Abstract
Transcription factors are believed to have a dominant role in acute myeloid leukemia (AML). This idea is supported by analysis of gene-knockout mice, which uncovered crucial roles of several transcription factors in normal hematopoiesis1, and of individuals with leukemia, in whom transcription factors are frequently downregulated or mutated2. However, analysis of knockout animals has not shown a direct link between abrogated transcription factors and the pathogenesis of AML. Sfpi1, encoding the lineage-specific transcription factor PU.1, is indispensable for normal myeloid and lymphoid development3,4. We found that mice carrying hypomorphic Sfpi1 alleles that reduce PU.1 expression to 20% of normal levels, unlike mice carrying homo- or heterozygous deletions of Sfpi1, developed AML. Unlike complete or 50% loss, 80% loss of PU.1 induced a precancerous state characterized by accumulation of an abnormal precursor pool retaining responsiveness to G-CSF with disruption of M- and GM-CSF pathways. Malignant transformation was associated with a high frequency of clonal chromosomal changes. Retroviral restoration of PU.1 expression rescued myeloid differentiation of mutant progenitors and AML blasts. These results suggest that tightly graded reduction, rather than complete loss, of a lineage-indispensable transcription factor can induce AML.
Similar content being viewed by others
Main
We previously identified a −14-kb upstream regulatory region (URE) in the Sfpi1 locus5. Here we used a cre-loxP-based homologous recombination strategy in mice to replace the URE with a neomycin-resistance cassette (Fig. 1a,b). Germline deletion of the URE after breeding the conditional mutants to CMV-cre deleter mice6 resulted in viable offspring at expected mendelian frequencies. Up to 8–10 weeks of age, these mice had no differences in size, behavior or reproductive ability compared with wild-type littermates. We found that PU.1 expression in the bone marrow of mice homozygous with respect to the ΔUREneo allele (referred to here as PU.1-knockdown mice) was reduced to 20% of wild-type levels (Fig. 1c–e). Thus, these mutants are an ideal in vivo model to study the consequences of low levels of PU.1.
(a) Map of Sfpi1 (encoding PU.1) indicating the location of the URE in DNase I–hypersensitive (HS) Cluster 1. Sfpi1 exons (filled boxes) and the direction of transcription (horizontal arrow) are symbolized. The targeting strategy is shown below. FRT sites are represented by open arrowheads and loxP sites by closed arrowheads. The orientations of the neo and tk cassettes are shown as open arrows. E, EcoRV restriction site; S, SspI restriction site. The location of the external probe A is indicated as a black horizontal bar. (b) Southern-blot analysis of EcoRV-SspI-digested mouse tail DNA. The DNA was hybridized with probe A, which recognizes a 9.7-kb wild-type fragment, a 4.7-kb loxP-targeted fragment and a 6.8-kb fragment indicating the knockdown allele (kd) that was generated after excision of the URE. (c) Northern blot showing downregulated Sfpi1 mRNA in bone marrow of Sfpi1kd/kd mice compared to Sfpi1+/+ littermates. Gapd was used as a loading control. (d) Quantitative real-time RT-PCR analysis of bone marrow RNA samples showed that Sfpi1 expression was reduced to 65% of wild-type level in heterozygotes (+/kd) and to 20% in preleukemic PU.1-knockdown (kd/kd) mice. Sfpi1 expression was also reduced in bone marrow of a PU.1-knockdown mouse with AML. Sfpi1 transcript numbers are shown as a percentage of 18S transcripts. (e) Western-blot analysis showing that PU.1 protein expression was five times lower in PU.1-knockdown mice than in wild-type controls. Total extracts equaling 106 cells from bone marrow (lanes 1–3) or fetal liver (lanes 4 and 5) were separated by SDS-PAGE, blotted onto a nitrocellulose membrane and probed with antibodies against PU.1 and actin. Lane 1, wild-type (+/+); lane 2, preleukemic PU.1-knockdown (kd/kd); lane 3, PU.1-knockdown with AML; lane 4, wild-type; lane 5, Sfpi1 knockout (−/−). Shown on the right is the position of a 40-kDa marker.
