Abstract
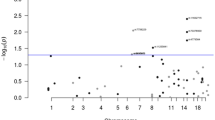
Using a multistage genetic association approach comprising 7,480 affected individuals and 7,779 controls, we identified markers in chromosomal region 8q24 associated with colorectal cancer. In stage 1, we genotyped 99,632 SNPs in 1,257 affected individuals and 1,336 controls from Ontario. In stages 2–4, we performed serial replication studies using 4,024 affected individuals and 4,042 controls from Seattle, Newfoundland and Scotland. We identified one locus on chromosome 8q24 and another on 9p24 having combined odds ratios (OR) for stages 1–4 of 1.18 (trend; P = 1.41 × 10−8) and 1.14 (trend; P = 1.32 × 10−5), respectively. Additional analyses in 2,199 affected individuals and 2,401 controls from France and Europe supported the association at the 8q24 locus (OR = 1.16, trend; 95% confidence interval (c.i.): 1.07–1.26; P = 5.05 × 10−4). A summary across all seven studies at the 8q24 locus was highly significant (OR = 1.17, c.i.: 1.12–1.23; P = 3.16 × 10−11). This locus has also been implicated in prostate cancer1,2,3.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Gudmundsson, J. et al. Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat. Genet. 39, 631–637 (2007).
Haiman, C.A. et al. Multiple regions within 8q24 independently affect risk for prostate cancer. Nat. Genet. 39, 638–644 (2007).
Yeager, M. et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat. Genet. 39, 645–649 (2007).
Lichtenstein, P. et al. Environmental and heritable factors in the causation of cancer–analyses of cohorts of twins from Sweden, Denmark and Finland. N. Engl. J. Med. 343, 78–85 (2000).
Cotterchio, M. et al. Ontario familial colon cancer registry: methods and first-year response rates. Chronic Dis. Can. 21, 81–86 (2000).
Fan, J.B. et al. Illumina universal bead arrays. Methods Enzymol. 410, 57–73 (2006).
Greenwood, C.M.T., Rangrej, J. & Sun, L. Optimal selection of markers for validation from genome-wide association studies. Genet. Epidemiol. 31, 396–407 (2007).
DerSimonian, R. & Laird, N. Meta-analysis in clinical trials. Control. Clin. Trials 7, 177–188 (1986).
Amundadottir, L.T. et al. A common variant associated with prostate cancer in European and African populations. Nat. Genet. 38, 652–658 (2006).
Wang, Y. et al. Allele quantification using molecular inversion probes (MIP). Nucleic Acids Res. 33, e183 (2005).
Nicolae, D.L., Wen, X., Voight, B.F. & Cox, N.J. Coverage and characteristics of the Affymetrix GeneChip Human Mapping 100K SNP set. PLoS Genet. 2, e67 (2006).
Sasieni, P.D. From genotypes to genes: doubling the sample size. Biometrics 53, 1253–1261 (1997).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Royal Stat. Soc. B 59, 289–300 (1995).
Acknowledgements
The authors thank D. Daftary and T. Selander at the Mount Sinai Hospital Biorepository for technical and administrative assistance with OFCCR samples. We also are grateful to A. Bélisle, S. Roland, M.-C. Tessier and D. Vincent of the McGill University and Génome Québec Innovation Centre for technical genotyping assistance. We acknowledge all those involved in recruitment and assembly of the biological and data resources of the Colorectal Cancer Genetic Susceptibility (COGS) study and the Scottish Colorectal Cancer Study (SOCCS), including the Edinburgh Wellcome Trust Clinical Research Facility and also the Family Practitioner Services Department, the Cancer Intelligence Unit of the Information and Statistics Division (ISD) and Scottish Cancer Registry Cancer, all of the Scottish Central National Health Service (NHS). C. Bonithon-Kopp, A. Pariente, B. Nalet and J. Lafon (Group d'Etude des Adenomas) and members of the Association Nationale des Gastroenterologues des Hopitaux Généraux. For administrative assistance, we acknowledge L. Blahut (Cancer Care Ontario). We are grateful to the nursing, laboratory and office staff throughout Edinburgh, at the Wellcome Trust Clinical Research Facility and at the central Scottish NHS departments, including Cancer Registry and the Scottish Cancer Intelligence Unit of ISD. Cancer Care Ontario, as the host organization to the ARCTIC Genome Project, acknowledges that this Project was funded by Genome Canada through the Ontario Genomics Institute, by Génome Québec, the Ministère