Abstract
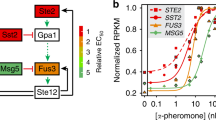
Cells must respond specifically to different environmental stimuli in order to survive. The signal transduction pathways involved in sensing these stimuli often share the same or homologous proteins. Despite potential cross-wiring, cells show specificity of response. We show, through modeling, that the physiological response of such pathways exposed to simultaneous and temporally ordered inputs can demonstrate system-level mechanisms by which pathways achieve specificity. We apply these results to the hyperosmolar and pheromone mitogen-activated protein (MAP) kinase pathways in the yeast Saccharomyces cerevisiae. These two pathways specifically sense osmolar and pheromone signals1,2,3, despite sharing a MAPKKK, Ste11, and having homologous MAPKs (Fus3 and Hog1). We show that in a single cell, the pathways are bistable over a range of inputs, and the cell responds to only one stimulus even when exposed to both. Our results imply that these pathways achieve specificity by filtering out spurious cross-talk through mutual inhibition. The variability between cells allows for heterogeneity of the decisions.
NOTE: In the version of this article initially published,the strain referred to as FUS3D63S on pp.411-412 of the main text and in the figure legend for Figure 5c-f should instead read 5c-f should instead read 5c-f FUS3D317G.The error has been corrected in the PDF version of the article.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
14 March 2007
NOTE: In the version of this article initially published,the strain referred to as FUS3D63S on pp.411-412 of the main text and in the figure legend for Figure 5c-f should instead read 5c-f should instead read 5c-f FUS3D317G.The error has been corrected in the PDF version of the article.
References
Hunter, T. & Plowman, G.D. The protein kinases of budding yeast: six score and more. Trends Biochem. Sci. 22, 18–22 (1997).
Posas, F., Takekawa, M. & Saito, H. Signal transduction by MAP kinase cascades in budding yeast. Curr. Opin. Microbiol. 1, 175–182 (1998).
Banuett, F. Signalling in the yeasts: an informational cascade with links to the filamentous fungi. Microbiol. Mol. Biol. Rev. 62, 249–274 (1998).
Whitmarsh, A.J. & Davis, R.J. Structural organization of MAP-kinase signaling modules by scaffold proteins in yeast and mammals. Trends Biochem. Sci. 23, 481–485 (1998).
Ferrell, J.E. Jr . & Machleder, E.M. The biochemical basis of an all-or-none switch in Xenopus oocytes. Science 280, 895–898 (1998).
Schwartz, M.A. & Madhani, H.D. Principles of MAP kinase signaling specificity in Saccharomyces cerevisiae. Annu. Rev. Genet. 38, 725–748 (2004).
Sprague, G.F. Jr . Control of MAP kinase signaling specificity or how not to go HOG wild. Genes Dev. 12, 2817–2820 (1998).
Pryciak, P.M. & Huntress, F.A. Membrane recruitment of the kinase cascade scaffold protein Ste5 by the Gbetagamma complex underlies activation of the yeast pheromone response pathway. Genes Dev. 12, 2684–2697 (1998).
Choi, K.Y., Satterberg, B., Lyons, D.M. & Elion, E.A. Ste5 tethers multiple protein kinases in the MAP kinase cascade required for mating in S. cerevisiae. Cell 78, 499–512 (1994).
Posas, F. & Saito, H. Osmotic activation of the HOG MAPK pathway via Ste11p MAPKKK: scaffold role of Pbs2p MAPKK. Science 276, 1702–1705 (1997).
Harris, K. et al. Role of scaffolds in MAP kinase pathway specificity revealed by custom design of pathway-dedicated signaling proteins. Curr. Biol. 11, 1815–1824 (2001).
O'Rourke, S.M. & Herskowitz, I. The Hog1 MAPK prevents cross talk between the HOG and pheromone response MAPK pathways in Saccharomyces cerevisiae. Genes Dev. 12, 2874–2886 (1998).
Posas, F. & Saito, H. Activation of the yeast SSK2 MAP kinase kinase kinase by the SSK1 two-component response regulator. EMBO J. 17, 1385–1394 (1998).
Posas, F. et al. Yeast HOG1 MAP kinase cascade is regulated by a multistep phosphorelay mechanism in the SLN1–YPD1-SSK1 “two-component” osmosensor. Cell 86, 865–875 (1996).
O'Rourke, S.M. & Herskowitz, I. Unique and redundant roles for HOG MAPK pathway components as revealed by whole-genome expression analysis. Mol. Biol. Cell 15, 532–542 (2004).
