Abstract
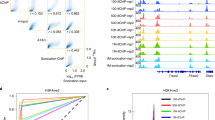
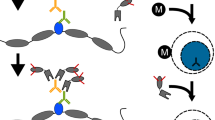
Chromatin immunoprecipitation (ChIP) defines the genomic distribution of proteins and their modifications but is limited by the cell numbers required (ideally >107). Here we describe a protocol that uses carrier chromatin and PCR, 'carrier' ChIP (CChIP), to permit analysis of as few as 100 cells. We assayed histone modifications at key regulator genes (such as Nanog, Pou5f1 (also known as Oct4) and Cdx2) by CChIP in mouse embryonic stem (ES) cells and in inner cell mass (ICM) and trophectoderm of cultured blastocysts. Activating and silencing modifications (H4 acetylation and H3K9 methylation) mark active and silent promoters as predicted, and we find close correlation between values derived from CChIP (1,000 ES cells) and conventional ChIP (5 × 107 ES cells). Studies on genes silenced in both ICM and ES cells (Cdx2, Cfc1, Hhex and Nkx2-2, also known as Nkx) show that the intensity of silencing marks is relatively diminished in ES cells, indicating a possible relaxation of some components of silencing on adaptation to culture.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Smith, A.G. Embryo-derived stem cells: of mice and men. Annu. Rev. Cell Dev. Biol. 17, 435–462 (2001).
Ralston, A. & Rossant, J. Genetic regulation of stem cell origins in the mouse embryo. Clin. Genet. 68, 106–112 (2005).
Chambers, I. et al. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113, 643–655 (2003).
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003).
Boyer, L.A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Arney, K.L. & Fisher, A.G. Epigenetic aspects of differentiation. J. Cell Sci. 117, 4355–4363 (2004).
Santos, F. & Dean, W. Epigenetic reprogramming during early development in mammals. Reproduction 127, 643–651 (2004).
Szutorisz, H. & Dillon, N. The epigenetic basis for embryonic stem cell pluripotency. Bioessays 27, 1286–1293 (2005).
Gilbert, N., Gilchrist, S. & Bickmore, W.A. Chromatin organization in the mammalian nucleus. Int. Rev. Cytol. 242, 283–336 (2005).
Merkenschlager, M. et al. Centromeric repositioning of coreceptor loci predicts their stable silencing and the CD4/CD8 lineage choice. J. Exp. Med. 200, 1437–1444 (2004).
Perry, P. et al. A dynamic switch in the replication timing of key regulator genes in embryonic stem cells upon neural induction. Cell Cycle 3, 1645–1650 (2004).
Sproul, D., Gilbert, N. & Bickmore, W.A. The role of chromatin structure in regulating the expression of clustered genes. Nat. Rev. Genet. 6, 775–781 (2005).
Wiblin, A.E., Cui, W., Clark, A.J. & Bickmore, W.A. Distinctive nuclear organisation of centromeres and regions involved in pluripotency in human embryonic stem cells. J. Cell Sci. 118, 3861–3868 (2005).
Margueron, R., Trojer, P. & Reinberg, D. The key to development: interpreting the histone code? Curr. Opin. Genet. Dev. 15, 163–176 (2005).
Nightingale, K.P., O'Neill, L.P. & Turner, B.M. Histone modifications: signalling receptors and potential elements of a heritable epigenetic code. Curr. Opin. Genet. Dev. 16, 125–136 (2006).
Baxter, J. et al. Histone hypomethylation is an indicator of epigenetic plasticity in quiescent lymphocytes. EMBO J. 23, 4462–4472 (2004).
Martens, J.H. et al. The profile of repeat-associated histone lysine methylation states in the mouse epigenome. EMBO J. 24, 800–812 (2005).
Rice, J.C. et al. Histone methyltransferases direct different degrees of methylation to define distinct chromatin domains. Mol. Cell 12, 1591–1598 (2003).
Schneider, R. et al. Histone H3 lysine 4 methylation patterns in higher eukaryotic genes. Nat. Cell Biol. 6, 73–77 (2004).
