Abstract
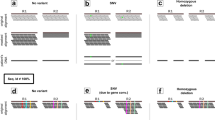
Structural and insertion-deletion (indel) variants have received considerable recent attention, partly because of their phenotypic consequences. Among these variants, the most common are small indels (∼1–30 bp). Identifying and genotyping indels using sequence traces obtained from diploid samples requires extensive manual review, which makes large-scale studies inconvenient. We report a new algorithm, implemented in available software (PolyPhred version 6.0), to help automate detection and genotyping of indels from sequence traces. The algorithm identifies heterozygous individuals, which permits the discovery of low-frequency indels. It finds 80% of all indel polymorphisms with almost no false positives and finds 97% with a false discovery rate of 10%. Additionally, genotyping accuracy exceeds 99%, and it correctly infers indel length in 96% of the cases. Using this approach, we identify indels in the HapMap ENCODE regions, providing the first report of these polymorphisms in this data set.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Sebat, J. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004).
Tuzun, E. et al. Fine-scale structural variation of the human genome. Nat. Genet. 37, 727–732 (2005).
Albertini, A.M., Hofer, M., Calos, M.P. & Miller, J.H. On the formation of spontaneous deletions: the importance of short sequence homologies in the generation of large deletions. Cell 29, 319–328 (1982).
Conrad, D.F., Andrews, T.D., Carter, N.P., Hurles, M.E. & Pritchard, J.K. A high-resolution survey of deletion polymorphism in the human genome. Nat. Genet. 38, 75–81 (2006).
Hinds, D.A., Kloek, A.P., Jen, M., Chen, X. & Frazer, K.A. Common deletions and SNPs are in linkage disequilibrium in the human genome. Nat. Genet. 38, 82–85 (2006).
McCarroll, S.A. et al. Common deletion polymorphisms in the human genome. Nat. Genet. 38, 86–92 (2006).
Bhangale, T.R., Rieder, M.J., Livingston, R.J. & Nickerson, D.A. Comprehensive identification and characterization of diallelic insertion-deletion polymorphisms in 330 human candidate genes. Hum. Mol. Genet. 14, 59–69 (2005).
Othman, M. et al. Identification and functional characterization of a novel 27-bp deletion in the macroglycopeptide-coding region of the GPIBA gene resulting in platelet-type von Willebrand disease. Blood 105, 4330–4336 (2005).
deSanctis, L. et al. Familial PAX8 small deletion (c.989_992delACCC) associated with extreme phenotype variability. J. Clin. Endocrinol. Metab. 89, 5669–5674 (2004).
Karban, A.S. et al. Functional annotation of a novel NFKB1 promoter polymorphism that increases risk for ulcerative colitis. Hum. Mol. Genet. 13, 35–45 (2004).
Lin, S.C. et al. Correlation between functional genotypes in the matrix metalloproteinases-1 promoter and risk of oral squamous cell carcinomas. J. Oral Pathol. Med. 33, 323–326 (2004).
Stenson, P.D. et al. Human Gene Mutation Database (HGMD): 2003 update. Hum. Mutat. 21, 577–581 (2003).
Nickerson, D.A., Tobe, V.O. & Taylor, S.L. PolyPhred: automating the detection and genotyping of single nucleotide substitutions using fluorescence-based resequencing. Nucleic Acids Res. 25, 2745–2751 (1997).
Stephens, M., Sloan, J.S., Robertson, P.D., Scheet, P. & Nickerson, D.A. Automating sequence-based detection and genotyping of SNPs from diploid samples. Nat. Genet. 38, 375–381 (2006).
Weckx, S. et al. novoSNP, a novel computational tool for sequence variation discovery. Genome Res. 15, 436–442 (2005).
Carlson, C.S. et al. Additional SNPs and linkage-disequilibrium analyses are necessary for whole-genome association studies in humans. Nat. Genet. 33, 518–521 (2003).
Livingston, R.J. et al. Pattern of sequence variation across 213 environmental response genes. Genome Res. 14, 1821–1831 (2004).
International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Manaster, C. et al. InSNP: a tool for automated detection and visualization of SNPs and InDels. Hum. Mutat. 26, 11–19 (2005).
