Abstract
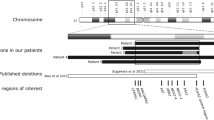
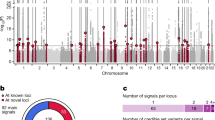
Dysfunction of the exocrine pancreas is observed in diabetes, but links between concurrent exocrine and endocrine pancreatic disease and contributing genetic factors are poorly characterized. We studied two families with diabetes and exocrine pancreatic dysfunction by genetic, physiological and in vitro functional studies. A genome-wide screen in Family 1 linked diabetes to chromosome 9q34 (maximal lod score 5.07). Using fecal elastase deficiency as a marker of exocrine pancreatic dysfunction refined the critical chromosomal region to 1.16 Mb (maximal lod score 11.6). Here, we identified a single-base deletion in the variable number of tandem repeats (VNTR)-containing exon 11 of the carboxyl ester lipase (CEL) gene, a major component of pancreatic juice and responsible for the duodenal hydrolysis of cholesterol esters. Screening subjects with maturity-onset diabetes of the young identified Family 2, with another single-base deletion in CEL and a similar phenotype with beta-cell failure and pancreatic exocrine disease. The in vitro catalytic activities of wild-type and mutant CEL protein were comparable. The mutant enzyme was, however, less stable and secreted at a lower rate. Furthermore, we found some evidence for an association between common insertions in the CEL VNTR and exocrine dysfunction in a group of 182 unrelated subjects with diabetes (odds ratio 4.2 (1.6, 11.5)). Our findings link diabetes to the disrupted function of a lipase in the pancreatic acinar cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Henderson, J.R. Why are the islets of Langerhans? Lancet 2, 469–470 (1969).
Cavalot, F. et al. Pancreatic elastase-1 in stools, a marker of exocrine pancreas function, correlates with both residual beta-cell secretion and metabolic control in type 1 diabetic subjects. Diabetes Care 27, 2052–2054 (2004).
Mitchell, R.M., Byrne, M.F. & Baillie, J. Pancreatitis. Lancet 361, 1447–1455 (2003).
Wang, F., Herrington, M., Larsson, J. & Permert, J. The relationship between diabetes and pancreatic cancer. Mol. Cancer 2, 4 (2003).
Lombardo, F. et al. Natural history of glucose tolerance, beta-cell function and peripheral insulin sensitivity in cystic fibrosis patients with fasting euglycemia. Eur. J. Endocrinol. 149, 53–59 (2003).
Stoffers, D.A., Zinkin, N.T., Stanojevic, V., Clarke, W.L. & Habener, J.F. Pancreatic agenesis attributable to a single nucleotide deletion in the human IPF1 gene coding sequence. Nat. Genet. 15, 106–110 (1997).
Sellick, G.S. et al. Mutations in PTF1A cause pancreatic and cerebellar agenesis. Nat. Genet. 36, 1301–1305 (2004).
Bellanne-Chantelot, C. et al. Clinical spectrum associated with hepatocyte nuclear factor-1beta mutations. Ann. Intern. Med. 140, 510–517 (2004).
Haumaitre, C. et al. Lack of TCF2/vHNF1 in mice leads to pancreas agenesis. Proc. Natl. Acad. Sci. USA 102, 1490–1495 (2005).
Edlund, H. Pancreatic organogenesis – developmental mechanisms and implications for therapy. Nat. Rev. Genet. 3, 524–532 (2002).
Hardt, P.D. et al. High prevalence of exocrine pancreatic insufficiency in diabetes mellitus. A multicenter study screening fecal elastase 1 concentrations in 1,021 diabetic patients. Pancreatology 3, 395–402 (2003).
Fajans, S.S., Bell, G.I. & Polonsky, K.S. Molecular mechanisms and clinical pathophysiology of maturity-onset diabetes of the young. N. Engl. J. Med. 345, 971–980 (2001).
Bengtsson-Ellmark, S.H. et al. Association between a polymorphism in the carboxyl ester lipase gene and serum cholesterol profile. Eur. J. Hum. Genet. 12, 627–632 (2004).
