Abstract
It has been 30 years since it was first proposed that the vertebrate genome evolved through several rounds of genome-wide duplications (polyploidizations)1. Despite rapid advances in genetics, including sequencing of the complete genomes of several divergent species, this hypothesis has not been tested rigorously and is still a matter of debate2. If polyploidizations occurred during chordate evolution, there should be a network of paralogous regions in the present-day jawed vertebrate (Gnathostomata) genomes3. Here we present an investigation of the major histocompatibility complex (MHC) paralogous regions, which we accomplished by characterizing the corresponding region in amphioxus by identifying nine anchor genes and sequencing both the anchor genes and the regions that flank them (a total of 400 kb). Phylogenetic analysis of 31 genes (including the anchor genes) in these regions shows that duplications occurred after the divergence of cephalochordates and vertebrates but before the Gnathostomata radiation. The distribution of human and amphioxus orthologs in their respective genomes and the relationship between these distributions support the en bloc duplication events. Our analysis represents the first step towards demonstrating that the human ancestral genome has undergone polyploidization. Moreover, reconstruction of the pre-duplicated region indicates that one of the duplicated regions retains the ancestral organization.
Similar content being viewed by others
Main
The genomic region in amphioxus that is equivalent to the human MHC paralogous regions has been defined through several steps, including: (i) choosing anchor genes, (ii) cloning their amphioxus equivalents, (iii) isolating the corresponding amphioxus genomic regions and analyzing neighboring genes (especially their phylogenetic relationships to the human genes) and (iv) studying the distribution of the human genes that are orthologous to the amphioxus genes.
We hypothesized that the MHC paralogous regions are the result of en bloc duplications that occurred before the Gnathostomata radiation. After these large-scale duplications, some of the duplicated genes probably returned to single-copy status or translocated to other chromosomal regions. We therefore used two approaches to select the anchor genes. The first approach is based on the analysis of the MHC paralogous regions. To identify paralogous genes that could result from these events, we selected only genes that duplicated in the expected window of time (to avoid having to work with paralogous genes that have different evolutionary histories). The second approach is designed for genes that have returned to single-copy status; this strategy is based on ancient conserved syntenies. Only the MHC region was used for this approach, owing to the lack of information for the other regions in nonmammalian vertebrate species.
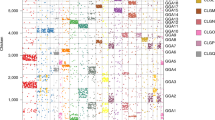
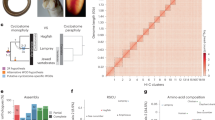
For the first approach, we chose 13 gene families, including AGPAT1,2; BAT1, DDX39; BRD2,3,4,T; C3, C4, C5; CACNA1A,B,E; COL11A2, COL5A1, COL11A1; NOTCH1,2,3,4; PBX1,2,3,4; RAB2L, RALGDS; RXRA,B,G; TNXB, HXB, TNR; VAV1,2,3 and ZNF297, ZNF297B (Fig. 1a). For the second approach, we chose five genes: PSMB8, PSMB9, PSMB10, TAP1 and TAP2 (Fig. 1b).
Two main approaches were used: the analysis of the human MHC paralogous regions and establishing the preserved syntenies between human and bony fishes (Actinopterygii). a , If the paralogous regions are the result of en bloc duplications, the observed conserved syntenies were present in the ancestral genomic region before the en bloc duplication. In the case of the MHC paralogous regions, this event probably occurred at least 420 Myr ago; thus, these syntenies are also at least this old. We expanded upon previous work13 by searching the paralogous regions to find new paralogous genes in the databases and investigating the phylogenetic relationships for all the paralogous genes. To eliminate duplications that would have occurred either too late or too recently, we kept only paralogous genes for which duplication occurred after the divergence of protostomes and deuterostomes and before the Osteichthyes radiation. The four human MHC paralogous regions are presented with the estimated sizes (according to the UCSC database; see Methods). The genes are ordered for the MHC (according to the UCSC database), but the position is arbitrary for the three other MHC paralogous regions. The borders for these last three regions are indicated in the square brackets. Genes of the human chromosome 1 paralogous region are dispersed in two regions on the short and long arms. This split is probably a late event, and the regions were probably linked in an ancestral state14. b , We analyzed sequences from the Human Genome Mapping Project (HGMP) fugu-sequencing project to determine which genes were present in both the human and pufferfish MHC genomic regions. The observed conserved syntenies probably represented an Osteichthyes ancestral state. These conserved syntenies have now been reported15 and have also been found in other teleostean16,17. We also selected the PSMB7,10 family, even though PSMB10 is not located in the human MHC region. It is nevertheless located in the teleost MHC15 and PSMB7 is in a MHC paralogous region, and so it is possible that PSMB10 has been translocated in the human phylum. The distances between the different genes are not shown to scale.
