Abstract
The proteins encoded by the classical HLA class I and class II genes in the major histocompatibility complex (MHC) are highly polymorphic and are essential in self versus non-self immune recognition. HLA variation is a crucial determinant of transplant rejection and susceptibility to a large number of infectious and autoimmune diseases1. Yet identification of causal variants is problematic owing to linkage disequilibrium that extends across multiple HLA and non-HLA genes in the MHC2,3. We therefore set out to characterize the linkage disequilibrium patterns between the highly polymorphic HLA genes and background variation by typing the classical HLA genes and >7,500 common SNPs and deletion-insertion polymorphisms across four population samples. The analysis provides informative tag SNPs that capture much of the common variation in the MHC region and that could be used in disease association studies, and it provides new insight into the evolutionary dynamics and ancestral origins of the HLA loci and their haplotypes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dupont, B. & Svejgaard, A. HLA and disease. Transplant. Proc. 9, 1271–1274 (1977).
Miretti, M.M. et al. A high-resolution linkage-disequilibrium map of the human major histocompatibility complex and first generation of tag single-nucleotide polymorphisms. Am. J. Hum. Genet. 76, 634–646 (2005).
Walsh, E.C. et al. An integrated haplotype map of the human major histocompatibility complex. Am. J. Hum. Genet. 73, 580–590 (2003).
Allcock, R.J. et al. The MHC haplotype project: a resource for HLA-linked association studies. Tissue Antigens 59, 520–521 (2002).
Horton, R. et al. Gene map of the extended human MHC. Nat. Rev. Genet. 5, 889–899 (2004).
Stewart, C.A. et al. Complete MHC haplotype sequencing for common disease gene mapping. Genome Res. 14, 1176–1187 (2004).
Malkki, M., Single, R., Carrington, M., Thomson, G. & Petersdorf, E. MHC microsatellite diversity and linkage disequilibrium among common HLA-A, HLA-B, DRB1 haplotypes: implications for unrelated donor hematopoietic transplantation and disease association studies. Tissue Antigens 66, 114–124 (2005).
Stenzel, A. et al. Patterns of linkage disequilibrium in the MHC region on human chromosome 6p. Hum. Genet. 114, 377–385 (2004).
Limm, T.M., Ashdown, M.L., Naughton, M.J., McGinnis, M.D. & Simons, M.J. HLA-DQA1 allele and suballele typing using noncoding sequence polymorphisms. Application to 4AOHW cell panel typing. Hum. Immunol. 38, 57–68 (1993).
Simons, M.J. et al. Strategy for definition of DR/DQ haplotypes in the 4AOHW cell panel using noncoding sequence polymorphisms. Hum. Immunol. 38, 69–74 (1993).
de Bakker, P.I.W. et al. Efficiency and power in genetic association studies. Nat. Genet. 37, 1217–1223 (2005).
Gonzalez-Neira, A. et al. The portability of tagSNPs across populations: a worldwide survey. Genome Res. 16, 323–330 (2006).
Monsuur, A.J. et al. Myosin IXB variant increases the risk of celiac disease and points toward a primary intestinal barrier defect. Nat. Genet. 37, 1341–1344 (2005).
Chadha, S. et al. Haplotype analysis of tumour necrosis factor receptor genes in 1p36: no evidence for association with systemic lupus erythematosus. Eur. J. Hum. Genet. 14, 69–78 (2006).
Vader, W. et al. The HLA-DQ2 gene dose effect in celiac disease is directly related to the magnitude and breadth of gluten-specific T cell responses. Proc. Natl. Acad. Sci. USA 100, 12390–12395 (2003).
Graham, R.R. et al. Visualizing human leukocyte antigen class II risk haplotypes in human systemic lupus erythematosus. Am. J. Hum. Genet. 71, 543–553 (2002).
Marsh, S.G. Nomenclature for factors of the HLA system, update June 2005. Tissue Antigens 66, 338–340 (2005).
Klein, J. Origin of major histocompatibility complex polymorphism: the trans-species hypothesis. Hum. Immunol. 19, 155–162 (1987).
Raymond, C.K. et al. Ancient haplotypes of the HLA Class II region. Genome Res. 15, 1250–1257 (2005).
Traherne, J.A. et al. Genetic analysis of completely sequenced disease-associated mhc haplotypes identifies shuffling of segments in recent human history. PLoS Genet 2, e9 (2006).
Froeschke, G. & Sommer, S. MHC class II DRB variability and parasite load in the striped mouse (Rhabdomys pumilio) in the Southern Kalahari. Mol. Biol. Evol. 22, 1254–1259 (2005).