At 2–3 months of age, PU.1-knockdown mice had more c-kitpos precursor cells (2–2.5 times more) and neutrophils (Gr-1pos) than wild-type littermates (Fig. 2a,b,d). The mutants also had moderately enlarged spleens (1.5–2 times bigger). Expansion of the progenitor cells and neutrophils was limited to bone marrow and spleen: peripheral white blood cell counts were normal (average of 11,300 leukocytes per μl versus 9,800 leukocytes per μl in wild-type littermates, n = 4 in each group; Supplementary Table 1 online). Notably, 33 of 34 PU.1-knockdown mice rapidly became moribund or died at 3–8 months of age (Fig. 2c), after developing a fatal, aggressive neoplastic disease that strongly resembled human AML. Macroscopically, leukemic mice developed severe splenomegaly (weights: 660–1,600 mg versus 80–150 mg in wild-type littermates) and hepatomegaly (weights: 3.0–7.2 g versus 1–1.5 g in wild-type littermates). Flow cytometric examinations of moribund mutants showed a vast accumulation of immature myeloid cells (c-kitpos Mac-1low Gr-1low) in bone marrow and spleens (Fig. 2a,b). Both organs had grossly disrupted architectures and hypercellularity due to expansion of myeloblastic cells (30–75% blasts and promyelocytes in bone marrow; Fig. 2e,f). In contrast, there were fewer B-lymphoid and erythroid cells (data not shown). We detected T-cell lymphomas that developed together with the myeloid leukemia in three PU.1-knockdown mice; in another two mice, lymphoma was the dominant disease (data not shown). In mice with AML, peripheral white-blood-cell counts were much higher (average of 75,200 leukocytes per μl, n = 4) than in wild-type mice; normal components were replaced by >40% blasts and immature myeloid forms (Fig. 2g and Supplementary Table 1 online). Hematocrits and platelets were within the normal ranges (data not shown). The aggressive malignant disease course was characterized by massive neoplastic organ infiltrations (Fig. 2h–j) with most invading cells staining positive for myeloperoxidase, indicative of granulocytic origin of tumor cells (Fig. 2i,k).
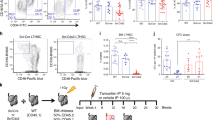
Flow cytometric analysis of single-cell suspensions from bone marrow (a) and spleen (b) of control and mutant mice before (preleukemic) and after (AML) the development of leukemia using indicated combinations of antibodies. Numbers in quadrants indicate the percentage of total cells. The results show a representative example of 4–6 mice per group. The presence of the c-kit antigen, as an early cell marker, on almost all Mac-1pos Gr-1pos myeloid cells indicates the immature phenotype of the neoplastic PU.1-knockdown (kd/kd) mice. (c) Cumulative survival of PU.1-knockdown (kd/kd) mice. n = 20 wild-type and 34 PU.1-knockdown mice. Mice were autopsied when visibly ill. No disease was observed in wild-type or heterozygous mice or in any mice before excision of the URE (which had the neomycin cassette incorporated in the locus). (d) Cytospin of spleen cells from a 3-month-old preleukemic PU.1-knockdown mouse showing the predominance of mature neutrophils (Wright-Giemsa stain, × 400). Cytospins of spleen (e) and bone marrow (f) cells from a PU.1-knockdown mouse with AML. The accumulation of blasts and myeloid precursors comprised almost the entire primary hematopoietic organs (Wright-Giemsa stain, × 400). (g) Blood smear showing large numbers of myeloblasts in the peripheral blood of a mutant after development of AML (Wright-Giemsa stain, ×400). (h) Massive neoblastic infiltration of the liver. Note the accumulation of myeloblasts in the blood vessel in the lower right corner (hematoxylin and eosin, ×100). (i) Peripheral lymph node densely populated with myeloid cells; most of the tumor cells stain positive for myeloperoxidase (×100). (j) Invasion of leukemia cells into kidney tissue (hematoxylin and eosin, ×200). (k) Neoplastic liver infiltration of a secondary recipient mouse after it was transplanted with cells from a leukemic PU.1-knockdown mouse (myeloperoxidase stain, ×100). (l) Engraftment of transplanted donor cells from leukemic PU.1-knockdown mice in nonirradiated NOD/SCID recipient mice led to complete displacement of host primary hematopoietic tissue. Diagnostic Southern blot of EcoRV-SspI-digested genomic DNA from mouse tails (controls: lane 1, wild-type (+/+); lane 2, heterozygous (+/kd); lane 3, homozygous knockdown (kd/kd)) and various organs from one recipient mouse (lane 4, bone marrow; lane 5, kidney; lane 6, tail; lane 7, spleen) using probe A.
To further test leukemogenicity, we intravenously injected 106 splenocytes from three PU.1-knockdown mice with AML into 12 nonirradiated NOD/SCID recipients. All the mice that received transplants developed progressive leukemia leading to death within 3–12 weeks. They had the same disease symptoms as donors did, including high peripheral white-blood-cell counts, hepatosplenomegaly and myeloblastic invasions of hematopoietic and nonhematopoietic organs (Fig. 2k and data not shown). Donor origin of leukemic cells in recipients was confirmed by Southern blots (Fig. 2l). We also transplanted the leukemia into normal nonirradiated wild-type mice. Despite decreased engraftment due to histocompatibility barriers, 3 of 12 transplanted noncongenic hosts developed lethal AML with massive progression of donor cells within 6–12 weeks (data not shown).
We found normal numbers of phenotypic hematopoietic stem cells (HSCs; c-kitpos Sca-1pos linneg) but an expanded progenitor compartment (c-kitpos Sca-1neg linneg) in preleukemic PU.1-knockdown mutants (Fig. 3a). Both HSCs and progenitors from PU.1-knockdown mice had reduced PU.1 expression (Fig. 3b). We could not separate mutant progenitors into distinct common myeloid progenitor, myeloid-erythroid progenitor and granulocyte-macrophage progenitor subpopulations7, suggestive of an abnormal progenitor compartment (Fig. 3a). Therefore, we sorted the combined progenitors as well as the HSCs to test differentiation potential in single-cell clonogenic assays. Erythroid and megakaryocytic colonies were normal, but myeloid differentiation was blocked at a blast-like stage under the conditions we used (Fig. 3c).
(a) Multiple color flow cytometry showing normal HSCs but accumulated progenitors in bone marrow of preleukemic PU.1-knockdown (kd/kd) mice. c-kitpos Sca-1pos linneg HSCs were phenotypically separated from c-kitpos Sca-1neg linneg progenitor cells, which were further split into common myeloid progenitors (c-kitpos Sca-1neg linneg FcgRII/IIIneg CD34pos), myeloid-erythroid progenitors (c-kitpos Sca-1neg linneg FcgRII/IIIneg CD34neg) and granulocyte-macrophage progenitors (c-kitpos Sca-1neg linneg FcgRII/IIIpos CD34pos). No differences were observed between wild-type and heterozygous (+/kd) mutant mice. (b) Multiplex of quantitative real-time RT-PCR using primers and probes specific for Sfpi1 (encoding PU.1) and 18S rRNA as a control. Sfpi1 (PU.1) transcript numbers are shown as percent of 18S transcripts in HSCs and progenitors. (c) Single cells from sorted HSCs and progenitors from wild-type and preleukemic PU.1-knockdown (kd/kd) bone marrow were cultured in the presence of 20 ng ml−1 of SCF, 10 ng ml−1 of TPO, 1 U ml−1 of EPO, 20 ng ml−1 of IL-3, 10 ng ml−1 of IL-11 and 10 ng ml−1 of GM-CSF. Colonies were counted and were identified by morphology and cytospins. The mixed populations from PU.1-knockdown mice contained normal erythroid and megakaryocytic cells in addition to myeloid blasts. (d) Normal myeloid differentiation induced with G-CSF, but not IL-3, IL-6 or SCF. Bone marrow cells from controls or preleukemic mutant mice were plated in methylcellulose containing either a mix of 10 ng ml−1 of IL-3, 10 ng ml−1 of IL-6 and 50 ng ml−1 of SCF or 10 ng ml−1 of G-CSF alone for 7 d. Individual colonies were picked and analyzed for cellular morphology. (e) Northern blot showing the absence of the mRNAs of the M-CSF receptor and the GM-CSF receptor α-chain, but not of the G-CSF receptor in preleukemic PU.1-knockdown mice. A total of 