We carried out PCR amplification and cloning of amphioxus orthologs of 11 of 18 of the chosen anchor genes families (BAT1, DDX39; BRD2,3,4,T; C3, C4, C5; CACNA1A,B,E; NOTCH1,2,3,4; PBX1,2,3,4; PSMB7; PSMB8; PSMB9; RAB2L, RALGDS and RXRA,B,G). In each case, we cloned a single amphioxus gene homologous to the vertebrate gene family. We used the PCR products as probes to clone the corresponding genomic region from a Branchiostoma floridae cosmid library, and thus obtained nine genes (amphioxus RAB2L/RALGDS and PSMB8 genes were not present in the library). One cosmid clone for each anchor gene was completely sequenced (except for RXR, with two sequenced cosmid clones) and analyzed for gene content. Forty-one genes were predicted from the ten sequenced cosmid clones (Table 1).
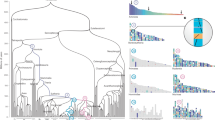
We used the sequences of each predicted gene or gene fragment to search for similar sequences in databases. We observed significant similarity with known genes, expressed-sequence tags (ESTs) or genomic sequences for 34 genes, 33 of which included human sequences. We built data sets with all available sequences, including ESTs and gene predictions (based on genomic sequences). We then carried out phylogenetic analysis and assigned orthology to 31 amphioxus genes: 9 anchor genes and 22 surrounding genes (see Fig. 2 and Web Fig. A online). All the data sets were treated to eliminate reconstruction biases. This was especially relevant, as several biases led to classical artifacts in phylogenetic reconstructions. It should be noted that the phylogenetic reconstructions which we believe to be false were supported by high bootstrap proportions, indicating that in the case of phylogenetic reconstructions that involve paralogous genes, bootstrap proportion may not be a good indicator of the validity of the anlysis.
Data sets were created using all available sequences, including EST and gene predictions (from genomic sequence). We constructed multiple alignments using ClustalX18 with the default parameters. When possible, the analysis was made at the domain level (Pfam domains). We used the partition homogeneity test19 to assess the congruence of two (or more) domains constituting two molecular data sets. When domains were not described, we conserved only the nonambiguous positions of the alignment. The paralogous groups were then analyzed separately using Gu′s program20, and the predicted critical amino-acid sites responsible for the functional divergence between two (or more) paralogous genes were removed. We also tested potential amino-acid composition bias (χ2 test with α=0.05). For the final reconstruction, three methods were used: maximum parsimony using PAUP*4.0 (ref. 21), neighbor-joining22 using MEGA v. 2.0, which is an updated version of MEGA23, and maximum likelihood using Tree-Puzzle 5.0 (ref. 24). The three topologies were compared using the Templeton test. All the trees are rooted at the midpoint unless indicated. Numbers indicate the bootstrap proportions25 (for 1,000 replicate data sets) in the following order (from upper values to lower ones): neighbor-joining, parsimony and maximum likelihood (the values correspond in this case to Quartet Puzzling support values, but they can be interpreted in much the same way as bootstrap proportions). Values less than 50 are indicated with an asterisk. An amphioxus picture represents an amphioxus sequence. Species names represent different groups: Actinopterygii: Cau, Cca, Dre, Ipu, Ola, Omy, Pfl, Pol, Tru; Amphibia: Ame, Rni, Xla; Bacteria: Bfr, Cte, Mvi, Ppr; Chondrichthyes: Dom, Gci; Echinodermata: Lva, Spu; Fungi: Sce, Spo; Hyperortia: Lja, Pma; Hyperotreti: Ebu, Mgl; Mammalia: Bta, Cgr, Chi, Cpo, Hsa, Mfa, Mmu, Ocu, Ptr, Rno, Rra, Ssc; Plants: Ath, Car, Mcr; Protostomians: Aam, Bma, Bmo, Cel, Der, Dme, Dsi, Lbl, Lcu, Lst, Ovo; Sauropsida: Cco, Gga; Nna; and Urochordata: Bsc, Cin, Hro, Pmi. EC, EST contig. Accession numbers of sequences used in this analysis are available at the Université de la Méditerranée website.
