Abstract
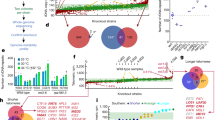
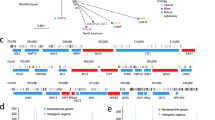
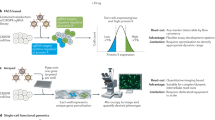
High similarity between yeast and human mitochondria allows functional genomic study of Saccharomyces cerevisiae to be used to identify human genes involved in disease1. So far, 102 heritable disorders have been attributed to defects in a quarter of the known nuclear-encoded mitochondrial proteins in humans2. Many mitochondrial diseases remain unexplained, however, in part because only 40–60% of the presumed 700–1,000 proteins involved in mitochondrial function and biogenesis have been identified3. Here we apply a systematic functional screen using the pre-existing whole-genome pool of yeast deletion mutants4,5,6 to identify mitochondrial proteins. Three million measurements of strain fitness identified 466 genes whose deletions impaired mitochondrial respiration, of which 265 were new. Our approach gave higher selection than other systematic approaches, including fivefold greater selection than gene expression analysis. To apply these advantages to human disorders involving mitochondria, human orthologs were identified and linked to heritable diseases using genomic map positions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Foury, F. Human genetic diseases: a cross-talk between man and yeast. Gene 195, 1–10 (1997).
DiMauro, S. & Schon, E.A. Nuclear power and mitochondrial disease. Nature Genet. 19, 214–215 (1998).
Wallace, D.C. Mitochondrial diseases in man and mouse. Science 283, 1482–1488 (1999).
Winzeler, E.A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Birrell, G.W., Giaever, G., Chu, A.M., Davis, R.W. & Brown, J.M. A genome-wide screen in Saccharomyces cerevisiae for genes affecting UV radiation sensitivity. Proc. Natl Acad. Sci. USA 98, 12608–12613 (2001).
Ooi, S.L., Shoemaker, D.D. & Boeke, J.D. A DNA microarray-based genetic screen for nonhomologous end-joining mutants in Saccharomyces cerevisiae. Science 294, 2552–2556 (2001).
Shoemaker, D.D., Lashkari, D.A., Morris, D., Mittmann, M. & Davis, R.W. Quantitative phenotypic analysis of yeast deletion mutants using a highly parallel molecular bar-coding strategy. Nature Genet. 14, 450–456 (1996).
Tzagoloff, A. & Myers, A.M. Genetics of mitochondrial biogenesis. Annu. Rev. Biochem. 55, 249–285 (1986).
Yaffe, M.P. in Methods in Enzymology Vol. 194 (eds Guthrie, C. & Fink, G.R.) 627–643 (Academic Press, San Diego, California, 1991).
Scharfe, C. et al. MITOP, the mitochondrial proteome database: 2000 update. Nucleic Acids Res. 28, 155–158 (2000).
Grivell, L.A. et al. Mitochondrial assembly in yeast. FEBS Lett. 452, 57–60 (1999).
Wood, V., Rutherford, K.M., Ivens, A., Rajandream, M.A. & Barrell, B. A re-annotation of the Saccharomyces cerevisiae genome. Comp. Funct. Genom. 2, 143–154 (2001).
Claros, M.G. & Vincens, P. Computational method to predict mitochondrially imported proteins and their targeting sequences. Eur. J. Biochem. 241, 779–786 (1996).
Andersson, S.G. et al. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396, 133–140 (1998).
Schwikowski, B., Uetz, P. & Fields, S. A network of protein–protein interactions in yeast. Nature Biotechnol. 18, 1257–1261 (2000).
Kumar, A. et al. Subcellular localization of the yeast proteome. Genes Dev. 16, 707–719 (2002).
Nishino, I., Spinazzola, A. & Hirano, M. Thymidine phosphorylase gene mutations in MNGIE, a human mitochondrial disorder. Science 283, 689–692 (1999).
Ashe, M.P., De Long, S.K. & Sachs, A.B. Glucose depletion rapidly inhibits translation initiation in yeast. Mol. Biol. Cell 11, 833–848 (2000).
DeRisi, J.L., Iyer, V.R. & Brown, P.O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686 (1997).
Hughes, T.R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000).
Boguski, M.S. & Schuler, G.D. ESTablishing a human transcript map. Nature Genet. 10, 369–371 (1995).
Mewes, H.W. et al. MIPS: a database for genomes and protein sequences. Nucleic Acids Res. 28, 37–40 (2000).
Hentati, A. et al. Linkage of 'pure' autosomal recessive familial spastic paraplegia to chromosome 8 markers and evidence of genetic locus heterogeneity. Hum. Mol. Genet. 3, 1263–1267 (1994).
Christodoulou, K. et al. Mapping of the second Friedreich's ataxia (FRDA2) locus to chromosome 9p23–p11: evidence for further locus heterogeneity. Neurogenetics 3, 127–132 (2001).
Kerrison, J.B. et al. Genetic heterogeneity of dominant optic atrophy, Kjer type: identification of a second locus on chromosome 18q12.2–12.3. Arch. Ophthalmol. 117, 805–810 (1999).
Assink, J.J. et al. A gene for X-linked optic atrophy is closely linked to the Xp11.4–Xp11.2 region of the X chromosome. Am. J. Hum. Genet. 61, 934–939 (1997).
Priest, J.M., Fischbeck, K.H., Nouri, N. & Keats, B.J. A locus for axonal motor-sensory neuropathy with deafness and mental retardation maps to Xq24–q26. Genomics 29, 409–412 (1995).
McMullan, T.F., Collins, A.R., Tyers, A.G. & Robinson, D.O. A novel X-linked dominant condition: X-linked congenital isolated ptosis. Am. J. Hum. Genet. 66, 1455–1460 (2000).
Malmgren, H. et al. Linkage mapping of a severe X-linked mental retardation syndrome. Am. J. Hum. Genet. 52, 1046–1052 (1993).
Acknowledgements
We thank M. Mindrinos, E. Allen, T. Neklesa, Q. Wang, W. Neupert and T. Meitinger for helpful advice and M. Trebo for help with preparing the supplementary website. This work was supported by the US National Institutes of Health (P.J.O. and R.W.D.) and the Bundesministerium für Bildung und Forschung (H.P.). L.M.S. was supported as a Howard Hughes Medical Institute predoctoral fellow and C.S. as a Deutsche Forschungsgemeinschaft postdoctoral fellow.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Steinmetz, L., Scharfe, C., Deutschbauer, A. et al. Systematic screen for human disease genes in yeast. Nat Genet 31, 400–404 (2002). https://doi.org/10.1038/ng929
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng929
This article is cited by
-
ECDEP: identifying essential proteins based on evolutionary community discovery and subcellular localization
BMC Genomics (2024)
-
From beer to breadboards: yeast as a force for biological innovation
Genome Biology (2024)
-
Yeast UPS1 deficiency leads to UVC radiation sensitivity and shortened lifespan
Antonie van Leeuwenhoek (2023)
-
Fit-Seq2.0: An Improved Software for High-Throughput Fitness Measurements Using Pooled Competition Assays
Journal of Molecular Evolution (2023)
-
Large-scale genomic and transcriptomic profiles of rice hybrids reveal a core mechanism underlying heterosis
Genome Biology (2022)