between July and August 20102,3. Here, we report spatially extensive measurements of methane concentrations and oxidation rates in the nine months following the spill. We show that although gas-rich deepwater plumes were a short-lived feature, water column concentrations of methane remained above background levels throughout the rest of the year. Rates of microbial methane oxidation peaked in the deepwater plumes in May and early June, coincident with a rapid rise in the abundance of known and new methane-oxidizing microbes. At this time, rates of methane oxidation reached up to 5,900 nmol l−1 d−1—the highest rates documented in the global pelagic ocean before the blowout4. Rates of methane oxidation fell to less than 50 nmol l−1 d−1 in late June, and continued to decline throughout the remainder of the year. We suggest the precipitous drop in methane consumption in late June, despite the persistence of methane in the water column, underscores the important role that physiological and environmental factors play in constraining the activity of methane-oxidizing bacteria in the Gulf of Mexico.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
28 April 2015
We would like to provide a more comprehensive description of sample handling and the timing of methane oxidation rate assays. The special circumstances of the Deepwater Horizon oil-well blowout required adaptation of standard operating procedures. As a result, methane oxidation rate assays on 248 of the 1,128 samples we took in our study were not initiated within 12–24 hours of sample collection (Supplementary Table 1), because some cruises lacked a radioisotope isolation van and samples had to be shipped to the University of Georgia for processing after the cruise ended. Specifically, on the RV Pelican, RV Walton Smith, RV Arctic Sunrise, and the RV Nancy Foster cruises, samples were stored for up to 29 days before initiation of rate assays. Sample storage times for these cruises are given in Supplementary Table 1. To evaluate potential issues arising from storage and shipping, we measured the concentration of dissolved oxygen and nutrients on the ship and after storage, and bottles were visually inspected for gas bubbles, and visible changes (for example, turbidity, aggregations) in microbial biomass. On board the ship, samples were stored without a headspace in a gas-tight bottle in a refrigerator set at 4 °C. The temperature of the refrigerator was checked at least every four hours to assure stability. We prepared samples for shipment to Athens (Georgia, USA) by placing them on a layer of Styrofoam peanuts (∼2.5 cm thick), which overlaid a layer of blue ice packs. Similarly, blue ice packs lined the sides of each cooler and were separated from the sample bottles by cardboard. A piece of cardboard was placed on top of the samples, and an ice pack was then placed on top of the cardboard. No warming occurred during shipment and samples remained at deep-sea temperatures (3–4 °C) until arrival in Athens. Several lines of evidence show that storage did not affect our results. First, visual assessment gave no indication of changes in microbial biomass (for example, blooms) during storage: there were no cell clumps, no visible turbidity, no microbial slime, and no growth on container walls, any of which could signal cell growth or death during storage. Second, no bottles went anoxic or exhibited the levels of nutrient or oxygen drawdown that would point towards a microbial population explosion during storage and/or shipment. Third, an increase of methanotroph biomass during storage would have increased the volumetric rates of methane oxidation and decreased the substrate concentration, both of which would increase the value of k, the specific methane oxidation rate. Our data show no such relationship between storage time and k; this is particularly noteworthy for the high-activity period in late May and early June. These observations are consistent with the current understanding of methanotrophs, which are fastidious and difficult to enrich and isolate, and tend to be inactive under stable (static) storage conditions1,2,3. Gulf waters, incubated in stirred bottles and monitored over time to track changes in methane concentration, have been used to estimate methane oxidation rates from the linear decrease in methane concentration over time in bottles during laboratory incubation1,2,3. Over periods up to 30 days, methane concentrations decreased linearly (with r2 varying from 0.995 to 0.973). These observations have been replicated multiple times2, and clearly show that methanotrophy depends upon turbulence: bottles incubated under static/stable conditions showed no significant methane consumption2. This reflects the common observation that headspace-free static storage or static incubation generally reflects lower microbial growth and activity4,5,6. Furthermore, methane oxidation rates determined from stored samples exhibited a similar pattern — as did the distribution of a key functional gene involved in aerobic methane oxidation, pmoA, measured in samples filtered and fixed on board the ship. Both rates (Fig. 1) and pmoA gene abundance (Fig. 2) increased by orders of magnitude in late May and early June and then relaxed back towards much lower values, although both remained above background values. As the molecular samples were filtered and fixed on board the ship, the pmoA gene abundance data cannot reflect a storage effect. We conclude that the likelihood of a methanotroph bloom during storage of these samples is exceedingly small. References 1. Bugna, G. C., Chanton, J. P., Young, J. E., Burnett, W. C. & Cable, P. H. The importance of groundwater discharge to the methane budget of nearshore and continental shelf waters of the NE Gulf of Mexico. Geochim. Cosmochim. Acta 60, 4735–4746 (1996). 2. Bugna, G. C. Methane as a Tracer of Submarine Groundwater Discharge into the NE Gulf of Mexico Masters thesis, Florida State Univ. (1995). 3. Kelley, C. A. Methane oxidation potential in the water column of two diverse coastal marine sites. Biogeochemistry 65, 105–120 (2003). 4. Lidstrom, M. E. Isolation and characterization of marine methanotrophs. A. Van Leeuw. J. Microb. 54, 189–199 (1988). 5. Wartiainen, I., Hestnes, A. G., McDonald, I. R. & Svenning, M. M. Methylobacter tundripaludum sp. nov., a methane-oxidizing bacterium from Arctic wetland soil on the Svalbard islands, Norway (78° N). Int. J. Sys. Evol. Microbiol. 56, 109–113 (2006). 6. Juergensmeyer, M. A., Nelson, E. S. & Juergensmeyer, E. A. Shaking alone, without concurrent aeration, affects the growth characteristics of Eschericia coli. Lett. Appl. Microbiol. 45, 179–183 (2007). Additional information Supplementary information is available in the online version of the Addendum.
