Abstract
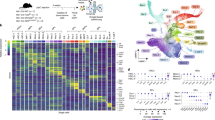
DNA methylation is associated with malignant transformation, but limitations imposed by genetic variability, tumor heterogeneity, availability of paired normal tissues and methodologies for global assessment of DNA methylation have limited progress in understanding the extent of epigenetic events in the initiation and progression of human cancer and in identifying genes that undergo methylation during cancer. We developed a mouse model of T/natural killer acute lymphoblastic leukemia that is always preceded by polyclonal lymphocyte expansion to determine how aberrant promoter DNA methylation and consequent gene silencing might be contributing to leukemic transformation. We used restriction landmark genomic scanning with this mouse model of preleukemia reproducibly progressing to leukemia to show that specific genomic methylation is associated with only the leukemic phase and is not random. We also identified Idb4 as a putative tumor-suppressor gene that is methylated in most mouse and human leukemias but in only a minority of other human cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Gilliland, D.G. & Tallman, M.S. Focus on acute leukemias. Cancer Cell 1, 417–420 (2002).
Jones, P.A. & Baylin, S.B. The fundamental role of epigenetic events in cancer. Nat. Rev. Genet. 3, 415–428 (2002).
Costello, J.F. et al. Aberrant CpG-island methylation has non-random and tumour-type-specific patterns. Nat. Genet. 24, 132–138 (2000).
Dai, Z. et al. Global methylation profiling of lung cancer identifies novel methylated genes. Neoplasia 3, 314–323 (2001).
Dai, Z. et al. An AscI boundary library for the studies of genetic and epigenetic alterations in CpG islands. Genome Res. 12, 1591–1598 (2002).
Smiraglia, D.J. et al. A new tool for the rapid cloning of amplified and hypermethylated human DNA sequences from restriction landmark genome scanning gels. Genomics 58, 254–262 (1999).
Yu, L. et al. A NotI-EcoRV promoter library for studies of genetic and epigenetic alterations in mouse models of human malignancies. Genomics 84, 647–660 (2004).
Chan, A.S. et al. Downregulation of ID4 by promoter hypermethylation in gastric adenocarcinoma. Oncogene 22, 6946–6953 (2003).
Rush, L.J. et al. Epigenetic profiling in chronic lymphocytic leukemia reveals novel methylation targets. Cancer Res. 64, 2424–2433 (2004).
Rush, L.J. et al. Novel methylation targets in de novo acute myeloid leukemia with prevalence of chromosome 11 loci. Blood 97, 3226–3233 (2001).
Andres-Barquin, P.J., Hernandez, M.C. & Israel, M.A. Id4 expression induces apoptosis in astrocytic cultures and is down-regulated by activation of the cAMP-dependent signal transduction pathway. Exp. Cell Res. 247, 347–355 (1999).
Fehniger, T.A. et al. Fatal leukemia in interleukin 15 transgenic mice follows early expansions in natural killer and memory phenotype CD8+ T cells. J. Exp. Med. 193, 219–231 (2001).
Costello, J.F., Smiraglia, D.J. & Plass, C. Restriction landmark genome scanning. Methods 27, 144–149 (2002).
Akama, T.O. et al. Restriction landmark genomic scanning (RLGS-M)-based genome-wide scanning of mouse liver tumors for alterations in DNA methylation status. Cancer Res. 57, 3294–3299 (1997).
Tateno, M. et al. Identification of a novel member of the snail/Gfi-1 repressor family, mlt 1, which is methylated and silenced in liver tumors of SV40 T antigen transgenic mice. Cancer Res. 61, 1144–1153 (2001).
Dallol, A. et al. Frequent epigenetic inactivation of the SLIT2 gene in gliomas. Oncogene 22, 4611–4616 (2003).
Dallol, A. et al. SLIT2, a human homologue of the Drosophila Slit2 gene, has tumor suppressor activity and is frequently inactivated in lung and breast cancers. Cancer Res. 62, 5874–5880 (2002).
Dallol, A., Morton, D., Maher, E.R. & Latif, F. SLIT2 axon guidance molecule is frequently inactivated in colorectal cancer and suppresses growth of colorectal carcinoma cells. Cancer Res. 63, 1054–1058 (2003).
Wang, B. et al. Induction of tumor angiogenesis by Slit-Robo signaling and inhibition of cancer growth by blocking Robo activity. Cancer Cell 4, 19–29 (2003).
Woronicz, J.D., Calnan, B., Ngo, V. & Winoto, A. Requirement for the orphan steroid receptor Nur77 in apoptosis of T-cell hybridomas. Nature 367, 277–281 (1994).
Liu, Z.G., Smith, S.W., McLaughlin, K.A., Schwartz, L.M. & Osborne, B.A. Apoptotic signals delivered through the T-cell receptor of a T-cell hybrid require the immediate-early gene nur77. Nature 367, 281–284 (1994).
Youn, H.D., Chatila, T.A. & Liu, J.O. Integration of calcineurin and MEF2 signals by the coactivator p300 during T-cell apoptosis. EMBO J. 19, 4323–4331 (2000).
Paine-Saunders, S., Viviano, B.L. & Saunders, S. GPC6, a novel member of the glypican gene family, encodes a product structurally related to GPC4 and is colocalized with GPC5 on human chromosome 13. Genomics 57, 455–458 (1999).
Filmus, J. Glypicans in growth control and cancer. Glycobiology 11, 19R–23R (2001).
Benezra, R. An intermolecular disulfide bond stabilizes E2A homodimers and is required for DNA binding at physiological temperatures. Cell 79, 1057–1067 (1994).
