Abstract
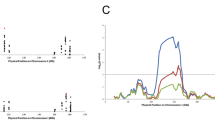
Multiple sclerosis is a common disease with proven heritability, but, despite large-scale attempts, no underlying risk genes have been identified. Traditional linkage scans have so far identified only one risk haplotype for multiple sclerosis (at HLA on chromosome 6), which explains only a fraction of the increased risk to siblings. Association scans such as admixture mapping have much more power, in principle, to find the weak factors that must explain most of the disease risk. We describe here the first high-powered admixture scan, focusing on 605 African American cases and 1,043 African American controls, and report a locus on chromosome 1 that is significantly associated with multiple sclerosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rife, D.C. Populations of hybrid origin as source material for the detection of linkage. Am. J. Hum. Genet. 6, 26–33 (1954).
Chakraborty, R. & Weiss, K.M. Admixture as a tool for finding linked genes and detecting that difference from allelic association between loci. Proc. Natl. Acad. Sci. USA 85, 9119–9123 (1988).
Stephens, J.C., Briscoe, D. & O'Brien, S.J. Mapping by admixture linkage disequilibrium in human populations: Limits and guidelines. Am. J. Hum. Genet. 55, 809–824 (1994).
Briscoe, D., Stephens, J.C. & O'Brien, S.J. Linkage disequilibrium in admixed populations: Applications in gene mapping. J. Hered. 85, 59–63 (1994).
McKeigue, P.M. Mapping genes underlying ethnic differences in disease risk by linkage disequilibrium in recently admixed populations. Am. J. Hum. Genet. 60, 188–196 (1997).
McKeigue, P.M. Mapping genes that underlie ethnic differences in disease risk: methods for detecting linkage in populations, by conditioning on parental admixture. Am. J. Hum. Genet. 63, 241–251 (1998).
Smith, M.W. et al. A high-density admixture map for disease gene discovery in African Americans. Am. J. Hum. Genet. 74, 1001–1013 (2004).
Patterson, N. et al. Methods for high-density admixture mapping of disease genes. Am. J. Hum. Genet. 74, 979–1000 (2004).
Hoggart, C.J., Shriver, M.D., Kittles, R.A., Clayton, D.G. & McKeigue, P.M. Design and analysis of admixture mapping studies. Am. J. Hum. Genet. 74, 965–978 (2004).
Montana, G. & Pritchard, J.K. Statistical tests for admixture mapping with case-control and cases-only data. Am. J. Hum. Genet. 75, 771–789 (2004).
Zhu, X. et al. Admixture mapping for hypertension loci with genome-scan markers. Nat. Genet. 37, 177–181 (2005).
Carlson, C.S. et al. Additional SNPs and linkage-disequilibrium analyses are necessary for whole-genome association studies in humans. Nat. Genet. 33, 518–521 (2003).
Reich, D. & Patterson, N. Will admixture mapping work to find disease genes? Phil. Trans. R. Soc. Lond. B 360, 1605–1607 (2005).
Kurtzke, J.F., Beebe, G.W. & Norman, J.E. Jr. Epidemiology of multiple sclerosis in U.S. veterans: 1. Race, sex, and geographic distribution. Neurology 29, 1228–1235 (1979).
Wallin, M.T., Page, W.F. & Kurtzke, J.F. Multiple sclerosis in US veterans of the Vietnam era and later military service: race, sex, and geography. Ann. Neurol. 55, 65–71 (2004).
The Transatlantic Multiple Sclerosis Genetics Cooperative. A meta-analysis of genomic screens in multiple sclerosis. A meta-analysis of genomic screens in multiple sclerosis. Mult. Scler. 7, 3–11 (2001).
Oksenberg, J.R. et al. Mapping multiple sclerosis susceptibility to the HLA-DR locus in African Americans. Am. J. Hum. Genet. 74, 160–167 (2004).
Hoggart, C.J. et al. Control of confounding of genetic associations in stratified populations. Am. J. Hum. Genet. 72, 1492–1504 (2003).
Quelvennec, E. et al. Genetic and functional studies in multiple sclerosis patients from Martinique attest for a specific and direct role of the HLA-DR locus in the syndrome. Tissue Antigens 61, 166–171 (2003).
Kelly, M.A. et al. An investigation of HLA-encoded genetic susceptibility to multiple sclerosis in subjects of Asian Indian and Afro-Caribbean ethnic origin. Tissue Antigens 45, 197–202 (1995).