References
Joye, S. B., MacDonald, I. R., Leifer, I. & Asper, V. Magnitude and oxidation potential of hydrocarbon gases released from the BP oil well blowout. Nature Geosci. 4, 160–164 (2011).
Kessler, J. D. et al. A persistent oxygen anomaly reveals the fate of spilled methane in the deep Gulf of Mexico. Science 331, 312–315 (2011).
Valentine, D. L. et al. Dynamic autoinoculation and the microbial ecology of a deep water hydrocarbon irruption. Proc. Natl Acad. Sci. USA 109, 20286–20291 (2012).
De Angelis, M. A., Lilley, M. d., Olson, E. J. & Baross, J. A. Methane oxidation in deep-sea hydrothermal plumes of the endeavour segment of the Juan de Fuca Ridge. Deep-Sea Res. I 40, 1169–1186 (1993).
Valentine, D. L. et al. Propane respiration jump-starts microbial response to a deep oil spill. Science 330, 208–211 (2010).
Camilli, R. et al. Tracking hydrocarbon plume transport and biodegradation at Deepwater Horizon. Science 330, 201–204 (2010).
Bowles, M. W., Samarkin, V. A., Bowles, K. M. & Joye, S. B. Weak coupling between sulfate reduction and the anaerobic oxidation of methane in methane-rich seafloor sediments during ex situ incubation. Geochim. Cosmochim. Acta 75, 500–519 (2011).
Kunze, E. Near-inertial wave propagation in geostrophic shear. J. Phys. Oceanogr. 15, 544–565 (1985).
Polzin, K. L., Toole, J. M., Ledwell, J. R. & Schmitt, R. W. Spatial variability of turbulent mixing in the Abyssal Ocean. Science 276, 93–96 (1997).
Sassen, R. et al. Thermogenic gas hydrates and hydrocarbon gases in complex chemosynthetic communities, Gulf of Mexico continental slope. Org. Geochem. 30, 485–497 (1999).
Wankel, S. D. et al. New constraints on methane fluxes and rates of anaerobic methane oxidation in a Gulf of Mexico brine pool via in situ mass spectrometry. Deep-Sea Res. II 57, 2022–2029 (2010).
Reeburgh, W. S. Oceanic methane biogeochemistry. Chem. Rev. 107, 486–513 (2007).
Segers, R. Methane production and methane consumption: A review of processes underlying wetland methane fluxes. Biogeochemistry 41, 21–51 (1998).
Knief, C., Kolb, S., Bodelier, P. L., Lipski, A. & Dunfield, P. F. The active methanotrophic community in hydromorphic soils changes in response to changing methane concentration. Environ. Microbiol. 8, 321–333 (2006).
Tavormina, P. L., Ussler, W. III, Joye, S. B., Harrison, B. K. & Orphan, V. J. Distributions of putative aerobic methanotrophs in diverse pelagic marine environments. ISME J. 4, 1–11 (2010).
Carini, S., Bano, N., LeCleir, G. & Joye, S. B. Aerobic methane oxidation and methanotroph community composition during seasonal stratification in Mono Lake, California (USA). Environ. Microbiol. 7, 1127–1138 (2005).
Rivers, A. R. et al. Transcriptional response of bathypelagic marine bacterioplankton to the Deepwater Horizon oil spill. ISME J. 7, 2315–2329 (2013).
Tavormina, P. L., Ussler, W. III & Orphan, V. J. Planktonic and sediment-associated aerobic methanotrophs in two seep systems along the North American margin. Appl. Environ. Microbiol. 74, 3985–3995 (2008).
Murase, J. & Frenzel, P. Selective grazing of methanotrophs by protozoa in a rice field soil. Fems Microbiol. Ecol. 65, 408–414 (2008).
Bastviken, D., Ejlertsson, J., Sundh, I. & Tranvik, L. Methane as a source of carbon and energy for lake pelagic food webs. Ecology 84, 969–981 (2003).
Bouvier, T. & del Giorgio, P. A. Key role of selective viral-induced mortality in determining marine bacterial community composition. Environ. Microbiol. 9, 287–297 (2007).
