Abstract
Auditory neuropathy is a particular type of hearing impairment in which neural transmission of the auditory signal is impaired, while cochlear outer hair cells remain functional. Here we report on DFNB59, a newly identified gene on chromosome 2q31.1–q31.3 mutated in four families segregating autosomal recessive auditory neuropathy. DFNB59 encodes pejvakin, a 352-residue protein. Pejvakin is a paralog of DFNA5, a protein of unknown function also involved in deafness. By immunohistofluorescence, pejvakin is detected in the cell bodies of neurons of the afferent auditory pathway. Furthermore, Dfnb59 knock-in mice, homozygous for the R183W variant identified in one DFNB59 family, show abnormal auditory brainstem responses indicative of neuronal dysfunction along the auditory pathway. Unlike previously described sensorineural deafness genes, all of which underlie cochlear cell pathologies, DFNB59 is the first human gene implicated in nonsyndromic deafness due to a neuronal defect.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Liberman, M.C. Single-neuron labeling in the cat auditory nerve. Science 216, 1239–1241 (1982).
Rubel, E.W. & Fritzsch, B. Auditory system development: primary auditory neurons and their targets. Annu. Rev. Neurosci. 25, 51–101 (2002).
Rouiller, E.M. Functional organization of the auditory pathways. in The Central Auditory System (eds. Ehret, G. & Romand, R.) 3–96 (Oxford Univ. Press, Oxford, 1997).
Romand, R. Modification of tonotopic representation in the auditory system during development. Prog. Neurobiol. 51, 1–17 (1997).
Sininger, Y.S. Identification of auditory neuropathy in infants and children. Semin. Hear. 23, 193–200 (2002).
Dirks, D.D., Morgan, D.E. & Ruth, R.A. Auditory brainstem response and electrocochleographic testing. in The Ear: Comprehensive Otology (eds. Canalis, R.F. & Lambert, P.R.) 231–241 (Lippincott Williams & Wilkins, Philadelphia, 2000).
Kemp, D.T. Otoacoustic emissions, their origin in cochlear function, and use. Br. Med. Bull. 63, 223–241 (2002).
Starr, A., Picton, T.W., Sininger, Y.S., Hood, L.J. & Berlin, C.I. Auditory neuropathy. Brain 119, 741–753 (1996).
Cone-Wesson, B. & Rance, G. Auditory neuropathy: a brief review. Curr. Opin. Otolaryngol. Head Neck Surg. 8, 421–425 (2000).
Starr, A., Sininger, Y.S. & Pratt, H. The varieties of auditory neuropathy. J. Basic Clin. Physiol. Pharmacol. 11, 215–230 (2000).
Pulleyn, L.J. et al. A new locus for autosomal recessive non-syndromal sensorineural hearing impairment (DFNB27) on chromosome 2q23-q31. Eur. J. Hum. Genet. 8, 991–993 (2000).
Van Laer, L. et al. Nonsyndromic hearing impairment is associated with a mutation in DFNA5 . Nat. Genet. 20, 194–197 (1998).
Katoh, M. & Katoh, M. Identification and characterization of human DFNA5L, mouse Dfna5l, and rat Dfna5l genes in silico. Int. J. Oncol. 25, 765–770 (2004).
Runkel, F. et al. The dominant alopecia phenotypes Bareskin, Rex-denuded, and Reduced Coat 2 are caused by mutations in gasdermin 3 . Genomics 84, 824–835 (2004).
Lunny, D.P. et al. Mutations in gasdermin 3 cause aberrant differentiation of the hair follicle and sebaceous gland. J. Invest. Dermatol. 124, 615–621 (2005).
Berezin, C. et al. CONSEQ: the identification of functionally and structurally important residues in protein sequences. Bioinformatics 20, 1322–1324 (2004).
Zheng, Q.Y., Johnson, K.R. & Erway, L.C. Assessment of hearing in 80 inbred strains of mice by ABR threshold analyses. Hear. Res. 130, 94–107 (1999).
Van Laer, L. et al. Mice lacking Dfna5 show a diverging number of cochlear fourth row outer hair cells. Neurobiol. Dis. 19, 386–399 (2005).
Durrant, J.D., Wang, J., Ding, D.L. & Salvi, R.J. Are inner or outer hair cells the source of summating potentials recorded from the round window? J. Acoust. Soc. Am. 104, 370–377 (1998).
Rance, G. Auditory neuropathy / dys-synchrony and its perceptual consequences. Trends Amplif. 9, 1–43 (2005).
Oertel, D. The role of timing in the brain stem auditory nuclei of vertebrates. Annu. Rev. Physiol. 61, 497–519 (1999).
Rapin, I. & Gravel, J. “Auditory neuropathy”: physiologic and pathologic evidence calls for more diagnostic specificity. Int. J. Pediatr. Otorhinolaryngol. 67, 707–728 (2003).
Yasunaga, S. et al. A mutation in OTOF, encoding otoferlin, a FER-1-like protein, causes DFNB9, a nonsyndromic form of deafness. Nat. Genet. 21, 363–369 (1999).
Varga, R. et al. Non-syndromic recessive auditory neuropathy is the result of mutations in the otoferlin (OTOF) gene. J. Med. Genet. 40, 45–50 (2003).
Rodríguez-Ballesteros, M. et al. Auditory neuropathy in patients carrying mutations in the otoferlin gene (OTOF). Hum. Mutat. 22, 451–456 (2003).
Schaffer, A.A. Faster linkage analysis computations for pedigrees with loops or unused alleles. Hum. Hered. 46, 226–235 (1996).
