Abstract
The knowledge of human genetic variation that will come from the human genome sequence makes feasible a polygenic approach to disease prevention, in which it will be possible to identify individuals as susceptible by their genotype profile and to prevent disease by targeting interventions to those at risk. There is doubt, however, regarding the magnitude of these genetic effects and thus the potential to apply them to either individuals or populations. We have therefore examined the potential for prediction of risk based on common genetic variation using data from a population-based series of individuals with breast cancer. The data are compatible with a log-normal distribution of genetic risk in the population that is sufficiently wide to provide useful discrimination of high- and low-risk groups. Assuming all of the susceptibility genes could be identified, the half of the population at highest risk would account for 88% of all affected individuals. By contrast, if currently identified risk factors for breast cancer were used to stratify the population, the half of the population at highest risk would account for only 62% of all cases. These results suggest that the construction and use of genetic-risk profiles may provide significant improvements in the efficacy of population-based programs of intervention for cancers and other diseases.
Similar content being viewed by others
Introduction
The human genome sequence is the starting point for the compilation of detailed information about the range of genetic differences among individuals. Whereas current approaches to the genetics of disease are based primarily on mendelian inheritance of single but uncommon predisposing genes, knowledge of the range of genetic variation across many loci in the population will allow a polygenic approach in which risks will be estimated from the combined effect of this variation. The prospect of a polygenic approach to common diseases has generated much attention. Some have claimed1,2 that greater understanding of genetic risk factors and their interactions with the environment will allow diseases to be predicted and to be prevented at both individual and population levels, by directing interventions at individuals shown to be at high risk. Others are less sure3,4,5; in particular, they question whether molecular testing for common genetic variants can have sufficient predictive power to be of practical use either for the individual or for defining risk groups in the population at large.
We therefore examined the potential for prediction of risk based on common genetic variation and compared this with the predictions that could be made using established risk factors. We addressed three questions: (i) What is the likely distribution of genetically determined risk in the population? (ii) What is the distribution of risk described by established risk factors? (iii) What are the implications of these risk distributions for effective targeting of intervention to individuals and within the population? We used breast cancer as a model to explore these questions.
Results
Distribution of genetic risk in population
Breast cancer, like other common cancers, shows familial clustering. Depending on age, the risk is typically increased by two- to threefold in first-degree relatives of an affected individual. Twin studies suggest that most of this familial aggregation results from inherited susceptibility6,7. The known predisposing genes for breast cancer, including BRCA1 and BRCA2, account for only 20–25% of this effect8. The number and properties of the genes that account for the remainder are unknown.
To investigate genetic models that best account for the familial aggregation of breast cancer not due to BRCA1 and BRCA2, we analyzed the occurrence of breast cancer in the relatives of affected individuals in the Anglian Breast Cancer Study, a population-based series of 1,484 affected individuals, all of whom were screened for mutations in BRCA1/2 (refs 9,10). Two models were found to fit the data well. The one best describing these data was a polygenic model in which susceptibility to breast cancer is conferred by a large number of alleles. The risk associated with any individual allele is small, but as the effects are multiplicative, a woman with several susceptibility alleles is at high risk. The model also fits well the pattern of breast cancer not due to BRCA mutations11 in a series of multiple-case families. It is probably an appropriate model for many common cancers and other diseases. The second model was that of a single common recessive allele (frequency 0.24). This model fit the population-based data well, but the fit for the multiple-case families was not as good. The risk allele was estimated to confer a relative risk of 21 for rare homozygotes compared with common homozygotes and heterozygotes, corresponding to a moderately high penetrance of 42%.
