Abstract
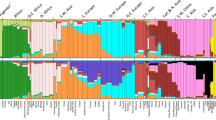
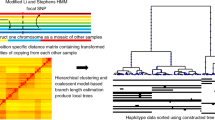
Large data sets on human genetic variation have been collected recently, but their usefulness for learning about history and natural selection has been limited by biases in the ways polymorphisms were chosen. We report large subsets of SNPs from the International HapMap Project1,2 that allow us to overcome these biases and to provide accurate measurement of a quantity of crucial importance for understanding genetic variation: the allele frequency spectrum. Our analysis shows that East Asian and northern European ancestors shared the same population bottleneck expanding out of Africa but that both also experienced more recent genetic drift, which was greater in East Asians.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
The International HapMap Consortium. The Phase II HapMap. (in the press).
Adams, A.M. & Hudson, R.R. Maximum-likelihood estimation of demographic parameters using the frequency spectrum of unlinked single-nucleotide polymorphisms. Genetics 168, 1699–1712 (2004).
Marth, G.T., Czabarka, E., Murvai, J. & Sherry, S.T. The allele frequency spectrum in genome-wide human variation data reveals signals of differential demographic history in three large world populations. Genetics 166, 351–372 (2004).
Pluzhnikov, A., Di Rienzo, A. & Hudson, R.R. Inferences about human demography based on multilocus analyses of noncoding sequences. Genetics 161, 1209–1218 (2002).
Reich, D.E. et al. Linkage disequilibrium in the human genome. Nature 411, 199–204 (2001).
Voight, B.F. et al. Interrogating multiple aspects of variation in a full resequencing data set to infer human population size changes. Proc. Natl. Acad. Sci. USA 102, 18508–18513 (2005).
Clark, A.G., Hubisz, M.J., Bustamante, C.D., Williamson, S.H. & Nielsen, R. Ascertainment bias in studies of human genome-wide polymorphism. Genome Res. 15, 1496–1502 (2005).
Nielsen, R., Hubisz, M.J. & Clark, A.G. Reconstituting the frequency spectrum of ascertained single-nucleotide polymorphism data. Genetics 168, 2373–2382 (2004).
Hinds, D.A. et al. Whole-genome patterns of common DNA variation in three human populations. Science 307, 1072–1079 (2005).
Garrigan, D., Mobasher, Z., Kingan, S.B., Wilder, J.A. & Hammer, M.F. Deep haplotype divergence and long-range linkage disequilibrium at xp21.1 provide evidence that humans descend from a structured ancestral population. Genetics 170, 1849–1856 (2005).
Williamson, S.H. et al. Simultaneous inference of selection and population growth from patterns of variation in the human genome. Proc. Natl. Acad. Sci. USA 102, 7882–7887 (2005).
Stephens, M., Sloan, J.S., Robertson, P.D., Scheet, P. & Nickerson, D.A. Automating sequence-based detection and genotyping of SNPs from diploid samples. Nat. Genet. 38, 375–381 (2006).
Prugnolle, F., Manica, A. & Balloux, F. Geography predicts neutral genetic diversity of human populations. Curr. Biol. 15, R159–R160 (2005).
Ramachandran, S. et al. Support from the relationship of genetic and geographic distance in human populations for a serial founder effect originating in Africa. Proc. Natl. Acad. Sci. USA 102, 15942–15947 (2005).
Conrad, D.F. et al. A worldwide survey of haplotype variation and linkage disequilibrium in the human genome. Nat. Genet. 38, 1251–1260 (2006).
Bowcock, A.M. et al. Drift, admixture, and selection in human evolution: a study with DNA polymorphisms. Proc. Natl. Acad. Sci. USA 88, 839–843 (1991).
Weir, B.S. & Cockerham, C.C. Estimating F-statistics for the analysis of population structure. Evolution Int. J. Org. Evolution 38, 1358–1370 (1984).
Becquet, C., Patterson, N., Stone, A.C., Przeworski, M. & Reich, D. Genetic structure of chimpanzee populations. PLoS Genet 3, e66 (2007) (doi:10.1371/journal.pgen.0030066).
Mellars, P. Going east: new genetic and archaeological perspectives on the modern human colonization of Eurasia. Science 313, 796–800 (2006).
Hewitt, G. The genetic legacy of the Quaternary ice ages. Nature 405, 907–913 (2000).
Plagnol, V. & Wall, J.D. Possible ancestral structure in human populations. PLoS Genet. 2, e105 (2006) (doi:10.1371/journal.pgen.0020105).
Lahr, M.M. & Foley, R. Multiple dispersals and modern human origins. Evol. Anthropol. 3, 48–60 (2005).
Stringer, C.B., Grun, R., Schwarcz, H.P. & Goldberg, P. ESR dates for the hominid burial site of Es Skhul in Israel. Nature 338, 756–758 (1989).
Vanhaereny, M. et al. Middle Paleolithic shell beads in Israel and Algeria. Science 312, 1785–1788 (2006).
Ning, Z., Cox, A.J. & Mullikin, J.C. SSAHA: a fast search method for large DNA databases. Genome Res. 11, 1725–1729 (2001).
The Chimpanzee Sequencing and Analysis Consortium. Initial sequence of the chimpanzee genome and comparison with the human genome. Nature 437, 69–87 (2005).
Lagarias, J.C., Reeds, J.A., Wright, M.H. & Wright, P.E. Convergence properties of the Nelder-Mead simplex method in low dimensions. SIAM J. Optim. 9, 112–147 (1998).
Lahiri, S.N. Resampling Methods for Dependent Data (Springer, New York, 2003).
Venter, J.C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Acknowledgements
We thank D. Altshuler, M. Bernstein, A. Keinan, E. Lander, M. Mandel, M. Mirazon Lahr, A. Price, M. Przeworski, S. Schaffner, C. Stringer and B. Weir for discussions, comments and assistance with stages of this study. We are grateful to G. Marth for sharing the multi-epoch model source code with us. Orangutan sequence traces were produced by the Genome Sequencing Center at Washington University School of Medicine (ftp://ftp.ncbi.nih.gov/pub/TraceDB/pongo_pygmaeus_abelii); we thank R. Wilson for permission to use these data. Sequence traces for the human ABC libraries used in this study were produced by Agencourt Biosciences Corporation and were obtained from ftp://ftp.ncbi.nih.gov/pub/TraceDB/homo_sapiens; we thank D. Smith and E. Eichler for permission to use these data. A.K. was supported by the Rothschild fellowship from Yad Hanadiv foundation. J.C.M. was supported by the Intramural Research Program of the National Human Genome Research Institute, National Institutes of Health (NIH). N.P. was supported by a career transition award from the NIH. D.R. was supported by NIH grant U01 HG004168 and a Burroughs Wellcome Career Development Award in the Biomedical Sciences.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Tables 1–5, Supplementary Figures 1 and 2, Supplementary Note (PDF 1007 kb)
Rights and permissions
About this article
Cite this article
Keinan, A., Mullikin, J., Patterson, N. et al. Measurement of the human allele frequency spectrum demonstrates greater genetic drift in East Asians than in Europeans. Nat Genet 39, 1251–1255 (2007). https://doi.org/10.1038/ng2116
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng2116