Fischle, W. et al. Regulation of HP1-chromatin binding by histone H3 methylation and phosphorylation. Nature 438, 1116–1122 (2005).
Sims, R.J. III et al. Human but not yeast CHD1 binds directly and selectively to histone H3 methylated at lysine 4 via its tandem chromodomains. J. Biol. Chem. 280, 41789–41792 (2005).
Gregory, R.I. et al. Inhibition of histone deacetylases alters allelic chromatin conformation at the imprinted U2af1-rs1 locus in mouse embryonic stem cells. J. Biol. Chem. 277, 11728–11734 (2002).
Hattori, N. et al. Epigenetic control of mouse Oct-4 gene expression in embryonic stem cells and trophoblast stem cells. J. Biol. Chem. 279, 17063–17069 (2004).
O'Neill, L.P. et al. X-linked genes in female embryonic stem cells carry an epigenetic mark prior to the onset of X inactivation. Hum. Mol. Genet. 12, 1783–1790 (2003).
Szutorisz, H. et al. Formation of an active tissue-specific chromatin domain initiated by epigenetic marking at the embryonic stem cell stage. Mol. Cell. Biol. 25, 1804–1820 (2005).
Handyside, A.H. & Hunter, S. A rapid procedure for visualising the inner cell mass and trophectoderm nuclei of mouse blastocysts in situ using polynucleotide-specific fluorochromes. J. Exp. Zool. 231, 429–434 (1984).
Harvey, M.B. & Kaye, P.L. Insulin increases the cell number of the inner cell mass and stimulates morphological development of mouse blastocysts in vitro. Development 110, 963–967 (1990).
Adjaye, J. et al. Primary differentiation in the human blastocyst: comparative molecular portraits of inner cell mass and trophectoderm cells. Stem Cells 23, 1514–1525 (2005).
Rugg-Gunn, P.J., Ferguson-Smith, A.C. & Pedersen, R.A. Epigenetic status of human embryonic stem cells. Nat. Genet. 37, 585–587 (2005).
Pokholok, D.K. et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122, 517–527 (2005).
Lachner, M., O'Sullivan, R.J. & Jenuwein, T. An epigenetic road map for histone lysine methylation. J. Cell Sci. 116, 2117–2124 (2003).
Schubeler, D. et al. The histone modification pattern of active genes revealed through genome-wide chromatin analysis of a higher eukaryote. Genes Dev. 18, 1263–1271 (2004).
Strumpf, D. et al. Cdx2 is required for correct cell fate specification and differentiation of trophectoderm in the mouse blastocyst. Development 132, 2093–2102 (2005).
Vakoc, C.R., Mandat, S.A., Olenchock, B.A. & Blobel, G.A. Histone H3 lysine 9 methylation and HP1gamma are associated with transcription elongation through mammalian chromatin. Mol. Cell 19, 381–391 (2005).
Solter, D. & Knowles, B.B. Immunosurgery of mouse blastocyst. Proc. Natl. Acad. Sci. USA 72, 5099–5102 (1975).
Jones, P.A. & Martienssen, R. A blueprint for a Human Epigenome Project: the AACR Human Epigenome Workshop. Cancer Res. 65, 11241–11246 (2005).
Hebbes, T.R., Thorne, A.W. & Crane-Robinson, C. A direct link between core histone acetylation and transcriptionally active chromatin. EMBO J. 7, 1395–1402 (1988).
O'Neill, L.P. & Turner, B.M. Histone H4 acetylation distinguishes coding regions of the human genome from heterochromatin in a differentiation-dependent but transcription-independent manner. EMBO J. 14, 3946–3957 (1995).
Orlando, V. Mapping chromosomal proteins in vivo by formaldehyde-crosslinked-chromatin immunoprecipitation. Trends Biochem. Sci. 25, 99–104 (2000).
Geisberg, J.V. & Struhl, K. Quantitative sequential chromatin immunoprecipitation, a method for analyzing co-occupancy of proteins at genomic regions in vivo. Nucleic Acids Res. 32, e151 (2004).
