Abstract
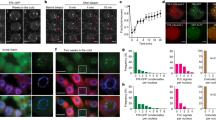
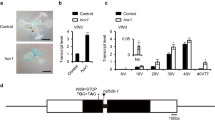
Vernalization is the process by which sensing a prolonged exposure to winter cold leads to competence to flower in the spring. In winter annual Arabidopsis thaliana accessions, flowering is suppressed in the fall by expression of the potent floral repressor FLOWERING LOCUS C (FLC)1. Vernalization promotes flowering via epigenetic repression of FLC2. Repression is accompanied by a series of histone modifications of FLC chromatin that include dimethylation of histone H3 at Lys9 (H3K9) and Lys27 (H3K27)3,4. Here, we report that A. thaliana LIKE HETEROCHROMATIN PROTEIN 1 (LHP1) is necessary to maintain the epigenetically repressed state of FLC upon return to warm conditions typical of spring. LHP1 is enriched at FLC chromatin after prolonged exposure to cold, and LHP1 activity is needed to maintain the increased levels of H3K9 dimethylation at FLC chromatin that are characteristic of the vernalized state.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Michaels, S.D. & Amasino, R.M. Memories of winter: vernalization and the competence to flower. Plant Cell Environ. 23, 1145–1154 (2000).
Sung, S. & Amasino, R.M. REMEMBERING WINTER: Toward a molecular understanding of vernalization. Annu. Rev. Plant Biol. 56, 491–508 (2005).
Sung, S. & Amasino, R.M. Vernalization in Arabidopsis thaliana is mediated by the PHD finger protein VIN3. Nature 427, 159–164 (2004).
Bastow, R. et al. Vernalization requires epigenetic silencing of FLC by histone methylation. Nature 427, 164–167 (2004).
Michaels, S.D. & Amasino, R.M. FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11, 949–956 (1999).
Sheldon, C.C. et al. The FLF MADS box gene: a repressor of flowering in Arabidopsis regulated by vernalization and methylation. Plant Cell 11, 445–458 (1999).
Gendall, A.R., Levy, Y.Y., Wilson, A. & Dean, C. The VERNALIZATION 2 gene mediates the epigenetic regulation of vernalization in Arabidopsis. Cell 107, 525–535 (2001).
Ringrose, L. & Paro, R. Epigenetic regulation of cellular memory by the Polycomb and Trithorax group proteins. Annu. Rev. Genet. 38, 413–443 (2004).
Schubert, D., Clarenz, O. & Goodrich, J. Epigenetic control of plant development by Polycomb-group proteins. Curr. Opin. Plant Biol. 8, 553–561 (2005).
Gaudin, V. et al. Mutations in LIKE HETEROCHROMATIN PROTEIN 1 affect flowering time and plant architecture in Arabidopsis. Development 128, 4847–4858 (2001).
Kotake, T., Takada, S., Nakahigashi, K., Ohto, M. & Goto, K. Arabidopsis TERMINAL FLOWER 2 gene encodes a heterochromatin protein 1 homolog and represses both FLOWERING LOCUS T to regulate flowering time and several floral homeotic genes. Plant Cell Physiol. 44, 555–564 (2003).
Maison, C. & Almouzni, G. HP1 and the dynamics of heterochromatin maintenance. Nat. Rev. Mol. Cell Biol. 5, 296–304 (2004).
Liu, L.P., Ni, J.Q., Shi, Y.D., Oakeley, E.J. & Sun, F.L. Sex-specific role of Drosophila melanogaster HP1 in regulating chromatin structure and gene transcription. Nat. Genet. 37, 1361–1366 (2005).
Nielsen, S.J. et al. Rb targets histone H3 methylation and HP1 to promoters. Nature 412, 561–565 (2001).
Nakahigashi, K., Jasencakova, Z., Schubert, I. & Goto, K. The Arabidopsis HETEROCHROMATIN PROTEIN1 homolog (TERMINAL FLOWER2) silences genes within euchromatic region but not genes positioned in heterochromatin. Plant Cel. Physiol. 46, 1747–1756 (2005).
Libault, M. et al. The Arabidopsis LHP1 protein is a component of euchromatin. Planta 222, 910–925 (2005).
Zemach, A. et al. Different domains control the localization and mobility of LIKE HETEROCHROMATIN PROTEIN1 in Arabidopsis nuclei. Plant Cell 18, 133–145 (2006).
