Abstract
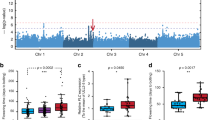
Light has an important role in modulating seedling growth and flowering time. We show that allelic variation at the PHYTOCHROME C (PHYC) photoreceptor locus affects both traits in natural populations of A. thaliana. Two functionally distinct PHYC haplotype groups are distributed in a latitudinal cline dependent on FRIGIDA, a locus that together with FLOWERING LOCUS C explains a large portion of the variation in A. thaliana flowering time1. In a genome-wide scan for association of 65 loci with latitude, there was an excess of significant P values, indicative of population structure. Nevertheless, PHYC was the most strongly associated locus across 163 strains, suggesting that PHYC alleles are under diversifying selection in A. thaliana. Our work, together with previous findings2,3,4,5, suggests that photoreceptor genes are major agents of natural variation in plant flowering and growth response.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Johanson, U. et al. Molecular analysis of FRIGIDA, a major determinant of natural variation in Arabidopsis flowering time. Science 290, 344–347 (2000).
Aukerman, M.J. et al. A deletion in the PHYD gene of the Arabidopsis Wassilewskija ecotype defines a role for phytochrome D in red/far-red light sensing. Plant Cell 9, 1317–1326 (1997).
El-Assal, S.E.-D., Alonso-Blanco, C., Peeters, A.J., Raz, V. & Koornneef, M.A. QTL for flowering time in Arabidopsis reveals a novel allele of CRY2. Nat. Genet. 29, 435–440 (2001).
Maloof, J.N. et al. Natural variation in light sensitivity of Arabidopsis. Nat. Genet. 29, 441–446 (2001).
Olsen, K.M. et al. Linkage disequilibrium mapping of Arabidopsis CRY2 flowering time alleles. Genetics 167, 1361–1369 (2004).
Shindo, C. et al. Role of FRIGIDA and FLOWERING LOCUS C in determining variation in flowering time of Arabidopsis. Plant Physiol. 138, 1163–1173 (2005).
Lempe, J. et al. Diversity of flowering responses in wild Arabidopsis thaliana strains. PLoS Genet. 1, e6 (2005).
Werner, J.D. et al. FRIGIDA-independent variation in flowering time of natural Arabidopsis thaliana accessions. Genetics 170, 1197–1207 (2005).
Werner, J.D. et al. Quantitative trait locus mapping and DNA array hybridization identify an FLM deletion as a cause for natural flowering-time variation. Proc. Natl. Acad. Sci. USA 102, 2460–2465 (2005).
Franklin, K.A., Davis, S.J., Stoddart, W.M., Vierstra, R.D. & Whitelam, G.C. Mutant analyses define multiple roles for phytochrome C in Arabidopsis photomorphogenesis. Plant Cell 15, 1981–1989 (2003).
Monte, E. et al. Isolation and characterization of phyC mutants in Arabidopsis reveals complex crosstalk between phytochrome signaling pathways. Plant Cell 15, 1962–1980 (2003).
Qin, M., Kuhn, R., Moran, S. & Quail, P.H. Overexpressed phytochrome C has similar photosensory specificity to phytochrome B but a distinctive capacity to enhance primary leaf expansion. Plant J. 12, 1163–1172 (1997).
Halliday, K.J., Thomas, B. & Whitelam, G.C. Expression of heterologous phytochromes A, B or C in transgenic tobacco plants alters vegetative development and flowering time. Plant J. 12, 1079–1090 (1997).
Weinig, C. et al. Novel loci control variation in reproductive timing in Arabidopsis thaliana in natural environments. Genetics 162, 1875–1884 (2002).
Wang, X., Korstanje, R., Higgins, D. & Paigen, B. Haplotype analysis in multiple crosses to identify a QTL gene. Genome Res. 14, 1767–1772 (2004).
Alonso-Blanco, C., El-Assal, S.E., Coupland, G. & Koornneef, M. Analysis of natural allelic variation at flowering time loci in the Landsberg erecta and Cape Verde Islands ecotypes of Arabidopsis thaliana. Genetics 149, 749–764 (1998).
Borevitz, J.O. et al. Quantitative trait loci controlling light and hormone response in two accessions of Arabidopsis thaliana. Genetics 160, 683–696 (2002).
Loudet, O., Chaillou, S., Camilleri, C., Bouchez, D. & Daniel-Vedele, F. Bay-0 x Shahdara recombinant inbred line population: a powerful tool for the genetic dissection of complex traits in Arabidopsis. Theor. Appl. Genet. 104, 1173–1184 (2002).
El-Lithy, M.E., Clerkx, E.J., Ruys, G.J., Koornneef, M. & Vreugdenhil, D. Quantitative trait locus analysis of growth-related traits in a new Arabidopsis recombinant inbred population. Plant Physiol. 135, 444–458 (2004).
Wolyn, D.J. et al. Light-response quantitative trait loci identified with composite interval and eXtreme array mapping in Arabidopsis thaliana. Genetics 167, 907–917 (2004).
Stinchcombe, J.R. et al. A latitudinal cline in flowering time in Arabidopsis thaliana modulated by the flowering time gene FRIGIDA. Proc. Natl. Acad. Sci. USA 101, 4712–4717 (2004).