Breitkreutz, A. & Tyers, M. MAPK signaling specificity: it takes two to tango. Trends Cell Biol. 12, 254–257 (2002).
Tsien, R.Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Hagen, D.C., McCaffrey, G. & Sprague, G.F. Jr . Pheromone response elements are necessary and sufficient for basal and pheromone-induced transcription of the FUS1 gene of Saccharomyces cerevisiae. Mol. Cell. Biol. 11, 2952–2961 (1991).
Campbell, R.E. et al. A monomeric red fluorescent protein. Proc. Natl. Acad. Sci. USA 99, 7877–7882 (2002).
Rep, M., Krantz, M., Thevelein, J.M. & Hohmann, S. The transcriptional response of Saccharomyces cerevisiae to osmotic shock. Hot1p and Msn2p/Msn4p are required for the induction of subsets of high osmolarity glycerol pathway-dependent genes. J. Biol. Chem. 275, 8290–8300 (2000).
Murray, J.D. Mathematical Biology (Springer, New York, 2002).
Hall, J.P. et al. The osmoregulatory pathway represses mating pathway activity in Saccharomyces cerevisiae: isolation of a FUS3 mutant that is insensitive to the repression mechanism. Mol. Cell. Biol. 16, 6715–6723 (1996).
Xiong, W. & Ferrell, J.E. Jr . A positive-feedback-based bistable 'memory module' that governs cell fate decision. Nature 426, 460–465 (2003).
Zhu, H. et al. Analysis of yeast protein kinases using protein chips. Nat. Genet. 26, 283–289 (2000).
Philips, J. & Herskowitz, I. Osmotic balance regulates cell fusion during mating in Saccharomyces cerevisiae. J. Cell Biol. 138, 961–974 (1997).
Gimeno, C.J., Ljungdahl, P.O., Styles, C.A. & Fink, G.R. Unipolar cell divisions in the yeast S. cerevisiae lead to filamentous growth: regulation by starvation and RAS. Cell 68, 1077–1090 (1992).
Bhattacharyya, R.P. et al. The Ste5 scaffold allosterically modulates signaling output of the yeast mating pathway. Science 311, 822–826 (2006).
Acknowledgements
We thank J. Weiner, P. Houston, K. Thorn, K. Duevel, L. Schneper, E. Xu and P. Hersen for help with experiments, R. Tsien and E. Winters for reagents, A. Sengupta, A. Murray and M. Tyers for helpful discussions and A. Regev, L. Garwin, K. Vestrepen, P. Swain, E. O'Shea, I. Nachman, N. Barkai and A. Amon for comments on the manuscript. This work was supported by grants from the NIH (J.R.B.), GRPW fellowship, Lucent Technologies (M.N.M.), Keck Futures Initiative (S.R.) and the FAS Center for Systems Biology (S.R. and M.N.M.). Requests for materials should be addressed to S.R. (sharadr@alcatel-lucent.com).
Author information
Authors and Affiliations
Contributions
S.R., J.R.B. and M.M. designed the experiments; M.M. and A.M. did the modeling and J.R.B., M.M., A.M. and S.R. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
The analysis of models. (PDF 887 kb)
Supplementary Fig. 2
Results from modeling. (PDF 525 kb)
Supplementary Fig. 3
Cell-to-cell variability. (PDF 732 kb)
Supplementary Fig. 4
Controls. (PDF 220 kb)
Supplementary Fig. 5
Protein blots. (PDF 290 kb)
Supplementary Fig. 6
Filamentous growth and pheromone response pathways and model. (PDF 494 kb)
Supplementary Fig. 7
Phase plot of pheromone and filamentous response as a function of the inputs. (PDF 384 kb)
Rights and permissions
About this article
Cite this article
McClean, M., Mody, A., Broach, J. et al. Cross-talk and decision making in MAP kinase pathways. Nat Genet 39, 409–414 (2007). https://doi.org/10.1038/ng1957
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1957
This article is cited by
-
Neural Differentiation of Mouse Neural Stem Cells as a Tool to Assess Developmental Neurotoxicity of Drinking Water in Taihu Lake
Biological Trace Element Research (2019)
-
Crosstalk and the evolvability of intracellular communication
Nature Communications (2017)
-
Negative Interactions and Feedback Regulations Are Required for Transient Cellular Response
Scientific Reports (2014)
-
Activation of stress signalling pathways enhances tolerance of fungi to chemical fungicides and antifungal proteins
Cellular and Molecular Life Sciences (2014)
-
Arabidopsis MAP kinase phosphatase 1 is phosphorylated and activated by its substrate AtMPK6
Plant Cell Reports (2011)