Lindroth, A.M. et al. Dual histone H3 methylation marks at lysines 9 and 27 required for interaction with CHROMOMETHYLASE3. EMBO J. 23, 4286–4296 (2004).
Lee, I. & Amasino, R.M. Effect of vernalization, photoperiod, and light quality on the flowering phenotype of Arabidopsis plants containing the FRIGIDA gene. Plant Physiol. 108, 157–162 (1995).
Mylne, J.S. et al. LHP1, the Arabidopsis homologue of HETEROCHROMATIN PROTEIN1, is required for epigenetic silencing of FLC. Proc. Natl. Acad. Sci. USA 103, 5012–5017 (2006).
Valverde, F. et al. Photoreceptor regulation of CONSTANS protein in photoperiodic flowering. Science 303, 1003–1006 (2004).
Huang, T., Bohlenius, H., Eriksson, S., Parcy, F. & Nilsson, O. The mRNA of the Arabidopsis gene FT moves from leaf to shoot apex and induces flowering. Science 309, 1694–1696 (2005).
Sheldon, C.C., Conn, A.B., Dennis, E.S. & Peacock, W.J. Different regulatory regions are required for the vernalization-induced repression of FLOWERING LOCUS C and for the epigenetic maintenance of repression. Plant Cell 14, 2527–2537 (2002).
Baumbusch, L.O. et al. The Arabidopsis thaliana genome contains at least 29 active genes encoding SET domain proteins that can be assigned to four evolutionarily conserved classes. Nucleic Acids Res. 29, 4319–4333 (2001).
Dejardin, J. et al. Recruitment of Drosophila Polycomb group proteins to chromatin by DSP1. Nature 434, 533–538 (2005).
He, Y., Doyle, M.R. & Amasino, R.M. PAF1-complex-mediated histone methylation of FLOWERING LOCUS C chromatin is required for the vernalization-responsive, winter-annual habit in Arabidopsis. Genes Dev. 18, 2774–2784 (2004).
Krogan, N.J. et al. The Paf1 complex is required for histone H3 methylation by COMPASS and Dot1p: linking transcriptional elongation to histone methylation. Mol. Cell 11, 721–729 (2003).
Kim, S.Y. et al. Establishment of the vernalization-responsive, winter-annual habit in Arabidopsis requires a putative histone H3 methyl transferase. Plant Cell 17, 3301–3310 (2005).
Jenuwein, T. & Allis, C.D. Translating the histone code. Science 293, 1074–1080 (2001).
Johnson, L., Cao, X. & Jacobsen, S. Interplay between two epigenetic marks. DNA methylation and histone H3 lysine 9 methylation. Curr. Biol. 12, 1360–1367 (2002).
Acknowledgements
We are grateful to S. Woody, R. Schmitz and R. Rodman for their comments on the manuscript. Research in the laboratory of R.M.A. was supported by the College of Agricultural and Life Sciences of the University of Wisconsin and by grants from the US Department of Agriculture National Research Initiative Competitive Grants Program and the National Science Foundation. Research in the laboratory of S.E.J. was supported by US National Institute of Health grant GM060398.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
LHP1 localization. (PDF 257 kb)
Supplementary Table 1
Primers used for ChIP and RT-PCR. (PDF 38 kb)
Rights and permissions
About this article
Cite this article
Sung, S., He, Y., Eshoo, T. et al. Epigenetic maintenance of the vernalized state in Arabidopsis thaliana requires LIKE HETEROCHROMATIN PROTEIN 1. Nat Genet 38, 706–710 (2006). https://doi.org/10.1038/ng1795
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1795
This article is cited by
-
Genome-wide screening and characterization of long noncoding RNAs involved in flowering/bolting of Lactuca sativa
BMC Plant Biology (2023)
-
Polycomb Repressive Complexes and Their Roles in Plant Developmental Programs, Particularly Floral Transition
Journal of Plant Biology (2023)
-
The metabolic changes that effect fruit quality during tomato fruit ripening
Molecular Horticulture (2022)
-
HEAT SHOCK TRANSCRIPTION FACTOR B2b acts as a transcriptional repressor of VIN3, a gene induced by long-term cold for flowering
Scientific Reports (2022)
-
Parallel reduction in flowering time from de novo mutations enable evolutionary rescue in colonizing lineages
Nature Communications (2022)