Lauer, K. The risk of multiple sclerosis in the U.S.A. in relation to sociogeographic features: a factor-analytic study. J. Clin. Epidemiol. 47, 43–48 (1994).
International Multiple Sclerosis Genetics Consortium. A high-density screen for linkage in multiple sclerosis. Am. J. Hum. Genet. 77, 454–467 (2005).
Cree, B.A. et al. Clinical characteristics of African Americans vs Caucasian Americans with multiple sclerosis. Neurology 63, 2039–2045 (2004).
McDonald, W.I. et al. Recommended diagnostic criteria for multiple sclerosis: guidelines from the International Panel on the Diagnosis of Multiple Sclerosis. Ann. Neurol. 50, 121–127 (2001).
Poser, C.M. et al. New diagnostic criteria for multiple sclerosis: guidelines for research protocols. Ann. Neurol. 13, 227–231 (1983).
Hosono, S. et al. Unbiased whole-genome amplification directly from clinical samples. Genome Res. 13, 954–964 (2003).
Fan, J.B. et al. Highly parallel SNP genotyping. Cold Spring Harb. Symp. Quant. Biol. 68, 69–78 (2003).
Tang, K. et al. Chip-based genotyping by mass spectrometry. Proc. Natl. Acad. Sci. USA 96, 10016–10020 (1999).
Excoffier, L. & Slatkin, M. Maximum-likelihood estimation of molecular haplotype frequencies in a diploid population. Mol. Biol. Evol. 12, 921–927 (1995).
Kong, X. et al. A combined linkage-physical map of the human genome. Am. J. Hum. Genet. 75, 1143–1148 (2004).
Acknowledgements
We thank the individuals with multiple sclerosis, their families and friends and others for permission to use their DNA in these studies; M. Williams for his role in building the collection of samples from African Americans; B. Henderson and S. Ingles for permission to use data from the prostate cancer study; D. Goldstein for frequency information from the Italian and Norwegian samples; and D. Goldstein and E. Lander for providing comments throughout the project. This work was supported by grants to D.R. from the Wadsworth Foundation and the US National Institute of Neurological Disorders and Stroke; to D.A.H. from the National Multiple Sclerosis Society; to J.R.O. from the US National Institute of Neurological Disorders and Stroke and the National Multiple Sclerosis Society; and to S.L.H. from the Nancy David and Montel Williams Foundations. D.R. is the recipient of a Burroughs-Wellcome Career Development Award in the Biomedical Sciences; B.A.C.C. is a Sylvia Lawry fellow of the National Multiple Sclerosis Society; N.P. has a US National Institutes of Health career transition award; D.A.H. has a US National Institute of Neurological Disorders and Stroke Javits Investigator Award; and P.L.D.J. is the William C Fowler scholar for MS Research and is supported by US National Institute of Neurological Disorders and Stroke grant and the Clinical Investigator Training program: Harvard-MIT Health Sciences and Technology–Beth Israel Deaconess Medical Center, in collaboration with Pfizer, Inc.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Results for the 1,166 SNPs used in the main MCMC analysis (analysis from Run 3). (XLS 435 kb)
Supplementary Table 2
Proof that case-control statistic is normally distributed. (PDF 44 kb)
Supplementary Table 3
Admixture statistics for the populations under study. (PDF 48 kb)
Supplementary Table 4
389 SNPs dropped from main analysis based on the linkage disequilibrium criteria. (XLS 77 kb)
Supplementary Table 5
Results by sample (analysis from run #10). (XLS 435 kb)
Rights and permissions
About this article
Cite this article
Reich, D., Patterson, N., Jager, P. et al. A whole-genome admixture scan finds a candidate locus for multiple sclerosis susceptibility. Nat Genet 37, 1113–1118 (2005). https://doi.org/10.1038/ng1646
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1646
This article is cited by
-
Towards a global view of multiple sclerosis genetics
Nature Reviews Neurology (2022)
-
Two genetic variants explain the association of European ancestry with multiple sclerosis risk in African-Americans
Scientific Reports (2020)
-
Replication analysis of variants associated with multiple sclerosis risk
Scientific Reports (2020)
-
Genomics of disease risk in globally diverse populations
Nature Reviews Genetics (2019)
-
Uveitis and Multiple Sclerosis: Potential Common Causal Mutations
Molecular Neurobiology (2019)