Abstract
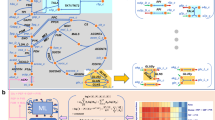
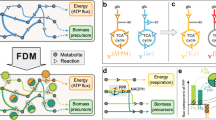
Qualitative theoretical approaches such as graph theory1,2 and stoichiometric analyses3,4,5,6 are beginning to uncover the architecture and systemic functions of complex metabolic reaction networks. At present, however, only a few, largely unproven quantitative concepts propose functional design principles of the global flux distribution7,8. As operational units of function, molecular fluxes determine the systemic cell phenotype by linking genes, proteins and metabolites to higher-level biological functions9. In sharp contrast to other 'omics' analyses, 'fluxome' analysis remained tedious10. By large-scale quantification of in vivo flux responses, we identified a robust flux distribution in 137 null mutants of Bacillus subtilis. On its preferred substrate, B. subtilis has suboptimal metabolism because regulators of developmental programs maintain a 'standby' mode that invests substantial resources in anticipation of changing environmental conditions at the expense of optimal growth. Network rigidity and robustness are probably universal functional design principles, whereas the standby mode may be more specific.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Arita, M. The metabolic world of Escherichia coli is not small. Proc. Natl. Acad. Sci. USA 101, 1543–1547 (2004).
Barabasi, A.L. & Oltvai, Z.N. Network biology: understanding the cell's functional organization. Nat. Rev. Genet. 5, 101–113 (2004).
Stelling, J., Klamt, S., Bettenbrock, K., Schuster, S. & Gilles, E.D. Metabolic network structure determines key aspects of functionality and regulation. Nature 420, 190–193 (2002).
Almaas, E., Kovacs, B., Vicsek, T., Oltvai, Z.N. & Barabasi, A.L. Global organization of metabolic fluxes in the bacterium Escherichia coli . Nature 427, 839–843 (2004).
Burgard, A.P., Nikolaev, E.V., Schilling, C.H. & Maranas, C.D. Flux coupling analysis of genome-scale metabolic network reconstructions. Genome Res. 14, 301–312 (2004).
Papin, J.A. et al. Comparison of network-based pathway analysis methods. Trends Biotechnol. 22, 400–405 (2004).
Segre, D., Vitkup, D. & Church, G.M. Analysis of optimality in natural and perturbed metabolic networks. Proc. Natl. Acad. Sci. USA 99, 15112–15117 (2002).
Edwards, J.S., Ibarra, R.U. & Palsson, B.O. In silico predictions of Escherichia coli metabolic capabilities are consistent with experimental data. Nat. Biotechnol. 19, 125–130 (2001).
Hellerstein, M.K. In vivo measurement of fluxes through metabolic pathways: the missing link in functional genomics and pharmaceutical research. Annu. Rev. Nutr. 23, 379–402 (2003).
Sauer, U. High-throughput phenomics: experimental methods for mapping fluxomes. Curr. Opin. Biotechnol. 15, 58–63 (2004).
Csete, M.E. & Doyle, J. Bow ties, metabolism and disease. Trends Biotechnol. 22, 446–450 (2004).
Fischer, E. & Sauer, U. A novel metabolic cycle catalyzes glucose oxidation and anaplerosis in hungry Escherichia coli . J. Biol. Chem. 278, 46446–46451 (2003).
Fischer, E. & Sauer, U. Metabolic flux profiling of Escherichia coli mutants in central carbon metabolism using GC-MS. Eur. J. Biochem. 270, 880–891 (2003).
Fischer, E., Zamboni, N. & Sauer, U. High-throughput metabolic flux analysis based on GC-MS derived 13C-constraints. Anal. Biochem. 325, 308–316 (2004).
Duetz, W.A. et al. Methods for intense aeration, growth, storage, and replication of bacterial strains in microtiter plates. Appl. Environ. Microbiol. 66, 2641–2646 (2000).
Zamboni, N. & Sauer, U. Knockout of the high-coupling cytochrome aa3 oxidase reduces TCA cycle fluxes in Bacillus subtilis . FEMS Microbiol. Lett. 226, 121–126 (2003).