Kuo, M.H. & Allis, C.D. In vivo cross-linking and immunoprecipitation for studying dynamic Protein:DNA associations in a chromatin environment. Methods 19, 425–433 (1999).
Robyr, D. & Grunstein, M. Genomewide histone acetylation microarrays. Methods 31, 83–89 (2003).
Suka, N., Suka, Y., Carmen, A.A., Wu, J. & Grunstein, M. Highly specific antibodies determine histone acetylation site usage in yeast heterochromatin and euchromatin. Mol. Cell 8, 473–479 (2001).
Gregory, R.I. & Feil, R. Analysis of chromatin in limited numbers of cells: a PCR-SSCP based assay of allele-specific nuclease sensitivity. Nucleic Acids Res. 27, e32 (1999).
Santos, F. et al. Epigenetic marking correlates with developmental potential in cloned bovine preimplantation embryos. Curr. Biol. 13, 1116–1121 (2003).
Schneider, I. Cell lines derived from late embryonic stages of Drosophila melanogaster. J. Embryol. Exp. Morphol. 27, 353–365 (1972).
Norris, D.P. et al. Evidence that random and imprinted Xist expression is controlled by preemptive methylation. Cell 77, 41–51 (1994).
O'Neill, L.P. et al. A developmental switch in H4 acetylation upstream of Xist plays a role in X chromosome inactivation. EMBO J. 18, 2897–2907 (1999).
Turner, B.M. & Fellows, G. Specific antibodies reveal ordered and cell-cycle-related use of histone-H4 acetylation sites in mammalian cells. Eur. J. Biochem. 179, 131–139 (1989).
White, D.A., Belyaev, N.D. & Turner, B.M. Preparation of site-specific antibodies to acetylated histones. Methods 19, 417–424 (1999).
Acknowledgements
We are grateful to S. Parnell for help with preparation of mRNA from small numbers of cells, to V. Gupta for advice on statistical analyses and to J. Frampton and W. Reik for comments on the manuscript. L.P.O. is a Royal Society Research Fellow and M.D.V. is supported by a PhD studentship from the University of Birmingham Medical School. This work was funded by Cancer Research UK and the Biotechnology and Biological Sciences Research Council (BBSRC).
Author information
Authors and Affiliations
Contributions
L.P.O.: validation and development of CChIP technique, study design, planning and carrying out experiments, data analysis, writing paper. M.D.V.: growth and dissection of embryos, carrying out experiments, data analysis, assistance in writing paper. B.M.T.: concept, study design, planning experiments, data analysis, writing paper.
Corresponding author
Ethics declarations
Competing interests
The CChIP procedure described in this paper is the subject of British patent application number 0601538.2 (B.M.T., L.P.O.).
Supplementary information
Supplementary Fig. 1
Levels of H4K16ac and H3K9me2 on silent genes in ES cells and ICM assayed by CChIP. (PDF 12 kb)
Supplementary Table 1
Levels of H4K16 acetylation and H3K9 dimethylation at selected genes and gene regions in ICM and ES cells. (PDF 11 kb)
Supplementary Table 2
Primers used for CChIP and expression analysis. (PDF 11 kb)
Rights and permissions
About this article
Cite this article
O'Neill, L., VerMilyea, M. & Turner, B. Epigenetic characterization of the early embryo with a chromatin immunoprecipitation protocol applicable to small cell populations. Nat Genet 38, 835–841 (2006). https://doi.org/10.1038/ng1820
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1820
This article is cited by
-
Laboratory methods to decipher epigenetic signatures: a comparative review
Cellular & Molecular Biology Letters (2021)
-
Dynamic regulation of Z-DNA in the mouse prefrontal cortex by the RNA-editing enzyme Adar1 is required for fear extinction
Nature Neuroscience (2020)
-
A PAX5–OCT4–PRDM1 developmental switch specifies human primordial germ cells
Nature Cell Biology (2018)