Lindquist, S., Bläckberg, L. & Hernell, O. Human bile salt-stimulated lipase has a high frequency of size variation due to a hypervariable region in exon 11. Eur. J. Biochem. 269, 759–767 (2002).
Higuchi, S., Nakamura, Y. & Saito, S. Characterization of a VNTR polymorphism in the coding region of the CEL gene. J. Hum. Genet. 47, 213–215 (2002).
Bjørkhaug, L. et al. Hepatocyte nuclear factor-1 alpha gene mutations and diabetes in Norway. J. Clin. Endocrinol. Metab. 88, 920–931 (2003).
Harding, H.P. & Ron, D. Endoplasmic reticulum stress and the development of diabetes: a review. Diabetes 51 (Suppl.) 3, S455–S461 (2002).
Kim, S.K. et al. Pbx1 inactivation disrupts pancreas development and in Ipf1-deficient mice promotes diabetes mellitus. Nat. Genet. 30, 430–435 (2002).
Iglesias, A. et al. Diabetes and exocrine pancreatic insufficiency in E2F1/E2F2 double-mutant mice. J. Clin. Invest. 113, 1398–1407 (2004).
Lombardo, D. Bile salt-dependent lipase: its pathophysiological implications. Biochim. Biophys. Acta 1533, 1–28 (2001).
Hui, D.Y. & Howles, P.N. Carboxyl ester lipase: structure-function relationship and physiological role in lipoprotein metabolism and atherosclerosis. J. Lipid Res. 43, 2017–2030 (2002).
Poorkhalkali, N. et al. Bile salt-stimulated lipase (BSSL) distribution in rat, mouse and transgenic mouse expressing human BSSL. Histochem. Cell Biol. 110, 367–376 (1998).
Auge, N. et al. Pancreatic bile salt-dependent lipase induces smooth muscle cells proliferation. Circulation 108, 86–91 (2003).
DiPersio, L.P., Fontaine, R.N. & Hui, D.Y. Identification of the active site serine in pancreatic cholesterol esterase by chemical modification and site-specific mutagenesis. J. Biol. Chem. 265, 16801–16806 (1990).
Aubert, E., Sbarra, V., Le Petit-Thevenin, J., Valette, A. & Lombardo, D. Site-directed mutagenesis of the basic N-terminal cluster of pancreatic bile salt-dependent lipase. Functional significance. J. Biol. Chem. 277, 34987–34996 (2002).
Aubert-Jousset, E., Sbarra, V. & Lombardo, D. Site-directed mutagenesis of the distal basic cluster of pancreatic bile salt-dependent lipase. J. Biol. Chem. 279, 39697–39704 (2004).
Downs, D., Xu, Y.Y., Tang, J. & Wang, C.S. Proline-rich domain and glycosylation are not essential for the enzymic activity of bile salt-activated lipase. Kinetic studies of T-BAL, a truncated form of the enzyme, expressed in Escherichia coli. Biochemistry 33, 7979–7985 (1994).
Gleghorn, L., Ramesar, R., Beighton, P. & Wallis, G. A mutation in the variable repeat region of the aggrecan gene (AGC1) causes a form of spondyloepiphyseal dysplasia associated with severe, premature osteoarthritis. Am. J. Hum. Genet. 77, 484–490 (2005).
Howles, P.N., Carter, C.P. & Hui, D.Y. Dietary free and esterified cholesterol absorption in cholesterol esterase (bile salt-stimulated lipase) gene-targeted mice. J. Biol. Chem. 271, 7196–7202 (1996).
Weng, W. et al. Intestinal absorption of dietary cholesteryl ester is decreased but retinyl ester absorption is normal in carboxyl ester lipase knockout mice. Biochemistry 38, 4143–4149 (1999).
Herman, G.E. Mouse models of human disease: lessons learned and promises to come. ILAR J. 43, 55–56 (2002).
Panicot, L., Mas, E., Thivolet, C. & Lombardo, D. Circulating antibodies against an exocrine pancreatic enzyme in type 1 diabetes. Diabetes 48, 2316–2323 (1999).