From the analysis of the phylogenetic relationships of the nine amphioxus anchor genes (Fig. 2), we conclude that the duplications that gave rise to the different anchor-gene families in vertebrates always occurred after the divergence of cephalochordates and vertebrates, estimated here at 766 Myr ago, but before the bony vertebrate (Osteichtyes) radiation—that is, 420 Myr ago—with two of them having a more restricted duplication range of between 766 Myr and 528 Myr ago (divergence time for the split between cartilaginous fishes (Chondrichthyes) and Osteichtyes4).
To better characterize the duplication process that led to the emergence of the MHC paralogous regions, we mapped the human orthologs of the genes surrounding the amphioxus anchor genes (Fig. 3) and analyzed the statistical significance of their distribution in the four human MHC paralogous regions on chromosomes 1, 6, 9 and 19. We did not include the human orthologs of the amphioxus anchor genes, to eliminate potential bias. Of the 32 genes predicted to be adjacent to the amphioxus anchor genes, 22 have at least one human ortholog. These 22 amphioxus genes have a total of 42 human orthologs. Of these 42 genes, 16 map in an MHC paralogous region. We estimate that the MHC paralogous regions contain 1,240 genes. Distribution of human orthologs to the amphioxus genes neighboring the anchor genes is significantly different (statistical test 1) from a random distribution for these four regions, indicating that there is an evolutionary link between the amphioxus genomic regions and the four human genomic regions. The same observation is reached when we remove the effect of the number of duplications in each gene family (for example, one human ortholog gene for the GPR54 family, five ortholog genes for the SIAT8 family). This observation suggests that the MHC paralogous regions derive from a single genomic region that duplicated en bloc after the cephalochordate/vertebrate split. Nevertheless, one could still argue that the MHC paralogous regions arose from independent duplications followed by independent gene re-localizations, from one genomic region to three others.
The ten sequenced clones are shown. A box represents each gene locus with the arrow indicating the direction of transcription. The introns are not shown. For the PSMB7,10 cosmid clone, two genes are overlapping, as the amphioxus TLR1,2,4,6 gene lies in an intron of the amphioxus PSMB7,10 gene. For each gene, the human orthologs to the amphioxus gene are shown with their respective chromosomal localizations. Anchor genes are indicated in blue and human genes that map in a MHC paralogous region are indicated in green.
To test this possibility, we evaluated the gene duplication history by examining the distribution of the human orthologs of the amphioxus genes that have at least two orthologs, one of which is in an MHC paralogous region (including the anchor genes). This would represent 25 independent duplications. Of the 25 duplicated genes, 19 map within a MHC paralogous region. If the probability of duplication in a given genomic region is the same for all the genomic regions, this possibility is rejected (statistical test 2). Thus, the most likely explanation of the link between the amphioxus genomic regions and the human MHC paralogous regions is that the four human genomic regions have a common genomic region ancestor that has duplicated en bloc after the divergence between cephalochordates and vertebrates. In addition, as the lower boundary for some of the anchor genes is the Gnathostomata speciation, this limit also constitutes the en bloc duplication lower bound. Thus, the MHC paralogous regions are genomic regions that duplicated en bloc between 766 Myr and 528 Myr ago.
Taking into consideration the most likely model to explain the link between the amphioxus regions and the human MHC paralogous regions, and using information from the organization of these genomic regions, we reconstructed the Gnathostomata ancestor proto-MHC region (Fig. 4a). To do this, we used all of the amphioxus cosmid clones sequences (because the duplication between all the anchor genes was compatible with an en bloc duplication process and the significance of the link between the amphioxus regions and the human MHC paralogous regions was calculated using all the genes from all the cosmid clones). None of the initially chosen anchor genes that have not been cloned in the amphioxus have been used, however, as their duplication history could not be established.
a , Putative reconstruction of the MHC paralogous genomic region evolution. By comparing the amphioxus regions and the human MHC paralogous regions and by using the available information described in the text, it is possible to reconstruct the Gnathostomata ancestor proto-MHC. Because the amphioxus regions were not linked, the state of the genomic region in a chordate ancestor is still unknown. The anchor genes are indicated in blue, and the amphioxus anchor-gene surrounding genes and their human orthologs are indicated in green. b , Substitution patterns among the human MHC paralogous genes. We used only the genes that have at least three copies in the MHC paralogous regions. For each of the genes, we compared the substitution rates of the different paralogous genes. The genes indicated in green have a substitution pattern that is significantly slower than the genes indicated in red. The hatched boxes correspond to missing paralogs. The relative position of the genes in the human MHC paralogous region is arbitrary.