Balasubramanian, R. et al. Oxidation of methane by a biological dicopper centre. Nature 456, 115–120 (2010).
Murrell, J. C., Gilbert, B. & McDonald, I. R. Molecular biology and regulation of methane monooxygenase. Arch. Microbiol. 173, 325–332 (2000).
Takeguchi, M., Ohashi, M. & Okura, I. Role of iron in particulate methane monooxygenase from Methylosinus trichosporium OB3b. Biometals 12, 123–129 (1999).
Bergquist, B. A. & Boyle, E. A. Dissolved iron in the tropical and subtropical Atlantic Ocean. Glob. Biogeochem. Cycles 20, GB1015 (2006).
Berson, O. & Lidstrom, M. E. Study of copper accumulation by the type I methanotroph Methylomicrobium albus BG8. Env. Sci. Tech. 30, 802–809 (1996).
Park, S., Shah, N. N., Taylor, R. T. & Droege, M. W. Batch cultivation of Methylosinus trichosporium OB3b I: Production of soluble methane monooxygenase. Biotechnol. Bioeng. 38, 423–433 (1991).
Joye, S. B., Bowles, M. W., Samarkin, V. A., Hunter, K. S. & Niemann, H. Biogeochemical signatures and microbial activity of different cold-seep habitats along the Gulf of Mexico deep slope. Deep-Sea Res. II 57, 1990–2001 (2010).
Mackelprang, R. et al. Metagenomic analysis of a permafrost microbial community reveals a rapid response to thaw. Nature 480, 368–371 (2011).
Sjostedt, J. et al. Recruitment of members from the rare biosphere of marine bacterioplankton communities after an environmental disturbance. Appl. Environ. Microbiol. 78, 1361–1369 (2012).
Acknowledgements
We thank C. Mann, A. Vossmeyer, J. Slaughter, C. Comerford, L. Potter, V. Samarkin and S. Cummings for assistance at sea and/or in the laboratory; M. Chistoserdova for providing advice on constructing qPCR primers and for providing pure cultures of methanotrophs; I. MacDonald, T. Treude and M. Chistoserdova provided constructive feedback on a previous version of this manuscript. Finally we thank the science parties and ship’s crews of RV Pelican, RV Nancy Foster, RV Walton Smith, RV Oceanus, RV Cape Hatteras, MY Arctic Sunrise and RV Atlantis. This work was supported by the NOAA Award NA07AR4300464 to the National Institute for Undersea Science and Technology (V.L.A., A.R.D. and S.B.J.), the Department of Energy (Gulf of Mexico Gas Hydrate Research Consortium, 07-11-036 to J.P.C.), the National Science Foundation (OCE-1043225 to S.B.J., OCE-0926699 to T.A.V. and OCE-1042934 to A.M.S.), the Gulf of Mexico Research Initiative (ECOGIG (S.B.J., V.L.A., A.B., J.P.C., A.R.D, J.P.M., C.D.M. and T.A.V.) and DEEP-C (J.P.C.)) and the Northern Gulf Institute (A.M.S.). Greenpeace and Texas A&M at Galveston facilitated the Arctic Sunrise expedition. This is ECOGIG contribution #192 and the data fall under GRIIDC accession number (R1.x132.134:0057).
Author information
Authors and Affiliations
Contributions
S.B.J., M.C-M., K.S.H., A-R.D., V.L.A., J.P.C., J.P.M., A.M.W., R.M.W.A., A.M.S., D.J.J., A.V. and T.A.V. collected the samples and/or carried out geochemical and microbial activity rate assays; M.C-M., L.M.N, P.T., J.J.B. and V.J.O. conducted the methanotroph gene analyses; S.B.J., C.D.M., M.C-M., P.T. and A.B. contributed to data reduction and analyses; S.B.J. led development of the manuscript; C.D.M., M.C-M., P.L.T. and V.J.O. made significant contributions and critical feedback was provided by the other co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2491 kb)
Supplementary Information
Supplementary Information (XLSX 57 kb)
Rights and permissions
About this article
Cite this article
Crespo-Medina, M., Meile, C., Hunter, K. et al. The rise and fall of methanotrophy following a deepwater oil-well blowout. Nature Geosci 7, 423–427 (2014). https://doi.org/10.1038/ngeo2156
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ngeo2156
This article is cited by
-
Financial liabilities and environmental implications of unplugged wells for the Gulf of Mexico and coastal waters
Nature Energy (2023)
-
Aerobic oxidation of methane significantly reduces global diffusive methane emissions from shallow marine waters
Nature Communications (2022)
-
Pelagic denitrification and methane oxidation in oxygen-depleted waters of the Louisiana shelf
Biogeochemistry (2021)
-
Reduced methane seepage from Arctic sediments during cold bottom-water conditions
Nature Geoscience (2020)
-
Influence of oil, dispersant, and pressure on microbial communities from the Gulf of Mexico
Scientific Reports (2020)