Sabeti, P.C. et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002).
Hill, W.G.R.A. Linkage disequilibrium in finite populations. Theor. Appl. Genet. 38, 226–231 (1968).
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
McVean, G.A. et al. The fine-scale structure of recombination rate variation in the human genome. Science 304, 581–584 (2004).
Myers, S., Bottolo, L., Freeman, C., McVean, G. & Donnelly, P. A fine-scale map of recombination rates and hotspots across the human genome. Science 310, 321–324 (2005).
Nei, M. (ed.). Molecular Evolutionary Genetics (Columbia Univ. Press, New York, 1987).
Voight, B.F., Kudaravalli, S., Wen, X. & Pritchard, J.K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Fry, B. Computational Information Design. Thesis, Massachusetts Institute of Technology, (2005).
Acknowledgements
The authors thank J. Oksenberg, P. De Jager and N. Walker for discussions and their critical reading of the manuscript. The authors are also grateful to B. Fry for technical assistance with the selection analysis. This project has been funded in whole or in part with federal funds from the US National Cancer Institute, National Institutes of Health (NIH), under contract N01-CO-12400. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government. This research was supported in part by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research. The Wellcome Trust supported the work of M.M., P.W., M.D., J.M., S.B., J.T., J.A.T. and P.D. The Juvenile Diabetes Research Foundation supported J.A.T. P.C.S. is funded by the Damon Runyon Cancer Fellowship. The International MS Genetics Consortium supported the work of D.H., S.G., M.P.V., and J.D.R. This work was also supported by grants from the National Institute of Diabetes and Digestive and Kidney Diseases and the National Institute of Allergy and Infectious Diseases (Autoimmunity Prevention Center grant U19 AI050864) to J.D.R.
Author information
Authors and Affiliations
Contributions
The study was designed by J.A.T., S.G., S.B., P.D. and J.D.R. Genotyping was performed by multiple groups (P.W., M.D., J.M., A.R., L.G., J.H., M.P.-V., S.S.M.). M.C. performed the HLA typing. A.J.M., C.W. and T.V. provided samples and genotype data for the cross-validation experiments. P.I.W.d.B., G.M., P.C.S., M.M., J.M., X.K., E.C.W. and T.G. performed analyses. The manuscript was written by P.I.W.d.B., G.M., P.C.S. and J.D.R., with contributions from J.A.T., D.A.H., M.J.D., M.C. and J.T. The genotyping, analysis and manuscript writing efforts of this international collaborative group were coordinated by J.D.R.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
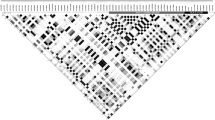
Supplementary Fig. 1
Allelic association between SNPs across the 7.5-Mb extended MHC region and HLA types at each gene for the combined population data (using the 5,754 SNPs that were typed in all populations and are polymorphic across the combined population samples). (PDF 2374 kb)
Supplementary Fig. 2
A region of the MHC, including BAK1 and HLA-DPA1, contains one of the top 20 candidates for selection based on the long-range haplotype test in the YRI. (PDF 313 kb)
Supplementary Table 1
Partial summary of established HLA associations and associations of contemporary interest. (PDF 16 kb)
Supplementary Table 2
Correlations between alleles at the six classical HLA loci typed in the study. (PDF 4 kb)
Supplementary Table 3
List of tags for HLA alleles. (PDF 70 kb)
Supplementary Table 4
Cross-panel performance of HLA tags. (PDF 65 kb)
Supplementary Table 5
List of top-ranking SNPs and haplotypes with evidence for recent positive selection using EHH-based methods. (PDF 74 kb)
Rights and permissions
About this article
Cite this article
de Bakker, P., McVean, G., Sabeti, P. et al. A high-resolution HLA and SNP haplotype map for disease association studies in the extended human MHC. Nat Genet 38, 1166–1172 (2006). https://doi.org/10.1038/ng1885
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1885
This article is cited by
-
Phenome-wide association study on miRNA-related sequence variants: the UK Biobank
Human Genomics (2023)
-
Comparison between qPCR and RNA-seq reveals challenges of quantifying HLA expression
Immunogenetics (2023)
-
Genetic variation in the human leukocyte antigen region confers susceptibility to Clostridioides difficile infection
Scientific Reports (2023)
-
HLA-SPREAD: a natural language processing based resource for curating HLA association from PubMed abstracts
BMC Genomics (2022)
-
HLA imputation and its application to genetic and molecular fine-mapping of the MHC region in autoimmune diseases
Seminars in Immunopathology (2022)