Ewing, B., Hillier, L., Wendl, M.C. & Green, P. Base-calling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res. 8, 175–185 (1998).
Locke, D.P. et al. Linkage disequilibrium and heritability of copy-number polymorphisms within duplicated regions of the human genome. Am. J. Hum. Genet. 79, 275–290 (2006).
Newman, T.L. et al. High-throughput genotyping of intermediate-size structural variation. Hum. Mol. Genet. 15, 1159–1167 (2006).
Klein, R.J. et al. Complement factor H polymorphism in age-related macular degeneration. Science 308, 385–389 (2005).
Ahn, J. et al. Cloning of the putative tumour suppressor gene for hereditary multiple exostoses (EXT1). Nat. Genet. 11, 137–143 (1995).
Rockman, M.V. et al. Positive selection on MMP3 regulation has shaped heart disease risk. Curr. Biol. 14, 1531–1539 (2004).
Eichler, E.E. Widening the spectrum of human genetic variation. Nat. Genet. 38, 9–11 (2006).
Weber, J.L. et al. Human diallelic insertion/deletion polymorphisms. Am. J. Hum. Genet. 71, 854–862 (2002).
Mills, R.E. et al. An initial map of insertion and deletion (INDEL) variation in the human genome. Genome Res. 16, 1182–1190 (2006).
Kruglyak, L. & Nickerson, D.A. Variation is the spice of life. Nat. Genet. 27, 234–236 (2001).
Ewing, B. & Green, P. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8, 186–194 (1998).
Gordon, D., Abajian, C. & Green, P. Consed: a graphical tool for sequence finishing. Genome Res. 8, 195–202 (1998).
Needleman, S.B. & Wunsch, C.D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J. Mol. Biol. 48, 443–453 (1970).
Acknowledgements
The authors thank the past and present members of the SeattleSNPs team for their efforts in variation discovery and the PolyPhred development team, including J. Sloan and P. Robertson. This work was supported by grants from the US National Institute of Health (HL66682 to D.A.N. and HG/LM02585 to M.S.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
PolyPhred is freely available for academic purposes, but a licensing fee is charged for commercial use, which predominantly funds further software and methods development.
Supplementary information
Supplementary Fig. 1
Trace signal patterns. (PDF 76 kb)
Supplementary Fig. 2
Lengths and LD characteristics of ENCODE indels. (PDF 285 kb)
Supplementary Fig. 3
Application of the DPA. (PDF 225 kb)
Supplementary Table 1
ENCODE indels found in different functional regions of genes. (PDF 49 kb)
Supplementary Table 2
Chromosomal locations of the 1,125 indels identified in the ENCODE regions. (PDF 200 kb)
Supplementary Table 3
The independent variables and their estimated effect parameters in the logistic regression model used for identifying heterozygous indel traces. (PDF 86 kb)
Supplementary Table 4
The independent variables and their estimated effect parameters in the logistic regression model used for identifying indel loci. (PDF 83 kb)
Rights and permissions
About this article
Cite this article
Bhangale, T., Stephens, M. & Nickerson, D. Automating resequencing-based detection of insertion-deletion polymorphisms. Nat Genet 38, 1457–1462 (2006). https://doi.org/10.1038/ng1925
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1925
This article is cited by
-
Development of Novel High-Resolution Melting-Based Assays for Genotyping Two Alu Insertion Polymorphisms (FXIIIB and PV92)
Molecular Biotechnology (2016)
-
DiSNPindel: improved intra-individual SNP and InDel detection in direct amplicon sequencing of a diploid
BMC Bioinformatics (2015)
-
Sequencing of Lp-PLA2-encoding PLA2G7 gene in 2000 Europeans reveals several rare loss-of-function mutations
The Pharmacogenomics Journal (2012)
-
The conserved WW-domain binding sites in Dystroglycan C-terminus are essential but partially redundant for Dystroglycan function
BMC Developmental Biology (2009)
-
Analysis of the tyrosine kinome in melanoma reveals recurrent mutations in ERBB4
Nature Genetics (2009)