Let us assume that one dominant susceptibility allele has been identified, and half the population carry the high risk allele. If 'high-risk' individuals have a breast cancer relative risk of 3 compared with 'low-risk' individuals, the high-risk group will have a relative risk of 1.5, and the low risk group will have a relative risk of 0.5 (1.5=3 × 0.5), giving an average relative risk of 1 for the population. Absolute risks of breast cancer will depend on underlying incidence rates. Using rates typical of northern Europe and the United States, the absolute risk of breast cancer by age 70 in the two risk groups is 2.9% and 8.4%, corresponding to an average population risk of 5.7% (ref. 12). It thus follows that 75% of all breast cancer cases will occur in high-risk women and 25% in low-risk women. An intervention targeted to high-risk women thus has the potential to reduce breast cancer morbidity by a maximum of 75%. Now let us assume there are two such susceptibility genes, each present in half the population and each conferring the same risks, which interact in a multiplicative manner. There will now be three risk groups in the population: one-quarter of the population will have no high-risk alleles and a relative risk of 0.25 (0.5 × 0.5), half the population will have one high-risk allele (relative risk of 0.75) and one-quarter of the population will have two high-risk alleles (relative risk of 2.25). The risk of breast cancer by age 70 in these three groups is 1.5%, 4.3% and 12.3%, respectively; the groups account for 8%, 23% and 69%, respectively, of all breast cancer cases. As the number of susceptibility genes increases, the number of risk groups will increase and risk in the population tends towards a continuous distribution. For example, with five susceptibility alleles, each conferring a different relative risk, there will be 32 different risk groups.
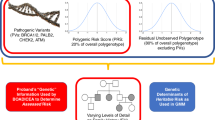
Under our polygenic model, the (continuous) distribution of risk in the population is predicted to be log-normal; that is, the logarithm of risk for all individuals in the population will follow a normal distribution (Fig. 1). A normal distribution is defined by its mean value and its standard deviation. In our analysis, the standard deviation of the log-normal distribution of genetic risk was estimated to be 1.2 (ref. 10). Once the standard deviation is defined, the mean of the distribution is set so that the arithmetical average risk (termed R in equations) is equal to 1 (see Methods). The standard deviation describes the variation in risks that can be defined within the population and thus is the key indicator of the power to discriminate individuals in groups at low or high risk.
Risks are shown on a log scale; the arithmetical average risk for the entire population has been set at 1.0 (see Methods). The risk distribution in individuals who will develop breast cancer (cases) is shifted to the right. The standard deviation describes the spread of risk between high and low values within the population, and thus the potential to discriminate different levels in different individuals.
It can be shown that the distribution of (initial) risk among affected individuals is also log-normal and has a simple relationship to the distribution of risk in the population (see Methods). The standard deviation of the log-normal distribution in individual cases is the same as in the population, but the average risk is higher (Fig. 1). (A complication occurs, because at older ages the distribution of risk in both the general population and among individual cases changes, as higher-risk individuals are more likely to have been 'eliminated'. We have allowed for this tendency and estimated the distribution of risk in individual cases as a whole averaged over all ages.)
The distribution of genetic risk in the population, for a standard deviation of 1.2, is shown in Fig. 1. The area under the curve gives the proportion of the population in any risk group. The risk to the highest quintile of the distribution is 40-fold higher than that of the lowest quintile. Also shown is the risk distribution in affected individuals according to their initial level of risk; that is, the risk distribution of women in the population who will subsequently develop breast cancer.
The proportion of the population that have a risk greater than a given level, and the proportion of cases that will occur within this high-risk subgroup, provide more useful information. These figures are obtained from the area under the population and case curves (Fig. 1) to the right of any given risk cut-off. Thus, for a standard deviation of 1.2, half the population have a relative risk of 0.46 or higher; this half of the population accounts for 88% of all individuals with breast cancer. Figure 2 shows the proportion of the population that have a risk above a given level and the proportion of cases above the same level of risk, where the risks have been converted to risk of breast cancer by age 70. From these curves, it can be seen that the 12% of the population at highest risk account for 50% of affected individuals. Another way of depicting these data is to plot the proportion of cases that occur in women above a given level of risk against the proportion of the population above that level of risk (Fig. 3).
Under the recessive model, there would be a single risk allele with a frequency of 0.24. Women with the at-risk genotype (RR=9.8) would thus comprise 5.8% of the population and account for 56% of cases. The remaining 94.2% of the population (RR=0.46) would account for 44% of cases. In light of the failure of genetic linkage studies to identify further breast cancer susceptibility genes, however, such a model seems less plausible than a polygenic model of inheritance13,14.
