Abstract
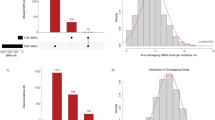
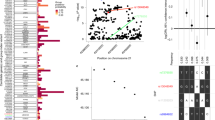
Previous studies have suggested more than 20 genetic intervals that are associated with susceptibility to type 1 diabetes (T1D)1,2, but identification of specific genes has been challenging and largely limited to known candidate genes. Here, we report evidence for an association between T1D and multiple single-nucleotide polymorphisms in 197 kb of genomic DNA in the IDDM5 interval. We cloned a new gene (SUMO4), encoding small ubiquitin-like modifier 4 protein, in the interval. A substitution (M55V) at an evolutionarily conserved residue of the crucial CUE domain of SUMO4 was strongly associated with T1D (P = 1.9 × 10−7). SUMO4 conjugates to IκBα and negatively regulates NFκB transcriptional activity. The M55V substitution resulted in 5.5 times greater NFκB transcriptional activity and ∼2 times greater expression of IL12B, an NFκB-dependent gene. These findings suggest a new pathway that may be implicated in the pathogenesis of T1D.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Onengut-Gumuscu, S. & Concannon, P. Mapping genes for autoimmunity in humans: type 1 diabetes as a model. Immunol. Rev. 190, 182–194 (2002).
Pociot, F. & McDermott, M.F. Genetics of type 1 diabetes mellitus. Genes Immun. 3, 235–249 (2002).
Twells, R.C. et al. Linkage and association mapping of the LRP5 locus on chromosome 11q13 in type 1 diabetes. Hum. Genet. 113, 99–105 (2003).
Twells, R.C. et al. The sequence and gene characterization of a 400-kb candidate region for IDDM4 on chromosome 11q13. Genomics 72, 231–242 (2001).
Nakagawa, Y. et al. Fine mapping of the diabetes-susceptibility locus, IDDM4, on chromosome 11q13. Am. J. Hum. Genet. 63, 547–556 (1998).
Eckenrode, S. et al. Fine-mapping of the type 1 diabetes locus (IDDM4) on chromosome 11q and evaluation of two candidate genes (FADD and GALN) by affected sibpair and linkage-disequilibrium analyses. Hum. Genet. 106, 14–18 (2000).
Luo, D.F. et al. Affected-sib-pair mapping of a novel susceptibility gene to insulin-dependent diabetes mellitus (IDDM8) on chromosome 6q25-q27. Am. J. Hum. Genet. 57, 911–919 (1995).
Luo, D.F. et al. Confirmation of three susceptibility genes to insulin-dependent diabetes mellitus: IDDM4, IDDM5 and IDDM8. Hum. Mol. Genet. 5, 693–698 (1996).
Owerbach, D. Physical and genetic mapping of IDDM8 on chromosome 6q27. Diabetes 49, 508–512 (2000).
Davies, J.L. et al. A genome-wide search for human type 1 diabetes susceptibility genes. Nature 371, 130–136 (1994).
Delepine, M. et al. Evidence of a non-MHC susceptibility locus in type I diabetes linked to HLA on chromosome 6. Am. J. Hum. Genet. 60, 174–187 (1997).
Jiang, Z., Ninomiya-Tsuji, J., Qian, Y., Matsumoto, K. & Li, X. Interleukin-1 (IL-1) receptor-associated kinase-dependent IL-1-induced signaling complexes phosphorylate TAK1 and TAB2 at the plasma membrane and activate TAK1 in the cytosol. Mol. Cell Biol. 22, 7158–7167 (2002).
Qian, Y., Commane, M., Ninomiya-Tsuji, J., Matsumoto, K. & Li, X. IRAK-mediated translocation of TRAF6 and TAB2 in the interleukin-1-induced activation of NFkappa B. J. Biol. Chem. 276, 41661–41667 (2001).
Takaesu, G. et al. TAB2, a novel adaptor protein, mediates activation of TAK1 MAPKKK by linking TAK1 to TRAF6 in the IL-1 signal transduction pathway. Mol. Cell 5, 649–658 (2000).
Takaesu, G. et al. Interleukin-1 (IL-1) receptor-associated kinase leads to activation of TAK1 by inducing TAB2 translocation in the IL-1 signaling pathway. Mol. Cell Biol. 21, 2475–2484 (2001).
Best, J.L. et al. SUMO-1 protease-1 regulates gene transcription through PML. Mol. Cell 10, 843–855 (2002).
Joseph, J., Tan, S.H., Karpova, T.S., McNally, J.G. & Dasso, M. SUMO-1 targets RanGAP1 to kinetochores and mitotic spindles. J. Cell Biol. 156, 595–602 (2002).