Jen, Y., Manova, K. & Benezra, R. Expression patterns of Id1, Id2, and Id3 are highly related but distinct from that of Id4 during mouse embryogenesis. Dev. Dyn. 207, 235–252 (1996).
Jen, Y., Weintraub, H. & Benezra, R. Overexpression of Id protein inhibits the muscle differentiation program: in vivo association of Id with E2A proteins. Genes Dev. 6, 1466–1479 (1992).
Kreider, B.L., Benezra, R., Rovera, G. & Kadesch, T. Inhibition of myeloid differentiation by the helix-loop-helix protein Id. Science 255, 1700–1702 (1992).
Shoji, W., Yamamoto, T. & Obinata, M. The helix-loop-helix protein Id inhibits differentiation of murine erythroleukemia cells. J. Biol. Chem. 269, 5078–5084 (1994).
Iavarone, A., Garg, P., Lasorella, A., Hsu, J. & Israel, M.A. The helix-loop-helix protein Id-2 enhances cell proliferation and binds to the retinoblastoma protein. Genes Dev. 8, 1270–1284 (1994).
Lasorella, A., Iavarone, A. & Israel, M.A. Id2 specifically alters regulation of the cell cycle by tumor suppressor proteins. Mol. Cell. Biol. 16, 2570–2578 (1996).
Lasorella, A., Uo, T. & Iavarone, A. Id proteins at the cross-road of development and cancer. Oncogene 20, 8326–8333 (2001).
Jen, Y., Manova, K. & Benezra, R. Each member of the Id gene family exhibits a unique expression pattern in mouse gastrulation and neurogenesis. Dev. Dyn. 208, 92–106 (1997).
Kee, Y. & Bronner-Fraser, M. Id4 expression and its relationship to other Id genes during avian embryonic development. Mech. Dev. 109, 341–345 (2001).
Bounpheng, M.A., Dimas, J.J., Dodds, S.G. & Christy, B.A. Degradation of Id proteins by the ubiquitin-proteasome pathway. FASEB J. 13, 2257–2264 (1999).
Welcsh, P.L. et al. BRCA1 transcriptionally regulates genes involved in breast tumorigenesis. Proc. Natl. Acad. Sci. USA 99, 7560–7565 (2002).
Beger, C. et al. Identification of Id4 as a regulator of BRCA1 expression by using a ribozyme-library-based inverse genomics approach. Proc. Natl. Acad. Sci. USA 98, 130–135 (2001).
Bellido, M. et al. Id4 is deregulated by a t(6;14)(p22;q32) chromosomal translocation in a B-cell lineage acute lymphoblastic leukemia. Haematologica 88, 994–1001 (2003).
Cheson, B.D. et al. National Cancer Institute-sponsored Working Group guidelines for chronic lymphocytic leukemia: revised guidelines for diagnosis and treatment. Blood 87, 4990–4997 (1996).
Wendel-Hansen, V. et al. Epstein-Barr virus (EBV) can immortalize B-cll cells activated by cytokines. Leukemia 8, 476–484 (1994).
Zhu, W.G. et al. Increased expression of unmethylated CDKN2D by 5-aza-2′-deoxycytidine in human lung cancer cells. Oncogene 20, 7787–7796 (2001).
Herman, J.G., Graff, J.R., Myohanen, S., Nelkin, B.D. & Baylin, S.B. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc. Natl. Acad. Sci. USA 93, 9821–9826 (1996).
Davuluri, R.V., Grosse, I. & Zhang, M.Q. Computational identification of promoters and first exons in the human genome. Nat. Genet. 29, 412–417 (2001).
Zhu, W.G., Lakshmanan, R.R., Beal, M.D. & Otterson, G.A. DNA methyltransferase inhibition enhances apoptosis induced by histone deacetylase inhibitors. Cancer Res. 61, 1327–1333 (2001).
Freund, R.J. & Wilson, W.J. Statistical Methods (Academic, San Diego, 1997).
Acknowledgements
We thank S. Liyanarachchi for statistical help. This work was supported in part by grants from the US Public Health Service (to C.P. and M.A.C.). C.P. and J.B. are Leukemia and Lymphoma Society of America Scholars.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Mouse RLGS master profile for enzyme combination NotI-EcoRV-HinfI. (PDF 4606 kb)
Rights and permissions
About this article
Cite this article
Yu, L., Liu, C., Vandeusen, J. et al. Global assessment of promoter methylation in a mouse model of cancer identifies ID4 as a putative tumor-suppressor gene in human leukemia. Nat Genet 37, 265–274 (2005). https://doi.org/10.1038/ng1521
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1521
This article is cited by
-
Methylation Mediated Downregulation of TOB1-AS1 and TOB1 Correlates with Malignant Progression and Poor Prognosis of Esophageal Squamous Cell Carcinoma
Digestive Diseases and Sciences (2023)
-
Downregulation of LINC00886 facilitates epithelial–mesenchymal transition through SIRT7/ELF3/miR-144 pathway in esophageal squamous cell carcinoma
Clinical & Experimental Metastasis (2022)
-
Aberrant methylation and downregulation of ZNF667-AS1 and ZNF667 promote the malignant progression of laryngeal squamous cell carcinoma
Journal of Biomedical Science (2019)
-
Aberrant hypermethylation-mediated downregulation of antisense lncRNA ZNF667-AS1 and its sense gene ZNF667 correlate with progression and prognosis of esophageal squamous cell carcinoma
Cell Death & Disease (2019)
-
Promoter hypermethylation-mediated downregulation of tumor suppressor gene SEMA3B and lncRNA SEMA3B-AS1 correlates with progression and prognosis of esophageal squamous cell carcinoma
Clinical & Experimental Metastasis (2019)