Abstract
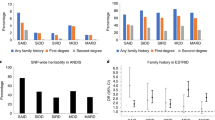
The Wellcome Trust Case Control Consortium (WTCCC) primary genome-wide association (GWA) scan1 on seven diseases, including the multifactorial autoimmune disease type 1 diabetes (T1D), shows associations at P < 5 × 10−7 between T1D and six chromosome regions: 12q24, 12q13, 16p13, 18p11, 12p13 and 4q27. Here, we attempted to validate these and six other top findings in 4,000 individuals with T1D, 5,000 controls and 2,997 family trios independent of the WTCCC study. We confirmed unequivocally the associations of 12q24, 12q13, 16p13 and 18p11 (Pfollow-up ≤ 1.35 × 10−9; Poverall ≤ 1.15 × 10−14), leaving eight regions with small effects or false-positive associations. We also obtained evidence for chromosome 18q22 (Poverall = 1.38 × 10−8) from a GWA study of nonsynonymous SNPs. Several regions, including 18q22 and 18p11, showed association with autoimmune thyroid disease. This study increases the number of T1D loci with compelling evidence from six to at least ten.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Smyth, D.J. et al. A genome-wide association study of nonsynonymous SNPs identifies a type 1 diabetes locus in the interferon-induced helicase (IFIH1) region. Nat. Genet. 38, 617–619 (2006).
Wang, W.Y., Barratt, B.J., Clayton, D.G. & Todd, J.A. Genome-wide association studies: theoretical and practical concerns. Nat. Rev. Genet. 6, 109–118 (2005).
Fisher, R.A. Correlation between relatives on the supposition of Mendelian inheritance. Trans. R. Soc. Edinb, 399–433 (1918).
Barton, N.H. & Keightley, P.D. Understanding quantitative genetic variation. Nat. Rev. Genet. 3, 11–21 (2002).
Hyttinen, V., Kaprio, J., Kinnunen, L., Koskenvuo, M. & Tuomilehto, J. Genetic liability of type 1 diabetes and the onset age among 22,650 young Finnish twin pairs: a nationwide follow-up study. Diabetes 52, 1052–1055 (2003).
Todd, J.A. A protective role of the environment in the development of type 1 diabetes? Diabet. Med. 8, 906–910 (1991).
Clayton, D.G. et al. Population structure, differential bias and genomic control in a large-scale, case-control association study. Nat. Genet. 37, 1243–1246 (2005).
Yamanouchi, J. et al. Interleukin-2 gene variation impairs regulatory T cell function and causes autoimmunity. Nat. Genet. 39, 329–337 (2007).
Todd, J.A. Statistical false positive or true disease pathway? Nat. Genet. 38, 731–733 (2006).
Chapman, J.M., Cooper, J.D., Todd, J.A. & Clayton, D.G. Detecting disease associations due to linkage disequilibrium using haplotype tags: a class of tests and the determinants of statistical power. Hum. Hered. 56, 18–31 (2003).
Lowe, C.E. et al. Cost-effective analysis of candidate genes using htSNPs: a staged approach. Genes Immun. 5, 301–305 (2004).
Ueda, H. et al. Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature 423, 506–511 (2003).
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
ten Hoeve, J. et al. Identification of a nuclear Stat1 protein tyrosine phosphatase. Mol. Cell. Biol. 22, 5662–5668 (2002).
Zelensky, A.N. & Gready, J.E. The C-type lectin-like domain superfamily. FEBS J. 272, 6179–6217 (2005).
Jones, R.B., Gordus, A., Krall, J.A. & MacBeath, G. A quantitative protein interaction network for the ErbB receptors using protein microarrays. Nature 439, 168–174 (2006).
Vella, A. et al. Localization of a type 1 diabetes locus in the IL2RA/CD25 region by use of tag single-nucleotide polymorphisms. Am. J. Hum. Genet. 76, 773–779 (2005).
