Abstract
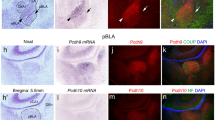
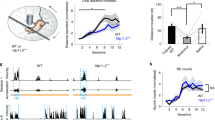
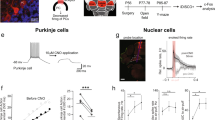
Mice homozygous for the cerebellar deficient folia (cdf) mutation are ataxic and have cerebellar hypoplasia and abnormal lobulation of the cerebellum. In the cerebella of cdf/cdf homozygous mice, approximately 40% of Purkinje cells are located ectopically in the white matter and inner granule-cell layer. Many hippocampal pyramidal cells are scattered in the plexiform layers, and those that are correctly positioned are less densely packed than are cells in wild-type mice. We show that fear conditioning and prepulse inhibition of the startle response are also disrupted in cdf/cdf mice. We identify a deletion on chromosome 6 that removes approximately 150 kb in the cdf critical region. The deletion includes part of Catna2, encoding αN-catenin, a protein that links the classical cadherins to the neuronal cytoskeleton. Expression of a Catna2 transgene in cdf/cdf mice restored normal cerebellar and hippocampal morphology, prepulse inhibition and fear conditioning. The findings suggest that catenin–cadherin cell-adhesion complexes are important in cerebellar and hippocampal lamination and in the control of startle modulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Cook, S.A., Bronson, R.T., Donahue, L.R., Ben-Arie, N. & Davisson, M.T. Cerebellar deficient folia (cdf): a new mutation on mouse Chromosome 6. Mamm. Genome 8, 108–112 (1997).
Swerdlow, N.R. & Geyer, M.A. Prepulse inhibition of acoustic startle in rats after lesions of the pedunculopontine tegmental nucleus. Behav. Neurosci. 107, 104–117 (1993).
Davis, M. & Astrachan, D.I. Conditioned fear and startle magnitude: effects of different footshock or backshock intensities used in training. J. Exp. Psychol. Anim. Behav. Process. 4, 95–103 (1978).
Park, C., Longo, C.M. & Ackerman, S.L. Genetic and physical mapping of the cerebellar deficient folia (cdf) locus on mouse Chromosome 6. Genomics 69, 135–138 (2000).
Uchida, N. et al. Mouse αN-catenin: two isoforms, specific expression in the nervous system, and chromosomal localization of the gene. Dev. Biol. 163, 75–85 (1994).
Rimm, D.L., Koslov, E.R., Kebriaei, P., Cianci, C.D. & Morrow, J.S. α1-catenin is an actin-binding and -bundling protein mediating the attachment of F-actin to the membrane adhesion complex. Proc. Natl Acad. Sci. USA. 92, 8813–8817 (1995).
Park, C., Finger, J.H., Cooper, J.A. & Ackerman, S.L. The cerebellar deficient folia (cdf) gene acts intrinsically in Purkinje cell migrations. Genesis 32, 32–41 (2002).
Beierbach, E., Park, C., Ackerman, S.L., Goldowitz, D. & Hawkes, R. Abnormal dispersion of a Purkinje cell subset in the mouse mutant cerebellar deficient folia (cdf). J. Comp. Neurol. 436, 42–51 (2001).
Ozol, K., Hayden, J.M., Oberdick, J. & Hawkes, R. Transverse zones in the vermis of the mouse cerebellum. J. Comp. Neurol. 412, 95–111 (1999).
Suzuki, S.C., Inoue, T., Kimura, Y., Tanaka, T. & Takeichi, M. Neuronal circuits are subdivided by differential expression of type-II classic cadherins in postnatal mouse brains. Mol. Cell. Neurosci. 9, 433–447 (1997).
Okabe, M., Ikawa, M., Kominami, K., Nakanishi, T. & Nishimune, Y. 'Green mice' as a source of ubiquitous green cells. FEBS Lett. 407, 313–319 (1997).
Uchida, N., Honjo, Y., Johnson, K.R., Wheelock, M.J. & Takeichi, M. The catenin/cadherin adhesion system is localized in synaptic junctions bordering transmitter release zones. J. Cell Biol. 135, 767–779 (1996).
Fannon, A.M. & Colman, D.R. A model for central synaptic junctional complex formation based on the differential adhesive specificities of the cadherins. Neuron 17, 423–434 (1996).
Colman, D.R. Neurites, synapses, and cadherins reconciled. Mol. Cell. Neurosci. 10, 1–6 (1997).
Serafini, T. An old friend in a new home: cadherins at the synapse. Trends Neurosci. 20, 322–323 (1997).
Radice, G.L. et al. Developmental defects in mouse embryos lacking N-cadherin. Dev. Biol. 181, 64–78 (1997).