10 μg of total RNA from bone marrow of 2-month-old mice was loaded per sample and probed as indicated. (f) Myc overexpression is associated with the leukemic phase in PU.1-knockdown mice. Expression levels of Myc in bone marrow were assessed by Northern blots. Lane 1, wild-type control; lane 2, preleukemic PU.1-knockdown mutant; lanes 3–5, PU.1-knockdown mice that developed AML. (g) Protein expression (upper panel) and phosphorylation (lower panel) of c-Myc in bone marrow (lanes 1–3) and spleen (lanes 4–6) extracts from wild-type (lanes 1 and 4) mice, a preleukemic PU.1-knockdown (kd/kd) mouse (lane 2) and PU.1-knockdown mice with AML (lanes 3, 5 and 6). Myc was overexpressed and phosphorylated (Myc-P) in most (examples in lanes 3 and 6), but not all (lane 5), tumors. NS, a nonspecific band that also served as a loading control.
Clonogenic assays using total bone marrow cells showed that mutant cells underwent profound changes in response to myeloid cytokines (Table 1). Cells from PU.1-knockdown mice with AML produced large numbers of colonies, even in the absence of any cytokines, further demonstrating their transformed, growth factor–independent nature. Cells from preleukemic PU.1-knockdown mice almost completely lacked M- and GM-CSF response but produced normal colony numbers with G-CSF or a mixture of IL-3, IL-6 and SCF. Differentiation of preleukemic mutant cells in the presence of IL-3, IL-6 and SCF was again blocked at a myeloblast-like stage; G-CSF alone induced generation of neutrophils (Fig. 3d), providing an explanation for the enhanced granulopoiesis observed in vivo. To examine whether the altered cytokine response of cells from PU.1-knockdown mice could be a consequence of differential growth factor receptor expression, we carried out northern-blot analysis (Fig. 3e). Expression of M- and GM-CSF receptors was substantially reduced in PU.1-knockdown mice, but G-CSF receptor was expressed normally. These findings were unexpected, because PU.1-knockout mice express markedly less RNA of all three receptors8 and do not respond to M-, GM- or G-CSF9,10. Haploinsufficient expression of PU.1 in heterozygous Sfpi1+/− mice is not sufficient to induce defects in cytokine receptor expression and response9,10. Thus, a narrow window of reduced (to ∼20%) PU.1 expression seems to create the dosage that selectively preserves specific growth factor signaling and survival while leading to the disruption of the response to other cytokines.
To identify cooperative changes that contribute to leukemogenesis in PU.1-knockdown mice, we carried out spectral karyotyping (SKY) analysis and fluorescence in situ hybridization (FISH). We found simple clonal chromosomal abnormalities in five of seven mice with AML (Table 2). A gain of chromosome 15 was observed in four cases (Supplementary Figs. 1 and 2 and Supplementary Table 2 online), and one case showed an interstitial deletion of chromosome 3. Trisomy 15 is the most common abnormality in mouse T-cell lymphomas, as well as some mouse models of myeloid leukemia (reviewed in ref. 11). Trisomy 8 is the most common abnormality in human AML and is homologous to mouse chromosome 15. The smallest region gained is mouse 15D2-3 or human 8q24, containing the Myc proto-oncogene, which is often overexpressed in human tumors. Notably, expression and activation of c-Myc was elevated in six of eight PU.1-knockdown mice with AML, suggestive of a functional role for overexpression of c-Myc in PU.1-initiated neoplastic transformation (Fig. 3f,g).