The most striking aspect of the evolution of the duplicated regions is the number of fixed genes. Indeed, the genomic region of chromosome 9 contains twice as many genes derived from the ancestral genomic region than any of three other regions (17 versus 7, 8 and 8 for the regions on chromosome 1, 6 and 19 respectively), and this is particularly true for the single-copy genes (9 versus 1, 2 and 1 for the regions on chromosome 1, 6 and 19 respectively). Thus, in terms of gene organization, the region on chromosome 9 is much more similar to the predicted ancestral genomic region than the three other MHC paralogous regions.
The plesiomorphy of the region on chromosome 9 is also noticeable at the gene level. Indeed, analysis of the substitution pattern of gene families having more than two paralogs in the MHC paralogous regions (six paralog families) shows that the genes in the region on chromosome 9 have lower substitution patterns than the three other paralogous regions (six genes over six; Fig. 4b). This is statistically significant (statistical test 3). The genes having fast substitution patterns are spread evenly over the other three regions.
Our study provides strong support for the hypothesis of en bloc duplication of the MHC paralogous regions and provides initial evidence that polyploidization occurred in our genome ancestor between 766 and 528 Myr ago—that is, after the split between cephalochordates and vertebrates and before the Gnathostomata radiation. Moreover, by reconstructing the region ancestral to the four MHC paralogous regions, we have uncovered an unexpected feature of the evolution of the duplicated regions: one region retains an ancestral state both in terms of organization and gene substitution patterns. Thus, the genome contains a large region apparently under negative selection, a concept usually applied to individual genes or gene segments; it will be useful to determine whether this finding can be extended to other regions of the genome. Our analysis underlines the importance of phylogeny and comparative genomics, not only for revealing the evolutionary history of our genome but also for increasing our understanding of genomic organization.
Methods
Definitions.
Two genes are orthologs if they diverged as a result of a speciation event; they are paralogs if they diverged because of a duplication event5. The term 'paralogous regions'refers to genomic regions that contain paralogous genes. The ratio [number of paralogous genes:size of the considered regions] had to be statistically significant so that the genomic regions could be considered paralogous regions (but as this ratio is rarely evaluated, this term is generally vague). Plesiomorphy means close to the ancestral state. Anchor genes are genes that are conserved between distant species and were used in this analysis to define the amphioxus region orthologous with respect to the presumed MHC paralogous regions.
Cloning of the anchor genes.
Adult Branchiostoma lanceolatum were collected near Marseilles, France and adult B. floridae were obtained from G. Luke (Univ. of Reading, UK). Total RNA was provided by F. Coulier (INSERM U119, France) and used for cDNA synthesis by random priming. We prepared DNA with classical protocols6. Branchiostoma belcheri genomic DNA was provided by H. Saiga (Tokyo Metropolitan Univ., Japan). For each selected family, we aligned sequences from all available species and designed degenerate oligonucleotide primers based on the conserved regions. We used the primers to amplify the homologs of these genes from either the Branchiostoma lanceolatum (Bla), floridae (Bfl) or belcheri (Bbe) genomic DNA or from the Bla RNA. In the case of the NOTCH1,2,3,4 family, we used a probe from M. Lardelli (Univ. of Adelaide, Australia), Bfl Amph26 cDNA, to clone the corresponding cosmid clones. For the C3,C4,C5 family, we designed oligonucleotides on a 500-bp EST from Bbe (obtained from S. Miho and N. Satoh, Kyoto University, Japan). For the PSMB7,10 family, we obtained the primer sequence from B. Magor (University of Alberta, Canada). PCR conditions and primer information are available upon request. PCR products at the expected size were gel-purified and directly sequenced with the degenerate oligonucleotides (Genome Express S.A.). We analyzed sequences by phylogenetic analysis or by direct comparison, depending on the sequence sizes.
Cloning of the genomic regions.