Haegel, H. et al. Lack of β-catenin affects mouse development at gastrulation. Development 121, 3529–3537 (1995).
Hirano, S., Kimoto, N., Shimoyama, Y., Hirohashi, S. & Takeichi, M. Identification of a neural α-catenin as a key regulator of cadherin function and multicellular organization. Cell 70, 293–301 (1992).
Nagafuchi, A., Ishihara, S. & Tsukita, S. The roles of catenins in the cadherin-mediated cell adhesion: functional analysis of E-cadherin-α-catenin fusion molecules. J. Cell Biol. 127, 235–245 (1994).
Torres, M. et al. An α-E-catenin gene trap mutation defines its function in preimplantation development. Proc. Natl Acad. Sci. USA 94, 901–906 (1997).
Gotz, M., Wizenmann, A., Reinhardt, S., Lumsden, A. & Price, J. Selective adhesion of cells from different telencephalic regions. Neuron 16, 551–564 (1996).
Provost, E. & Rimm, D.L. Controversies at the cytoplasmic face of the cadherin-based adhesion complex. Curr. Opin. Cell Biol. 11, 567–572 (1999).
Verhage, M. et al. Synaptic assembly of the brain in the absence of neurotransmitter secretion. Science 287, 864–869 (2000).
Maren, S. Neurobiology of Pavlovian fear conditioning. Annu. Rev. Neurosci. 24, 897–931 (2001).
Swerdlow, N.R., Geyer, M.A. & Braff, D.L. Neural circuit regulation of prepulse inhibition of startle in the rat: current knowledge and future challenges. Psychopharmacology 156, 194–215 (2001).
Davis, M. Anatomic and physiologic substrates of emotion in an animal model. J. Clin. Neurophysiol. 15, 378–387 (1998).
Niwa, H., Yamamura, K. & Miyazaki, J. Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene 108, 193–199 (1991).
Morgan, J.G., Dolganov, G.M., Robbins, S.E., Hinton, L.M. & Lovett, M. The selective isolation of novel cDNAs encoded by the regions surrounding the human interleukin 4 and 5 genes. Nucleic Acids Res. 20, 5173–5179 (1992).
Lynn, R.B., Bechtold, L.S. & Miselis, R.R. Ultrastructure of bombesin-like immunoreactive nerve terminals in the nucleus of the solitary tract and the dorsal motor nucleus. J. Auton. Nerv. Syst. 62, 174–182 (1997).
Falls, W.A. Fear-potentiated startle in mice. in Current Protocols in Neuroscience (eds Crawley, J.N., Gerfen, C., McKay, R., Rogawski, M., Sibley, D. & Skolnick, P.) (Wiley, New York, in press).
Acknowledgements
We thank M. Takeichi for the NCAT5 antibody, T. Lufkin for the IRES-lacZ plasmid; J. Miyazaki for the pCAGGS plasmid; Q.-Y. Zheng for ABR analysis; L. Bechtold, P. Finger and R. Bronson for assistance with electron microscopy; the Microchemistry Facility for DNA sequencing; W. Frankel, P. Nishina and J. Willott for comments on the manuscript; R. Burgess for discussions and J. Smith for figure preparation. This work was supported by a grant from the National Institutes of Health (National Institute of Neurological Disorders and Stroke) to S.L.A., a National Center for Research Resources Center of Biomedical Research Excellence grant, an NIH subcontract to W.A.F., an NIH postdoctoral fellowship to J.H.F., a grant from the Molecular Medicine Research Group Program of the Korean Ministry of Science and Technology to C.P. and an institutional Cancer Center Support Grant from the National Cancer Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Park, C., Falls, W., Finger, J. et al. Deletion in Catna2, encoding αN-catenin, causes cerebellar and hippocampal lamination defects and impaired startle modulation. Nat Genet 31, 279–284 (2002). https://doi.org/10.1038/ng908
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng908
This article is cited by
-
Infant inhibited temperament in primates predicts adult behavior, is heritable, and is associated with anxiety-relevant genetic variation
Molecular Psychiatry (2021)
-
Force-dependent allostery of the α-catenin actin-binding domain controls adherens junction dynamics and functions
Nature Communications (2018)
-
Integrative genomic analysis of methylphenidate response in attention-deficit/hyperactivity disorder
Scientific Reports (2018)
-
Biallelic loss of human CTNNA2, encoding αN-catenin, leads to ARP2/3 complex overactivity and disordered cortical neuronal migration
Nature Genetics (2018)
-
Cattle genome-wide analysis reveals genetic signatures in trypanotolerant N’Dama
BMC Genomics (2017)