Distribution of risk due to established risk factors
Several risk factors that do not require molecular genotyping (although they may in part have a genetic basis) are already used to stratify individuals into high- and low-risk groups for breast cancer. We have used data on 3,209 individuals with breast cancer, from the same population-based study that we used for the genetic risk estimates, to estimate the risk distribution for breast cancer in pre-menopausal women that is provided by these established risk factors15. We included age at menarche (under 13 years versus 13 years and older), number of full-term pregnancies, age at first full-term pregnancy, oral contraception use (current, past, never) and family history. The relative risk associated with each factor was assumed to act independently, with several risk factors interacting multiplicatively. The distribution of relative risk in the affected individuals was found to be approximately log-normally distributed, with a standard deviation of 0.3. It follows that the distribution of risk in the population determined by these risk factors should also be log-normal, with the same standard deviation. The width of this distribution corresponds to a 3.5-fold difference in risk between the highest quintile to the lowest, but it is considerably narrower than the distribution, with a standard deviation of 1.2 predicted for genetic risk factors.
In practice, the estimated power of the genetic risk distribution is an upper limit, because some disease-associated genes may prove difficult to detect; this will reduce the width of the distribution and thus the predictive value. We therefore recalculated the model assuming that genes responsible for half the variation in genetic risk could be identified. The risk to the highest quintile of the distribution is now 12-fold higher than that of the lowest quintile (Table 1). The discriminatory power of the distribution based on established risk factors with a standard deviation of 0.3 is also compared with that of risk factors with a standard deviation of 1.2 (Fig 3).
Discussion
The validity of our results depends on the validity of the segregation analysis10, a detailed critique of which is beyond the scope of this paper. The only other segregation analysis that has taken into account the contribution of BRCA1 and BRCA2 to familial aggregation found evidence of a single high-penetrance recessive gene16. We found that a recessive model fit the population-based data well10, but the fit for the multiple-case families was not as good11. In addition, a recent meta-analysis found that the familial risk of breast cancer to siblings is similar to that to mothers, suggesting that any recessive component is at best small.
Thus, the possibility that genetic susceptibility to breast cancer is due to several loci, each conferring a modest independent risk, seems reasonable. In practice, the number of loci involved will be finite, but once there are more than four to five loci the distribution of risk will be similar to that of the polygenic model, except at the extreme tails. A key aspect of the model is standard deviation, as this determines the power of the risk distribution to discriminate high- and low-risk individuals. The estimate of standard deviation is specified by the segregation analysis and is also close to that predicted by other studies of familial risk. Assuming the relative risk of breast cancer in siblings to be equal to 217, the predicted standard deviation of the (log) genetic risk distribution is also 1.2 (see Methods). The familial relative risk for many other common cancers is also around 2, which suggests that the distribution of risk for these cancers will be similar to that which we observed for breast cancer. The potential benefits of an approach targeted to high-risk disease prevention are thus also likely to be similar.
The assumption that the putative polygenes act in a multiplicative manner may not be correct. It has been argued, on the basis of the ratio of risks to monozygotic and dizygotic twins of cancer cases, that an additive model provides the best fit for most common cancers, including breast cancer14. The effect of an additive model would be to reduce the standard deviation of the risk distribution from 1.2 to 1.05, reducing slightly the predictive power of genetic testing. By contrast, a recent analysis of twin data reported by Peto and Mack7 found a very high incidence in the monozygotic twin of affected individuals, more consistent with a muliplicative model, and concluded that a high proportion, and perhaps the majority, of breast cancers arise in a susceptible minority of women. If this were true, the discriminatory power could be substantially improved.
The practical use of risk information has previously been considered in two contexts: that of the individual, and that of the population18. In both cases, our analysis suggests that a risk profile that is based on the combination of known genotype and other risk factors is likely to provide risk discrimination that has practical value for health care. Whether genetic testing in whole populations would be socially or economically acceptable remains unknown, and will probably depend on whether useful action can be seen to result. But it does seem clear that using combinations of risk factors may overcome many of the limitations of using single risk factors, which has caused skepticism regarding the usefulness of molecular genotyping for common, low-risk genes4.