Schmid, K.J. et al. Evidence for a large-scale population structure of Arabidopsis thaliana from genome-wide single nucleotide polymorphism markers. Theor. Appl. Genet. 112, 1104–1114 (2006).
Caicedo, A.L., Stinchcombe, J.R., Olsen, K.M., Schmitt, J. & Purugganan, M.D. Epistatic interaction between Arabidopsis FRI and FLC flowering time genes generates a latitudinal cline in a life history trait. Proc. Natl. Acad. Sci. USA 101, 15670–15675 (2004).
Wagner, D. & Quail, P.H. Mutational analysis of phytochrome B identifies a small COOH-terminal-domain region critical for regulatory activity. Proc. Natl. Acad. Sci. USA 92, 8596–8600 (1995).
Xu, Y., Parks, B.M., Short, T.W. & Quail, P.H. Missense mutations define a restricted segment in the C-terminal domain of phytochrome A critical to its regulatory activity. Plant Cell 7, 1433–1443 (1995).
Reed, J.W., Nagpal, P., Poole, D.S., Furuya, M. & Chory, J. Mutations in the gene for the red/far-red light receptor phytochrome B alter cell elongation and physiological responses throughout Arabidopsis development. Plant Cell 5, 147–157 (1993).
Takano, M. et al. Distinct and cooperative functions of phytochromes A, B, and C in the control of deetiolation and flowering in rice. Plant Cell 17, 3311–3325 (2005).
Mathews, S. & Donoghue, M.J. The root of angiosperm phylogeny inferred from duplicate phytochrome genes. Science 286, 947–950 (1999).
White, G.M., Hamblin, M.T. & Kresovich, S. Molecular evolution of the phytochrome gene family in sorghum: changing rates of synonymous and replacement evolution. Mol. Biol. Evol. 21, 716–723 (2004).
Devos, K.M., Beales, J., Ogihara, Y. & Doust, A.N. Comparative sequence analysis of the Phytochrome C gene and its upstream region in allohexaploid wheat reveals new data on the evolution of its three constituent genomes. Plant Mol. Biol. 58, 625–641 (2005).
Acknowledgements
We thank the Nottingham Arabidopsis Stock Center for seed stocks of SALK T-DNA lines donated by J. Ecker and colleagues. We thank C. Lanz for help with sequencing, and K. Bomblies and Y.L. Guo for help with phylogenetic analysis. We thank K. Bomblies, V. Grbic, Y. Kobayashi, J. Lempe and S. Russell for discussion and critical reading of the manuscript. This work was supported by a European Molecular Biology Organization (EMBO) Long-Term Fellowship to S.B., a US National Institutes of Health (NIH) Postdoctoral Fellowship to T.P.M., an NIH grant (GM62932) to J.C. and D.W., and by the Max Planck Society, of which D.W. is a director. J.C. is a Howard Hughes Medical Institute (HHMI) Investigator.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Markers used for mapping of the early flowering phenotype in Fr-2. (PDF 92 kb)
Supplementary Fig. 2
Flowering behavior of populations segregating for different PHYC alleles. (PDF 91 kb)
Supplementary Fig. 3
Latitudinal cline of PHYC alleles. (PDF 81 kb)
Supplementary Table 1
PHYC haplotypes of A. thaliana strains. (PDF 111 kb)
Supplementary Table 2
ANOVA of latitude by PHYC haplotype group interaction on residual variation in flowering time at 23 °C in short days after accounting for FRI functionality. (PDF 64 kb)
Supplementary Table 3
p-values obtained for different models testing for association with latitude (Latitude), FRI dependent interaction with latitude (Interaction), hypocotyl length under GA treatment (GAHypo) and total leaf number in short days (TLNSD) for 67 markers across 163 strains. (PDF 221 kb)
Supplementary Table 4
p-values obtained for two different models testing for latitudinal association with a completely independent set of markers across 56 strains. (PDF 204 kb)
Supplementary Table 5
Oligonucleotide primers. (PDF 68 kb)
Supplementary Table 6
Strains sequenced for PHYC. (PDF 87 kb)
Rights and permissions
About this article
Cite this article
Balasubramanian, S., Sureshkumar, S., Agrawal, M. et al. The PHYTOCHROME C photoreceptor gene mediates natural variation in flowering and growth responses of Arabidopsis thaliana. Nat Genet 38, 711–715 (2006). https://doi.org/10.1038/ng1818
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1818
This article is cited by
-
Indirect genetic effects are shaped by demographic history and ecology in Arabidopsis thaliana
Nature Ecology & Evolution (2023)
-
A base substitution in OsphyC disturbs its Interaction with OsphyB and affects flowering time and chlorophyll synthesis in rice
BMC Plant Biology (2022)
-
Molecular variation in a functionally divergent homolog of FCA regulates flowering time in Arabidopsis thaliana
Nature Communications (2020)
-
Natural variation in genes potentially involved in plant architecture and adaptation in switchgrass (Panicum virgatum L.)
BMC Evolutionary Biology (2018)
-
Single nucleotide polymorphisms in a regulatory site of VRN-A1 first intron are associated with differences in vernalization requirement in winter wheat
Molecular Genetics and Genomics (2018)