Zamboni, N. et al. The Bacillus subtilis yqjI gene encodes the NADP+-dependent 6-P-gluconate dehydrogenase in the pentose phosphate pathway. J. Bacteriol. 186, 4528–4534 (2004).
Msadek, T. When going gets tough: survival strategies and environmental signaling networks in Bacillus subtilis . Trends Microbiol. 7, 201–207 (1999).
Servant, P., Le Coq, D. & Aymerich, S. CcpN (YqzB), a regulator for CcpA-independent catabolite repression of Bacillus subtilis gluconeogenic genes. Mol. Microbiol. 55, 1435–1451 (2005).
Sauer, U. et al. Metabolic fluxes in riboflavin-producing Bacillus subtilis . Nat. Biotechnol. 15, 448–452 (1997).
Moritz, B., Striegel, K., De Graaf, A.A. & Sahm, H. Kinetic properties of the glucose-6-phosphate and 6-phosphogluconate dehydrogenases from Corynebacterium glutamicum and their application for predicting pentose phosphate pathway flux in vivo . Eur. J. Biochem. 267, 3442–3452 (2000).
Zamboni, N. et al. Transient expression and flux changes during a shift from high to low riboflavin production in continuous cultures of Bacillus subtilis . Biotechnol. Bioeng. 89, 219–232 (2005).
Dauner, M., Storni, T. & Sauer, U. Bacillus subtilis metabolism and energetics in carbon-limited and carbon-excess chemostat culture. J. Bacteriol. 183, 7308–7317 (2001).
Sonenshein, A.L., Hoch, J.A. & Losick, R. Bacillus subtilis and its closest relatives. From genes to cells. (ASM Press, Washington, DC, 2002).
Pfeiffer, T., Schuster, S. & Bonhoeffer, S. Cooperation and competition in the evolution of ATP-producing pathways. Science 292, 504–507 (2001).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Ibarra, R.U., Edwards, J.S. & Palsson, B.O. Escherichia coli K-12 undergoes adaptive evolution to achieve in silico predicted optimal growth. Nature 420, 186–189 (2002).
Nudler, E. & Mironov, A.S. The riboswitch control of bacterial metabolism. Trends Biochem. Sci. 29, 11–17 (2004).
Stelling, J., Sauer, U., Szallasi, Z., Doyle III, F.J. & Doyle, J. Robustness of cellular functions. Cell 118, 675–685 (2004).
Dauner, M. & Sauer, U. Stoichiometric growth model for riboflavin-producing Bacillus subtilis . Biotechnol. Bioeng. 76, 132–143 (2001).
Acknowledgements
We thank S. Aymerich, S. Bonhoeffer, H Hennecke, T. Pfeiffer and J. Stelling for critical comments on the manuscript and S. Aymerich, K. Kobayashi and N. Ogasawara for providing B. subtilis mutants. This work was supported in part by the Roche Research Foundation (E.F.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Mutants used in this study. (XLS 37 kb)
Supplementary Table 2
Relative and absolute fluxes during aerobic batch growth on glucose in wild-type B. subtilis and the 137 viable mutants. (XLS 424 kb)
Rights and permissions
About this article
Cite this article
Fischer, E., Sauer, U. Large-scale in vivo flux analysis shows rigidity and suboptimal performance of Bacillus subtilis metabolism. Nat Genet 37, 636–640 (2005). https://doi.org/10.1038/ng1555
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1555
This article is cited by
-
Rational design of GDP‑d‑mannose mannosyl hydrolase for microbial l‑fucose production
Microbial Cell Factories (2023)
-
In silico-guided metabolic engineering of Bacillus subtilis for efficient biosynthesis of purine nucleosides by blocking the key backflow nodes
Biotechnology for Biofuels and Bioproducts (2022)
-
Understanding and mathematical modelling of cellular resource allocation in microorganisms: a comparative synthesis
BMC Bioinformatics (2021)
-
Addressing uncertainty in genome-scale metabolic model reconstruction and analysis
Genome Biology (2021)
-
Carbon flux through photosynthesis and central carbon metabolism show distinct patterns between algae, C3 and C4 plants
Nature Plants (2021)