For example, with respect to individual risk, a single gene that conferred a relative risk of breast cancer of 1.5-fold (the size of effect that seems plausible from reported studies19) would increase the risk of breast cancer by age 70 from 5.7% to 85% for an individual from the UK. By contrast, a genotypic risk profile might identify one woman in 30 who has a risk by age 70 of 20% or more (Fig. 2). Little is known about how individuals will perceive and respond to such risks, but the discriminatory power of the polygenic risk profile is clear.
At the population level, the effects are even more striking. Under the genetic model, 12% of the population have a risk of breast cancer of 1 in 10 or more by age 70, and half the total breast cancer incidence falls within that 12% of the population (Fig. 2). Different cut-offs can be chosen, to suit a specific purpose, to give the best combination of high risk and proportion of total breast cancer incidence that is included within the high-risk group. A single genotypic marker would, by contrast, provide far weaker discrimination; for example, a dominant predisposing allele with frequency 10% and relative risk of 1.5 would result in 26% of cancer incidence occurring in the 19% of the population who carried at least one allele—very poor enrichment, in terms of targeting interventions. An important feature of the high-risk groups defined by the model is that most of the individuals within them will be at risk because of the combined effect of several predisposing alleles. This implies that interventions that are based on specific mechanisms of predisposition will individually deal with only a proportion of the excess cancer risk, and that except for predisposing genes with major effects, generic interventions are more likely to be appropriate.
Risk profiles may also be used to define low-risk groups. Only 12% of breast cancer incidence falls within the 50% of women at lowest risk (Fig. 2). Exclusion of low-risk groups from interventions, if it were socially acceptable, might be cost-effective. For example, screening of the whole population by mammography should reduce breast cancer mortality by approximately 30% (ref. 20). If mammography were offered only to the half of the population in the highest-risk group (per the genetic profile), total mortality from breast cancer would still be reduced by 26%—a 'loss' of only 4%. There would be additional benefits, as the benefit:harm ratio is likely to be improved by targeting to the high-risk group. (These arguments assume that the efficacy of any intervention is independent of genotype; if that is not the case, the benefit of genotyping may be greater or smaller, depending on whether the cancers in high-risk individuals are more or less responsive to the intervention).
These arguments and examples assume that all of the genetic factors that contribute to the estimated risk distribution can be identified and typed. In practice, this goal is some way off. Nevertheless, the results (Table 1) suggest that even if only half the risk factors were typed, useful discrimination of risk might be possible. Our findings also suggest that the power of risk profiles based solely on the currently available 'classical' risk factors is quite weak, and that, in the future, genotypic data may have a decisive role.
Methods
Equations.
Under the polygenic model, the distribution of risk (R) in the population at birth is log-normal. The distribution is defined by the mean value (μ) and its standard deviation (σ). The parameter μ is an arbitrary constant; in our examples we set μ=−σ2/2 so that the arithmetical average population risk (R) is equal to 1.
The distribution of risk among cases of the disease at young ages is also log-normal and has a simple relationship to the distribution of risk in the population (μ and σ).
For a log-normally distributed risk R in the population,

that is, the density of Y, the logarithm of the risk, is given by:

As risk of disease is proportional to R (= ey), the distribution of risk (on the y scale) in cases is given by:

This is also a normal distribution with parameters (μ+σ2,σ2). Thus, the risk distribution in the cases has the same shape as in the general population, but shifted (on a log-scale) by σ2.
The RR of disease in monozygotic twins (λmonozygotic) and siblings (λsibling) are related to each other and to the predicted s.d. of the polygenic log-normal risk distribution by the equation:

Assuming λsibling to be equal to 2, as estimated by many observational epidemiologic studies17, this equation solves to predict an s.d. of 1.2.
References
Bell, J. The new genetics in clinical practice. Br. Med. J. 316, 618–620 (1998).