Cohen-Salmon, M., El-Amraoui, A., Leibovici, M. & Petit, C. Otogelin: a glycoprotein specific to the acellular membranes of the inner ear. Proc. Natl. Acad. Sci. USA 94, 14450–14455 (1997).
Letunic, I. et al. SMART 4.0: towards genomic data integration. Nucleic Acids Res. 32, D142–D144 (2004).
Cokol, M. et al. Finding nuclear localization signals. EMBO Rep. 1, 411–415 (2000).
Kent, W.J. BLAT–the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Edgar, R.C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5, 113 (2004).
Guindon, S. & Gascuel, O. A simple, fast and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003).
Lee, E.C. et al. A highly efficient Escherichia coli-based chromosome engineering system adapted for recombinogenic targeting and subcloning of BAC DNA. Genomics 73, 56–62 (2001).
Liu, P., Jenkins, N.A. & Copeland, N.G. A highly efficient recombineering-based method for generating conditional knockout mutations. Genome Res. 13, 476–484 (2003).
Kress, C., Vandormael-Pournin, S., Baldacci, P., Cohen-Tannoudji, M. & Babinet, C. Nonpermissiviness for mouse embryonic stem (ES) cell derivation circumvented by a single backcross to 129/Sv strain: establishment of ES cell lines bearing the Omd conditional lethal mutation. Mamm. Genome 9, 998–1001 (1998).
Matise, M.P., Auerbach, W. & Joyner, A. Production of targeted embryonic stem cell clones. in Gene Targeting: a Practical Approach (ed. A. Joyner) 101–132 (Oxford Univ. Press, Oxford, 1999).
Lallemand, Y., Luria, V., Haffner-Krausz, R. & Lonai, P. Maternally expressed PGK-cre transgene as a tool for early and uniform activation of the Cre site-specific recombinase. Transgenic Res. 7, 105–112 (1998).
Steel, K.P. & Hardisty, R. Assessing hearing, vision and balance in mice. in What's Wrong With My Mouse? New Interplays Between Mouse Genes and Behavior 26–38 (Society for Neuroscience, Washington D.C., 1996).
Küssel-Andermann, P. et al. Vezatin, a novel transmembrane protein, bridges myosin VIIA to the cadherin-catenins complex. EMBO J. 19, 6020–6029 (2000).
Adato, A. et al. Interactions in the network of Usher syndrome type 1 proteins. Hum. Mol. Genet. 14, 347–356 (2005).
Acknowledgements
The authors are grateful to all the families and clinicians involved in the study for their collaboration. We thank the staff at the Pasteur Institute of Iran and the Specialized Education Centers for their help in collecting subject samples; T. Hutchin and R.F. Mueller for generously sharing DNA samples from individuals belonging to the family that defined the DFNB27 locus; N.G. Copeland for gifts of bacterial strains and plasmids; S. Nouaille, S. Chardenoux and F. Thouron for technical help; P. Roux for advice on confocal microscopy; M. Cohen-Salmon and S. Safieddine for advice on immunohistofluorescence techniques; J. Levilliers and A. Hafidi for discussion and J.P. Hardelin for critical reading of the manuscript. S.D. is grateful for the support of the Letten Saugstad Foundation. F.J.d.C was a recipient of a Marie Curie postdoctoral fellowship. L.V.L. is a postdoctoral fellow of the Flemish Fonds voor Wetenschappelijk Onderzoek (FWO). This work was supported by the European Commission FP6 Integrated Project EuroHear (LSHG-CT-2004-512063) and by Fondation Louis-Jeantet.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
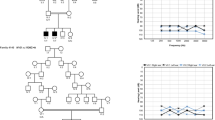
Pedigrees of DFNB59 families 700 and 710. (PDF 116 kb)
Supplementary Fig. 2
Phylogenetic tree shwoing the evolutionary relationships between pejvakin and the remaining members of the DFNA5-gasdermin-MLZE protein family. (PDF 251 kb)
Supplementary Fig. 3
Scanning electron micrographs of IHCs and OHCs at equivalent positions in the medial part of the cochlear duct of P30 Dfnb59+/+ and Dfnb59tm1 Ugds/tm1 Ugds mice. (PDF 405 kb)
Supplementary Fig. 4
Expression of pejvakin in the murine organ of Corti. (PDF 135 kb)
Supplementary Fig. 5
Examples of ABR waveforms in response to 20-kHz tone bursts at 80 dB SPL. (PDF 111 kb)
Supplementary Fig. 6
Strategy for targeted replacement of the wild-type Dfnb59 allele with Dfnb59tm1 Ugds (PDF 90 kb)
Supplementary Table 1
Sequences of primers used to amplify DFNB59 exons. (PDF 102 kb)
Rights and permissions
About this article
Cite this article
Delmaghani, S., del Castillo, F., Michel, V. et al. Mutations in the gene encoding pejvakin, a newly identified protein of the afferent auditory pathway, cause DFNB59 auditory neuropathy. Nat Genet 38, 770–778 (2006). https://doi.org/10.1038/ng1829
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1829
This article is cited by
-
A dominant variant in apoptosis-related gene XKR8 is relevant to hereditary auditory neuropathy
Journal of Translational Medicine (2023)
-
Impaired AIF-CHCHD4 interaction and mitochondrial calcium overload contribute to auditory neuropathy spectrum disorder in patient-iPSC-derived neurons with AIFM1 variant
Cell Death & Disease (2023)
-
Pore-forming proteins as drivers of membrane permeabilization in cell death pathways
Nature Reviews Molecular Cell Biology (2023)
-
Gasdermins and pyroptosis in the kidney
Nature Reviews Nephrology (2023)
-
The gasdermin protein family: emerging roles in gastrointestinal health and disease
Nature Reviews Gastroenterology & Hepatology (2023)