Brand, O.J. et al. Association of the interleukin-2 receptor alpha (IL-2Ra)/CD25 gene region with Graves' disease using a multilocus test and tag SNPs. Clin. Endocrinol. 66, 508–512 (2007).
Bottini, N., Vang, T., Cucca, F. & Mustelin, T. Role of PTPN22 in type 1 diabetes and other autoimmune diseases. Semin. Immunol. 18, 207–213 (2006).
Smyth, D. et al. Replication of an association between the lymphoid tyrosine phosphatase locus (LYP/PTPN22) with type 1 diabetes, and evidence for its role as a general autoimmunity locus. Diabetes 53, 3020–3023 (2004).
Dardalhon, V. et al. CD226 is specifically expressed on the surface of Th1 cells and regulates their expansion and effector functions. J. Immunol. 175, 1558–1565 (2005).
Kato, H. et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441, 101–105 (2006).
Bersaglieri, T. et al. Genetic signatures of strong recent positive selection at the lactase gene. Am. J. Hum. Genet. 74, 1111–1120 (2004).
Brassat, D. et al. Multifactor dimensionality reduction reveals gene-gene interactions associated with multiple sclerosis susceptibility in African Americans. Genes Immun. 7, 310–315 (2006).
Clayton, D., Leung, H. & An, R. Package for analysis of whole-genome association studies. Hum. Hered. 64, 45–51 (2007).
Plagnol, V., Cooper, J.D., Todd, J.A. & Clayton, D.G. A method to address differential bias in genotyping in large scale association studies. PLoS Genet., published online 5 April 2007 (doi:10.1371/journal.pgen.0030074.eor).
Cordell, H.J. & Clayton, D.G. A unified stepwise regression procedure for evaluating the relative effects of polymorphisms within a gene using case/control or family data: application to HLA in type 1 diabetes. Am. J. Hum. Genet. 70, 124–141 (2002).
Hulbert, E.M. et al. T1DBase: integration and presentation of complex data for type 1 diabetes research. Nucleic Acids Res. 35, D742–D746 (2007).
Hardenbol, P. et al. Highly multiplexed molecular inversion probe genotyping: over 10,000 targeted SNPs genotyped in a single tube assay. Genome Res. 15, 269–275 (2005).
Acknowledgements
This work was funded by the Juvenile Diabetes Research Foundation International and the Wellcome Trust. We gratefully acknowledge the participation of all the patients, control subjects and family members and thank the Human Biological Data Interchange and Diabetes UK for the USA and UK multiplex families, respectively, the Norwegian Study Group for Childhood Diabetes for the collection of Norwegian families (D. Undlien and K. Rønningen), D. Savage, C. Patterson, D. Carson and P. Maxwell for the Northern Irish samples. GET1FIN (J. Tuomilehto, L. Kinnunen, E. Tuomilehto-Wolf, V. Harjutsalo and T. Valle) thank the Academy of Finland, the Sigrid Juselius Foundation and the JDRF for funding. We acknowledge use of the DNA from the 1958 British Birth Cohort collection, funded by the Medical Research Council and Wellcome Trust, and we thank D. Strachan and P. Burton for their help. We also thank The Avon Longitudinal Study of Parents and Children laboratory in Bristol, including S. Ring, R. Jones, M. Pembrey and W. McArdle for preparing and providing the control DNA samples. We thank colleagues at Affymetrix for help and advice in genotyping and T. Willis, M. Faham and P. Hardenbol for the molecular inversion probe technology. We thank the Wellcome Trust for funding the AITD UK national collection; all doctors and nurses in Birmingham, Bournemouth, Cambridge, Cardiff, Exeter, Leeds, Newcastle and Sheffield for recruitment of patients and J. Franklyn, S. Pearce (Newcastle) and P. Newby (Birmingham) for preparing and providing DNA samples on Graves' disease patients. We thank V. Everett, G. Scholz and G. Dolman for information technology support. T1D DNA samples were prepared by K. Bourget, S. Duley, M. Hardy, S. Hawkins, S. Hood, E. King, T. Mistry, A. Simpson, S. Wood, P. Lauder, S. Clayton, F. Wright and C. Collins. We thank L. Peterson for helpful discussions. C.W. is supported by the British Heart Foundation. S. Nejentsev is a Diabetes Research and Wellness Foundation Non-Clinical Fellow.