Beaudet, A.L. 1998 ASHG presidential address. Making genomic medicine a reality. Am. J. Hum. Genet. 64, 1–13 (1999).
Friend, S.H. How DNA microarrays and expression profiling will affect clinical practice. Br. Med. J. 319, 1306–1307 (1999).
Holtzman, N.A. & Marteau, T.M. Will genetics revolutionize medicine? N. Engl. J. Med. 343, 141–144 (2000).
Vineis, P., Schulte, P. & McMichael, A.J. Misconceptions about the use of genetic tests in populations. Lancet 357, 709–712 (2001).
Lichtenstein, P. et al. Environmental and heritable factors in the causation of cancer—analyses of cohorts of twins from Sweden, Denmark, and Finland. N. Engl. J. Med. 343, 78–85 (2000).
Peto, J. & Mack, T.M. High constant incidence in twins and other relatives of women with breast cancer. Nature Genet. 26, 411–414 (2000).
Easton, D.F. How many more breast cancer predisposition genes are there? Breast Cancer Res. 1, 14–17 (1999).
Anglian Breast Cancer Study Group. Prevalence and penetrance of BRCA1 and BRCA2 in a population based series of breast cancer cases. Br. J. Cancer 83, 1301–1308 (2000).
Antoniou, A.C. et al. Evidence for further breast cancer susceptibility genes in addition to BRCA1 and BRCA2 in a population based study. Genet. Epidemiol. 21, 1–18 (2001).
Antoniou, A.C. et al. A comprehensive model for familial breast cancer incorporating BRCA1, BRCA2 and other genes. Br. J. Cancer in press (2001).
Pharoah, P.D.P. & Mackay, J. Absolute risk of breast cancer in women at increased risk: a more useful clinical measure than relative risk? The Breast 7, 255–259 (1998).
Vaittinen, P. & Hemminki, K. Risk factors and age-incidence relationships for contralateral breast cancer. Int. J. Cancer 88, 998–1002 (2000).
Risch, N. The genetic epidemiology of cancer: interpreting family and twin studies and their implications for molecular genetic approaches. Cancer. Epidemiol. Biomarkers Prev. 10, 733–741 (2001).
Kelsey, J.L., Gammon, M.D. & John, E.M. Reproductive factors and breast cancer. Epidemiol. Rev. 15, 36–47 (1993).
Cui, J. et al. After BRCA1 and BRCA2—what next? Multifactorial segregation analyses of three-generation, population-based Australian families affected by female breast cancer. Am. J. Hum. Genet. 68, 420–431 (2001).
Pharoah, P.D.P., Day, N.E., Duffy, S., Easton, D.F. & Ponder, B.A.J. Family history and the risk of breast cancer: a systematic review and meta-analysis. Int. J. Cancer 71, 800–809 (1997).
Rose, G. Sick individuals and sick populations. Int. J. Epidemiol. 14, 32–38 (1985).
Dunning, A.M. et al. A systematic review of genetic polymorphisms and breast cancer risk. Cancer. Epidemiol. Biomarkers Prev. 8, 843–854 (1999).
Kerlikowske, K., Grady, D., Rubin, S.M., Sandrock, C. & Ernster, V.L. Efficacy of screening mammography. A meta-analysis. J.A.M.A. 273, 149–154 (1995).
Acknowledgements
P.P. is a Senior Clinical Research Fellow, D.F.E. is a Principal Fellow and B.A.J.P. is a Gibb Fellow of Cancer Research UK. The Public Health Genetics Unit is funded by the Eastern Regional Office of the National Health Service Executive. Research in the Strangeways Laboratories is supported by a grant from the National Lottery Board and by program grants from Cancer Research UK and the Medical Research Council. We thank H. Burton, C. Brayne, M. Austin and N. Day for their helpful comments on early drafts of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pharoah, P., Antoniou, A., Bobrow, M. et al. Polygenic susceptibility to breast cancer and implications for prevention. Nat Genet 31, 33–36 (2002). https://doi.org/10.1038/ng853
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng853