Abstract
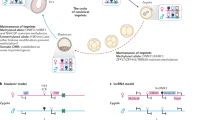
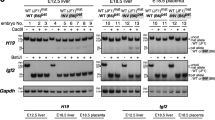
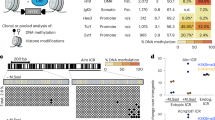
Imprinted genes are expressed from only one of the parental chromosomes and are marked epigenetically by DNA methylation and histone modifications1,2,3,4,5,6,7,8,9. The imprinting center 2 (IC2) on mouse distal chromosome 7 is flanked by several paternally repressed genes, with the more distant ones imprinted exclusively in the placenta. We found that most of these genes lack parent-specific DNA methylation, and genetic ablation of methylation does not lead to loss of their imprinting in the trophoblast (placenta). The silent paternal alleles of the genes are marked in the trophoblast by repressive histone modifications (dimethylation at Lys9 of histone H3 and trimethylation at Lys27 of histone H3), which are disrupted when IC2 is deleted, leading to reactivation of the paternal alleles. Thus, repressive histone methylation is recruited by IC2 (potentially through a noncoding antisense RNA) to the paternal chromosome in a region of at least 700 kb and maintains imprinting in this cluster in the placenta, independently of DNA methylation. We propose that an evolutionarily older imprinting mechanism limited to extraembryonic tissues was based on histone modifications, and that this mechanism was subsequently made more stable for use in embryonic lineages by the recruitment of DNA methylation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Reik, W. & Walter, J. Genomic imprinting: parental influence on the genome. Nat. Rev. Genet. 2, 21–32 (2001).
Ferguson-Smith, A.C. & Surani, M.A. Imprinting and the epigenetic asymmetry between parental genomes. Science 293, 1086–1089 (2001).
Sleutels, F. & Barlow, D.P. The origins of genomic imprinting in mammals. Adv. Genet. 46, 119–163 (2002).
Verona, R.I., Mann, M.R. & Bartolomei, M.S. Genomic imprinting: intricacies of epigenetic regulation in clusters. Annu. Rev. Cell. Dev. Biol. 19, 237–259 (2003).
Li, E., Beard, C. & Jaenisch, R. Role for DNA methylation in genomic imprinting. Nature 366, 362–365 (1993).
Bourc'his, D., Xu, G.L., Lin, C.S., Bollman, B. & Bestor, T.H. Dnmt3L and the establishment of maternal genomic imprints. Science 294, 2536–2539 (2001).
Kaneda, M. et al. Essential role for de novo DNA methyltransferase Dnmt3a in paternal and maternal imprinting. Nature 429, 900–903 (2004).
Grandjean, V., O'Neill, L., Sado, T., Turner, B. & Ferguson-Smith, A. Relationship between DNA methylation, histone H4 acetylation and gene expression in the mouse imprinted Igf2-H19 domain. FEBS Lett. 488, 165–169 (2001).
Fournier, C. et al. Allele-specific histone lysine methylation marks regulatory regions at imprinted mouse genes. EMBO J. 21, 6560–6570 (2002).
Howell, C.Y. et al. Genomic imprinting disrupted by a maternal effect mutation in the Dnmt1 gene. Cell 104, 829–838 (2001).
Shemer, R., Birger, Y., Riggs, A.D. & Razin, A. Structure of the imprinted mouse Snrpn gene and establishment of its parental-specific methylation pattern. Proc. Natl. Acad. Sci. USA 94, 10267–10272 (1997).
Dao, D. et al. Multipoint analysis of human chromosome 11p15/mouse distal chromosome 7: inclusion of H19/IGF2 in the minimal WT2 region, gene specificity of H19 silencing in Wilms' tumorigenesis and methylation hyper-dependence of H19 imprinting. Hum. Mol. Genet. 8, 1337–1352 (1999).
Caspary, T., Cleary, M.A., Baker, C.C., Guan, X.J. & Tilghman, S.M. Multiple mechanisms regulate imprinting of the mouse distal chromosome 7 gene cluster. Mol. Cell. Biol. 18, 3466–3474 (1998).
Tanaka, M. et al. Parental origin-specific expression of Mash2 is established at the time of implantation with its imprinting mechanism highly resistant to genome-wide demethylation. Mech. Dev. 87, 129–142 (1999).
Engemann, S. et al. Sequence and functional comparison in the Beckwith-Wiedemann region: implications for a novel imprinting centre and extended imprinting. Hum. Mol. Genet. 9, 2691–2706 (2000).