We used the B. floridae cosmid genomic DNA library no. 117 from the Resource Center of the German Human Genome (RZPD, Berlin, Germany)7. The library was hybridized according to the RZPD protocols. We confirmed positivity of the clones by colony hybridizations, PCR amplification on purified insert DNA, Southern blots of insert digestions and partial sequencing of the insert with the degenerate oligonucletides. The positive clones are MPMGc117C0356 for BAT1, DDX39; MPMGc117 (G0971, D1293 and A156) for BRD2,3,4,T; MPMGc117C0472 for CACNA1A,B,E; MPMGc117B2065 for C3,C4,C5; MPMGc117 (E1080 and A2375) for NOTCH1,2,3,4; MPMGc117 (N2261, F1971, E1113, I0640, K1330 and H0247) for PBX1,2,3,4; MPMGc117K0849 for PSMB5,8; MPMGc117 (K0348 and N1476) for PSMB7,10; MPMGc117 (C2425 and D1853) for RXRA,B,G(1) and MPMGc117H1057 for RXRA,B,G(2).
Clone sequencing.
Cosmids were sequenced according to previously described protocols8. The names of the sequenced clones are given in Table 1.
Sequence analysis.
The gene nomenclature was set according to NCBI LocusLink and the HUGO gene nomenclature committee. We used the Blast Search Updater perl script9 to search against databases (NR, dbEST and HTGS). We clustered ESTs using the CAP3 sequence assembly program10. We predicted domains using Pfam11 protein search. Transmembrane domains were evidenced using TMHMM12. Details of the phylogenetic analysis, the description of the analysis of each data set and the divergence time estimation are available on the Université de la Méditerranée website (see below). Human gene mapping was achieved by comparing data from the NCBI LocusLink and from the Draft Human Genome Browser of UCSC (University of California of Santa Cruz; 7 October 2000 assembly). For all the analyzed genes, the mappings were compatible (except for GPR54; see the Université de la Méditerranée website). Although the two mappings were closely related, we used the UCSC mapping as it comes from an assembly and is, in most cases, more accurate. The mapping was generally obtained by searching with the official gene symbol, except for several genes for which accession numbers were used (see the Université de la Méditerranée website).
Statistical significance.
All of the genomic regions were defined using the UCSC browser (both for the size and the gene number). We mapped all of the human orthologs to the amphioxus genes. The more distant genes were used to define the genomic region. To get a more accurate estimate for the gene number of each genomic region, we summed the human genome browser known genes and predicted genes. The detail of the estimation as well as all the statistical tests are available at the Université de la Méditerranée website.
GenBank accession numbers.
BAT1, DDX39 cosmid clone, AF391287; BRD2,3,4,T cosmid clone, AF391288; C3,C4,C5 cosmid clone, AF391289; CACNA1A,B,E cosmid clone, AF391290; NOTCH1,2,3,4 cosmid clone, AF391291; PBX1,2,3,4 cosmid clone, AF391292; PSMB5,8 cosmid clone, AF391293; PSMB7,10 clone, AF391294; RXRA,B,G(1) cosmid clone, AF391295; RXRA,B,G(2) cosmid clone, AF391296.
URLs.
HMMER package, http://hmmer.wustl.edu/; Vector NTI Viewer 4.0.1, http://www.informaxinc.com; Université de la Méditerranée website, http://evolution.luminy.univ-mrs.fr/duplications/.
Note: Supplementary information is available on the Nature Genetics website.
References
Ohno, S. Evolution by Gene Duplication (Springer, Berlin, 1970).
Wolfe, K.H. Yesterday′s polyploids and the mystery of diploidization. Nature Rev. Genet. 2, 333–341 (2001).
Lundin, L.G. Evolution of the vertebrate genome as reflected in paralogous chromosomal regions in man and the house mouse. Genomics 16, 1–19 (1993).
Kumar, S. & Hedges, S.B. A molecular timescale for vertebrate evolution. Nature 392, 917–920 (1998).
Fitch, W.S. Distinguishing homologous from analogous proteins. Syst. Zool. 19, 99–113 (1970).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning (Cold Spring Harbor Laboratory Press, 1989).
Zehetner, G. & Lehrach, H., The Reference Library System—sharing biological material and experimental data. Nature 367, 489–491 (1994).
Shiina, T. et al. Genomic anatomy of a premier major histocompatibility complex paralogous region on chromosome 1q21-q22. Genome Res. 11, 789–802 (2001).
Boone, M. & Upton, C. BLAST Search Updater: a notification system for new database matches. Bioinformatics 16, 1054–1055 (2000).
Huang, X. & Madan, A. CAP3: a DNA sequence assembly program. Genome Res. 9, 868–877 (1999).
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Res. 28, 263–266 (2000).