Melchior, F. & Hengst, L. SUMO-1 and p53. Cell Cycle 1, 245–249 (2002).
Rogers, R.S., Horvath, C.M. & Matunis, M.J. SUMO modification of STAT1 and its role in PIAS-mediated inhibition of gene activation. J. Biol. Chem. 278, 30091–30097 (2003).
Ross, S., Best, J.L., Zon, L.I. & Gill, G. SUMO-1 modification represses Sp3 transcriptional activation and modulates its subnuclear localization. Mol. Cell 10, 831–842 (2002).
Tian, S., Poukka, H., Palvimo, J.J. & Janne, O.A. Small ubiquitin-related modifier-1 (SUMO-1) modification of the glucocorticoid receptor. Biochem. J. 367, 907–911 (2002).
Desterro, J.M., Rodriguez, M.S. & Hay, R.T. SUMO-1 modification of IkappaBalpha inhibits NF-kappaB activation. Mol. Cell 2, 233–239 (1998).
Matunis, M.J. On the road to repair: PCNA encounters SUMO and ubiquitin modifications. Mol. Cell 10, 441–442 (2002).
Karin, M. How NF-kappaB is activated: the role of the IkappaB kinase (IKK) complex. Oncogene 18, 6867–6874 (1999).
May, M.J. & Ghosh, S. Signal transduction through NF-kappa B. Immunol. Today 19, 80–88 (1998).
Matsuda, A. et al. Large-scale identification and characterization of human genes that activate NF-kappaB and MAPK signaling pathways. Oncogene 22, 3307–3318 (2003).
Baldwin, A.S. Jr. The NF-kappa B and I kappa B proteins: new discoveries and insights. Annu. Rev. Immunol. 14, 649–683 (1996).
Deng, G.Y., Muir, A., Maclaren, N.K. & She, J.X. Association of LMP2 and LMP7 genes within the major histocompatibility complex with insulin-dependent diabetes mellitus: population and family studies. Am. J. Hum. Genet. 56, 528–534 (1995).
Spielman, R.S., McGinnis, R.E. & Ewens, W.J. Transmission test for linkage disequilibrium: the insulin gene region and insulin-dependent diabetes mellitus (IDDM). Am. J. Hum. Genet. 52, 506–516 (1993).
Wang, C.Y. et al. Molecular cloning and characterization of a novel gene family of four ancient conserved domain proteins (ACDP). Gene 306, 37–44 (2003).
Acknowledgements
We thank R.T. Hay for the NFκB-dependent luciferase reporter (3enh conA luc) and IκBα expression construct, R. McIndoe for suggestions about this project and J. Gu for discussion of manuscript preparation. This study was supported in part by grants from the Combined Intramural Grants Program, the Juvenile Diabetes Research Foundation (C.Y.W.) and the National Institute of Child Health and Development (J.X.S.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
LD of SNPs flanking the IDDM5 interval. (PDF 92 kb)
Supplementary Fig. 2
Pooled DNA sequencing results for the M55V of SUMO4. (PDF 103 kb)
Supplementary Fig. 3
Semi-quantitative Western analysis of IL-p40 in stimulated/unstimulated PBMC lysates from individuals studied in Figure 4. (PDF 123 kb)
Supplementary Table 1
Case-control association results for 001Msp. (PDF 53 kb)
Supplementary Table 2
Family-based association results for 001Msp. (PDF 90 kb)
Supplementary Table 3
Family-based association results for 268Hha. (PDF 89 kb)
Supplementary Table 4
Family-based association results for 012Taq. (PDF 89 kb)
Supplementary Table 5
Case-control association results for M55V. (PDF 44 kb)
Supplementary Table 6
Family-based association results for M55V. (PDF 89 kb)
Rights and permissions
About this article
Cite this article
Guo, D., Li, M., Zhang, Y. et al. A functional variant of SUMO4, a new IκBα modifier, is associated with type 1 diabetes. Nat Genet 36, 837–841 (2004). https://doi.org/10.1038/ng1391
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1391
This article is cited by
-
Mechanisms and functions of SUMOylation in health and disease: a review focusing on immune cells
Journal of Biomedical Science (2024)
-
The emerging roles of SUMOylation in pulmonary diseases
Molecular Medicine (2023)
-
DeSUMOylation of a Verticillium dahliae enolase facilitates virulence by derepressing the expression of the effector VdSCP8
Nature Communications (2023)
-
SUMO monoclonal antibodies vary in sensitivity, specificity, and ability to detect types of SUMO conjugate
Scientific Reports (2022)
-
SUMOylation targeting mitophagy in cardiovascular diseases
Journal of Molecular Medicine (2022)