Author information
Authors and Affiliations
Consortia
Contributions
Members of the WTCCC are listed in the Supplementary Note. J.A.T. participated in the conception, design and coordination of the study; data analysis and drafting of the manuscript. N.M.W. managed the data and helped coordinate the study. J.D.C. analyzed data and drafted the manuscript. D.J.S. genotyped the nsSNP study and contributed to follow-up genotyping of the nsSNP and WTCCC studies, sequencing and genotyping of IL2 and SOCS1, data analysis and drafting of the manuscript. K.D. contributed to follow-up genotyping of the nsSNP and WTCCC studies, sequencing and genotyping of PTPN2 and data analysis. V.P. developed the nsSNP scoring algorithm and contributed to data analysis. R.B. genotyped the nsSNP study and contributed to follow-up genotyping of the nsSNP study. S. Nejentsev sequenced KIAA0350 and contributed to its bioinformatics analysis, participated in the follow-up genotyping of the WTCCC study and genotyped and analyzed 12q24 SNPs. S.F.F. genotyped the nsSNP study and contributed to follow-up genotyping of the nsSNP study. F.P. sequenced and genotyped CIITA. C.E.L. sequenced and genotyped IL21. J.S.S. genotyped and analyzed the gvSNPs. J.P.H. genotyped CD226 SNPs. L.Z. contributed to follow-up genotyping of the WTCCC scan and bioinformatics analysis. J.H.M.Y. contributed to follow-up genotyping of the WTCCC study. A.V. genotyped the IL2RB tag SNPs. S. Nutland, H.E.S., H.S., G.C., M.M. and W.M. were responsible for the DNA. L.J.S., B.H., O.S.B. and A.A.C.L. provided bioinformatics support. N.R.O. managed subject exclusions and SNP exclusions and the database for the nsSNP study. J.A. and E.A. provided T1DBase support. H.-T.L. and C.W. produced Supplementary Figure 1 and provided statistical support. J.M.M.H. performed statistical analysis; C.G. and C.I.-T. collected the Romanian families; J. Tuomilehto, L. Kinnunen, E. Tuomilehto-Wolf, V. Harjutsalo and T. Valle of GET1FIN collected the Finnish families; M.J.S., J.M.H. and S.C.L.G. provided the Graves' disease cases and genotyping of rs1990760; WTCCC carried out the 500,000-SNP GWA study; D.B.D collected the T1D cases; L.S.W. discovered the CD226 nsSNP splice sequence alterations and contributed to the overall planning of the study and D.G.C. participated in the conception, design and coordination of the study; data analysis and drafting of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Association localization plots. (PDF 906 kb)
Supplementary Fig. 2
Cross-species alignment of KIAA0350. (PDF 399 kb)
Supplementary Table 1
The locus-specific sibling recurrence-risk ratio of type 1 diabetes susceptibility variants. (PDF 125 kb)
Supplementary Table 2
Type 1 diabetes association analyses of SNPs chosen for follow-up from the WTCCC and nsSNP studies in case-controls and families. (PDF 122 kb)
Supplementary Table 3
Further genotyping from the 18p11 and 12q24 regions. (PDF 33 kb)
Supplementary Table 4
A summary of geographically variable nsSNPs and association analyses in type 1 diabetes. (PDF 58 kb)
Supplementary Table 5
Graves' disease case-control association analyses. (PDF 67 kb)
Supplementary Table 6
DNA collections and references. (PDF 24 kb)
Supplementary Table 7
Concordance between the two GWA studies and TaqMan genotyping. (PDF 35 kb)
Rights and permissions
About this article
Cite this article
Todd, J., Walker, N., Cooper, J. et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat Genet 39, 857–864 (2007). https://doi.org/10.1038/ng2068
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng2068