Fitzpatrick, G.V., Soloway, P.D. & Higgins, M.J. Regional loss of imprinting and growth deficiency in mice with a targeted deletion of KvDMR1. Nat. Genet. 32, 426–431 (2002).
Yatsuki, H. et al. Domain regulation of imprinting cluster in Kip2/Lit1 subdomain on mouse chromosome 7F4/F5: large-scale DNA methylation analysis reveals that DMR-Lit1 is a putative imprinting control region. Genome Res. 12, 1860–1870 (2002).
Lei, H. et al. De novo DNA cytosine methyltransferase activities in mouse embryonic stem cells. Development 122, 3195–3205 (1996).
Sado, T. et al. X inactivation in the mouse embryo deficient for Dnmt1: distinct effect of hypomethylation on imprinted and random X inactivation. Dev. Biol. 225, 294–303 (2000).
Umlauf, D et al. Imprinting along the Kcnq1 domain on mouse chromosome 7 involves repressive histone methylation and recruitment of Polycomb group complexes. Nat. Genet. advance online publication, 31 October 2004 (doi:10.1038/ngXXX).
Mager, J., Montgomery, N.D., de Villena, F.P. & Magnuson, T. Genome imprinting regulated by the mouse Polycomb group protein Eed. Nat. Genet. 33, 502–507 (2003).
Wutz, A. & Jaenisch, R. A shift from reversible to irreversible X inactivation is triggered during ES cell differentiation. Mol. Cell 5, 695–705 (2000).
Heard, E. Recent advances in X-chromosome inactivation. Curr. Opin. Cell. Biol. 16, 247–255 (2004).
Zwart, R., Sleutels, F., Wutz, A., Schinkel, A.H. & Barlow, D.P. Bidirectional action of the Igf2r imprint control element on upstream and downstream imprinted genes. Genes Dev. 15, 2361–2366 (2001).
Kanduri, C. et al. A differentially methylated imprinting control region within the Kcnq1 locus harbors a methylation-sensitive chromatin insulator. J. Biol. Chem. 277, 18106–18110 (2002).
Wilkins, J.F. & Haig, D. What good is genomic imprinting: the function of parent-specific gene expression. Nat. Rev. Genet. 4, 359–368 (2003).
Killian, J.K. et al. M6P/IGF2R imprinting evolution in mammals. Mol. Cell 5, 707–716 (2000).
Cleary, M.A. et al. Disruption of an imprinted gene cluster by a targeted chromosomal translocation in mice. Nat. Genet. 29, 78–82 (2001).
Paulsen, M. et al. Sequence conservation and variability of imprinting in the Beckwith-Wiedemann syndrome gene cluster in human and mouse. Hum. Mol. Genet. 9, 1829–1841 (2000).
Smilinich, N.J. et al. A maternally methylated CpG island in KvLQT1 is associated with an antisense paternal transcript and loss of imprinting in Beckwith-Wiedemann syndrome. Proc. Natl. Acad. Sci. USA 96, 8064–8069 (1999).
Acknowledgements
We thank E. Li for Dnmt1 knockout mice, Y. Zhang for antibodies, G. Fitzpatrick for help with the KvDMR1 knockout, F. Cerrato for help with allele-specific RT-PCR and F. Santos and G. Kelsey for advice and discussion. This work was supported by the Biotechnology and Biological Sciences Research Council and the Medical Research Council (to W.R. and W.D.) and by the Cancer Centre Support Grant and the National Cancer Institute (to M.H.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
CpG island methylation of IC2 imprinted gene cluster in mouse embryonic and extraembryonic cell lineages. (PDF 572 kb)
Rights and permissions
About this article
Cite this article
Lewis, A., Mitsuya, K., Umlauf, D. et al. Imprinting on distal chromosome 7 in the placenta involves repressive histone methylation independent of DNA methylation. Nat Genet 36, 1291–1295 (2004). https://doi.org/10.1038/ng1468
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1468
This article is cited by
-
Drug-induced loss of imprinting revealed using bioluminescent reporters of Cdkn1c
Scientific Reports (2023)
-
Orientation of mouse H19 ICR affects imprinted H19 gene expression through promoter methylation-dependent and -independent mechanisms
Communications Biology (2021)
-
Zfp57 inactivation illustrates the role of ICR methylation in imprinted gene expression during neural differentiation of mouse ESCs
Scientific Reports (2021)
-
Usefulness of methylation-specific multiplex ligation-dependent probe amplification for identification of parental origin of triploidy
Journal of Human Genetics (2020)
-
Mechanisms of early placental development in mouse and humans
Nature Reviews Genetics (2020)