Sonnhammer, E.L., von Heijne, G. & Krogh, A. A hidden Markov model for predicting transmembrane helices in protein sequences. Proc. Int. Conf. Intell. Syst. Mol. Biol. 6, 175–182 (1998).
Kasahara, M., Nakaya, J., Satta, Y. & Takahata, N. Chromosomal duplication and the emergence of the adaptive immune system. Trends Genet. 13, 90–92 (1997).
Maresco, D.L. et al. Localization of FCGR1 encoding Fcgamma receptor class I in primates: molecular evidence for two pericentric inversions during the evolution of human chromosome 1. Cytogenet. Cell Genet. 82, 71–74 (1998).
Clark, M.S., Shaw, L., Kelly, A., Snell, P. & Elgar, G. Characterization of the MHC class I region of the Japanese pufferfish (Fugu rubripes). Immunogenetics 52, 174–185 (2001).
Hansen, J.D., Strassburger, P., Thorgaard, G.H., Young, W.P. & Du Pasquier, L., Exprssion, linkage, and polymorphism of MHC-related genes in rainbow trout, Oncorhynchus mykiss . J. Immunol. 163, 774–786 (1999).
Takami, K., Zaleska-Rutczynska, Z., Figueroa, F. & Klein, J. Linkage of LMP, TAP, and RING3 with Mhc class I rather than class II genes in the zebrafish. J. Immunol. 159, 6052–6060 (1997).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Farris, J.S., Kallersjo, M., Kluge, A.G. & Bult, C. Testing significance of incongruence. Cladistics 10, 315–319 (1994).
Gu, X. Statistical methods for testing functional divergence after gene duplication. Mol. Biol. Evol. 16, 1664–1674 (1999).
Swofford, D.L. PAUP* . Phylogenetic Analysis Using Parsimony (*and other Methods). Version 4. (Sinauer Associates, Sunderland, Massachusetts, 2000).
Saitou, N. & Nei, M. The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Kumar, S., Tamura, K. & Nei, M. MEGA: Molecular Evolutionary Genetics Analysis software for microcomputers. Comput. Appl. Biosci. 10, 189–191 (1993).
Strimmer, K. & von Haeseler, A. Quartet puzzling: a quartet maximum likelihood method for reconstructing tree topologies. Mol. Biol. Evol. 13, 964–969 (1996).
Felsenstein, J. Confidence limits on phylogenenies: an approach using the bootstrap. Evolution 39, 783–791 (1985).
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78–94 (1997).
Solovyev, V. & Salamov, A., The Gene-Finder computer tools for analysis of human and model organisms genome sequences. Proc. Int. Conf. Intell. Syst. Mol. Biol. 5, 294–302 (1997).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Gelfand, M.S., Mironov, A.A. & Pevzner, P.A. Gene recognition via spliced sequence alignment. Proc. Natl Acad. Sci. USA 93, 9061–9066 (1996).
Guigo, R., Agarwal, P., Abril, J.F., Burset, M. & Fickett, J.W. An assessment of gene prediction accuracy in large DNA sequences. Genome Res. 10, 163–1642 (2000).
Acknowledgements
We would like to thank E. Faure for help in cloning the amphioxus RXR gene; M. Flajnik, C. Mawas, F. Birg, J. Berry, D. Birnbaum and L. Du Pasquier for critical reading of the manuscript; E. Danchin for helpful discussions, as well as all the members of the lab for their support; and M. Clark for the Takifugu rubripes BRD2 sequence. L. Abi-Rached is an M.R.T. fellow. This work was supported by INSERM.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Rights and permissions
About this article
Cite this article
Abi-Rached, L., Gilles, A., Shiina, T. et al. Evidence of en bloc duplication in vertebrate genomes. Nat Genet 31, 100–105 (2002). https://doi.org/10.1038/ng855
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng855
This article is cited by
-
Genomic view of the origins of cell-mediated immunity
Immunogenetics (2023)
-
Reconstruction of proto-vertebrate, proto-cyclostome and proto-gnathostome genomes provides new insights into early vertebrate evolution
Nature Communications (2021)
-
Recent genome duplications facilitate the phenotypic diversity of Hb repertoire in the Cyprinidae
Science China Life Sciences (2021)
-
Transcriptome Ortholog Alignment Sequence Tools (TOAST) for phylogenomic dataset assembly
BMC Evolutionary Biology (2020)
-
Deeply conserved synteny resolves early events in vertebrate evolution
Nature Ecology & Evolution (2020)