Lillioja, S. et al. Insulin resistance and insulin secretory dysfunction as precursors of non-insulin-dependent diabetes mellitus. Prospective studies of Pima Indians. N. Engl. J. Med. 329, 1988–1992 (1993).
Goda, K. et al. Pancreatic volume in type 1 and type 2 diabetes mellitus. Acta Diabetol. 38, 145–149 (2001).
Vaxillaire, M. et al. A gene for maturity onset diabetes of the young (MODY) maps to chromosome 12q. Nat. Genet. 9, 418–423 (1995).
Icks, A., Haastert, B., Giani, G. & Rathmann, W. Low fecal elastase-1 in type I diabetes mellitus. Z. Gastroenterol. 39, 823–830 (2001).
Rathmann, W. et al. Low faecal elastase 1 concentrations in type 2 diabetes mellitus. Scand. J. Gastroenterol. 36, 1056–1061 (2001).
Verine, A., Le Petit-Thevenin, J., Panicot-Dubois, L., Valette, A. & Lombardo, D. Phosphorylation of the oncofetal variant of the human bile salt-dependent lipase. identification of phosphorylation site and relation with secretion process. J. Biol. Chem. 276, 12356–12361 (2001).
Acknowledgements
We thank L. Bindoff, G. Eide, G. Fluge and H. Immervoll for discussions; L. Marselli and G. Weir for sharing unpublished information; S. Aanderud and E. Husebye for providing outpatients; B.H. Berge and L. Aasmul for technical assistance; E. Aubert-Jousset for catalytic constant determinations; B. Mallet and L. Benkoel for assays of C-reactive protein, and orosomucoid and immunohistochemistry experiments, respectively; and the families involved in the study. This work was supported in part by funds from Helse Vest, Haukeland University Hospital and Innovest (to the Department of Pediatrics, Haukeland University Hospital, Bergen), from the University of Bergen, the Meltzer Foundation, Norwegian Diabetes Association and Health & Rehabilitation (to the Institute of Clinical Medicine, University of Bergen, Bergen, Norway) and from INSERM and the Université de la Méditerranée (to the INSERM U-559, Marseille, France).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Pedigree of studied branches of Family 2. (PDF 20 kb)
Supplementary Fig. 2
Pancreas morphology investigated by abdominal CT. (PDF 116 kb)
Supplementary Fig. 3
Sequencing a CEL polymorphism to investigate mutant CEL expression in fibroblasts. (PDF 63 kb)
Supplementary Table 1
Markers with two-point lod scores >1.17 (P < 0.01) in the genome scan of the core pedigree of 61 subjects in Family 1. (PDF 16 kb)
Supplementary Table 2
Two-point lod scores at different theta values between diabetes or elastase deficiency and chromosome 9 saturation markers in the extended pedigree of 184 subjects in Family 1. (PDF 74 kb)
Supplementary Table 3
Clinical characteristics of patients in Family 1 with mutation in CEL. (PDF 21 kb)
Supplementary Table 4
Clinical characteristics of patients in Family 2 with mutation in CEL. (PDF 22 kb)
Rights and permissions
About this article
Cite this article
Ræder, H., Johansson, S., Holm, P. et al. Mutations in the CEL VNTR cause a syndrome of diabetes and pancreatic exocrine dysfunction. Nat Genet 38, 54–62 (2006). https://doi.org/10.1038/ng1708
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1708
This article is cited by
-
Monogenic diabetes
Nature Reviews Disease Primers (2023)
-
Monogenic Diabetes Reported in South Asians: A Systematic Review
Journal of the Indian Institute of Science (2023)
-
Clinical and genetic characteristics of CEL-MODY (MODY8): a literature review and screening in Chinese individuals diagnosed with early-onset type 2 diabetes
Endocrine (2023)
-
Incomplete penetrance and variable expressivity in monogenic diabetes; a challenge but also an opportunity
Reviews in Endocrine and Metabolic Disorders (2023)
-
Abnormal exocrine–endocrine cell cross-talk promotes β-cell dysfunction and loss in MODY8
Nature Metabolism (2022)