du Dévelopement Économique et Régional et de la Recherche du Québec and the Ontario Institute for Cancer Research (B.W.Z., T.J.H., C.M.T.G, S.G. and M.C.). This work was supported through collaboration and cooperative agreements with the Colon Cancer Family Registry and principal investigators supported by the US National Cancer Institute, US National Institutes of Health under RFA CA-95-011, including S.G. at the Ontario Registry for Studies of Familial Colorectal Cancer (U01 CA076783) and J.P. at the Seattle Colorectal Cancer Family Registry (U01 CA074794). The content of this manuscript does not necessarily reflect the views or policies of the National Cancer Institute or any of the collaborating institutions or investigators in the Colon Familial Registry, nor does mention of trade names, commercial products, or organizations imply endorsement by the US government or the Colon CFR. This work was supported through collaboration and cooperative agreement with the Newfoundland Familial Colon Cancer Registry at the Memorial University of Newfoundland (B.Y., R.G., J.G.). The work in Scotland was supported by Cancer Research UK (C348/A3758, C48/A6361), the UK Medical Research Council (G0000657-53203) and the Scottish Executive Chief Scientist's Office (K/OPR/2/2/D333, CSO CZB/4/94) (M.D.) and by a Centre Grant from the Digestive Disorders Foundation (http://www.corecharity.org.uk). Support in France came from the French Ministry of Research, Fondation de France (S.O., C.B.P.), Projet Hospitalier de Researche Clinique (PHRC) AOM01-006 (G.T.), Ligue Nationale contre le Cancer (G.T.), Groupement des Enterprises Francaises dans la Lutte contre le Cancer (GEFLUC) (S.B.) and the European Commission (E.R.). Support is acknowledged from the National Program for Complex Data Structures (Canada) (R.K.) and the Centre for Applied Genomics (Toronto) (C.G.).
Author information
Authors and Affiliations
Contributions
B.W.Z., C.M.T.G., R.K., S.G., M.C. and T.J.H. devised and executed the original study design and prepared this manuscript. J.R., A.T., S.M.F., J.P., S.O., T.C., E.C., V.F., P.L., S.S., S.R., J.-F.O., F.R. and A.M. assisted in data analysis. R.S., P.C., S.B., A.M.O., G.Z., M.J., J.M., B.Y., R.G., J.G., M.E.M.P., H.C., H.B., M.S., E.T., C.B.-P., B.B., E.R., S.K. and S.J.C. provided insights and G.T., M.G.D., J.P., P.N. and E.P. assisted in sample procurement and study design and interpretation. A.T., S.M.F., J. Prendergast, M.E.M.P., H.C. and M.G.D. designed and undertook stages 3 and 4 of the study using the Scottish sample and data resource and contributed to writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
B.W.Z. is the founder and equity holder of Arctic Dx, a new Canadian biotechnology company that has received a license in the field of diagnostics for intellectual property generated by several Canadian institutions involved in this research. Arctic Dx currently has no financiers or employees.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1, Supplementary Table 3, Supplementary Table 4, Supplementary Figure 1, Supplementary Methods (PDF 151 kb)
Supplementary Table 2
Phase 2 validation genotyping at 76 sites. (XLS 157 kb)
Supplementary Table 5
Selected SNPs genotyped using the Illumina GoldenGate Assay in stage 1 of the ARCTIC project. (XLS 197 kb)
Rights and permissions
About this article
Cite this article
Zanke, B., Greenwood, C., Rangrej, J. et al. Genome-wide association scan identifies a colorectal cancer susceptibility locus on chromosome 8q24. Nat Genet 39, 989–994 (2007). https://doi.org/10.1038/ng2089
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng2089
This article is cited by
-
Identification and functional validation of HLA-C as a potential gene involved in colorectal cancer in the Korean population
BMC Genomics (2022)
-
Interactions between folate intake and genetic predictors of gene expression levels associated with colorectal cancer risk
Scientific Reports (2022)
-
Variation rs9929218 and risk of the colorectal Cancer and adenomas: A meta-analysis
BMC Cancer (2021)
-
Evaluating the prognostic performance of a polygenic risk score for breast cancer risk stratification
BMC Cancer (2021)
-
Genetic risk factors for colorectal cancer in multiethnic Indonesians
Scientific Reports (2021)