To determine whether the reduction in PU.1 expression was directly responsible for the PU.1-knockdown phenotype, we restored PU.1 expression in mutant cells using a bicistronic retrovirus expressing PU.1 and green fluorescent protein (GFP). PU.1 expression by this retroviral construct correlates with GFP expression12, as confirmed by western blotting (data not shown). We transduced bone marrow from preleukemic PU.1-knockdown mice and separated GFP-positive (high PU.1 expression) from GFP-negative (hypomorphic PU.1 expression) cells. We assessed the myeloid differentiation potential of both fractions in the presence of IL-3, IL-6 and SCF by analyzing cell-surface marker expression (Fig. 4a) and cellular morphology (data not shown). The GFP-positive progenitors differentiated into mature macrophages (Gr-1neg Mac-1pos) and neutrophils (Gr-1high Mac-1high), similar to wild-type controls. In contrast, the GFP-negative population gave rise to immature granulocytic cells (Gr-1pos Mac-1neg-low).
(a) Bone marrow cells from preleukemic PU.1-knockdown (kd/kd) mice were transduced by a retrovirus expressing bicistronic PU.1iresGFP. Cells were then sorted by fluorescence-activated cell sorting for GFP expression, split into GFP-positive and GFP-negative fractions, cultured in IL-3, IL-6 and SCF for 7 d and analyzed for myeloid cell surface markers by flow cytometry. Wild-type control cells were mock-transduced and kept under same culturing conditions. (b) Bone marrow cells from control and preleukemic PU.1-knockdown mice were infected with retroviruses expressing either GFP alone or PU.1iresGFP. Clonogenic assays were done in the presence of GM-CSF. Numbers of mutant colonies after PU.1 restoration were comparable to those in control mice. (c) PU.1-knockdown colonies transfected with the PU.1 virus were GFP-positive and produced mature macrophages and granulocytes comparable to those of wild-type controls (see Fig. 3d). (d) Bone marrow of a PU.1-knockdown mouse suffering from AML was infected with the GFP control virus or with the PU.1iresGFP virus. GFP-positive cells were sorted by fluorescence-activated cell sorting, plated in methylcellulose containing 10 ng ml−1 of IL-3 and analyzed after 7 d. Colony shape is shown in left panels as phase-contrast images, and cellular morphology is shown in right panels by Wright-Giemsa stains. Colony numbers of control and PU.1-infected cells were equal.
We next examined whether restoration of PU.1 expression could rescue the response of preleukemic PU.1-knockdown progenitor cells to GM- and M-CSF (Fig. 4b,c; shown for GM-CSF conditions). Control cells infected with control or PU.1-expressing retrovirus formed comparable numbers of colonies in GM- or M-CSF, whereas mutant cells produced colonies only after transduction with the retrovirus expressing PU.1 and not with the control virus. Most notably, whereas transduction with control virus of bone marrow from PU.1-knockdown mice with AML led to cells that were arrested at a myeloblastic stage in IL-3, restoration of PU.1 expression in leukemic cells induced dispersed colonies of mature macrophages and granulocytes (Fig. 4d). Thus, reintroduction of PU.1 was sufficient to restore normal myeloid differentiation of leukemic PU.1-knockdown cells. These data indicate that downregulated expression of PU.1 is directly responsible for the disorder of the PU.1-knockdown mice and suggest that introduction of PU.1 into blasts of individuals with AML could be a therapeutic option to restore myeloid differentiation.
Our results show that reduced PU.1 expression induces AML in mice with high frequency and short latency. The importance of PU.1 concentrations on lineage fate decisions during normal hematopoiesis has been noted before12. Haploinsufficiency of PU.1 triggered granulocyte production in a G-CSF-null background13, in agreement with our observation of accelerated granulopoiesis in the presence of hypomorphic levels of PU.1. Like mice deficient in other transcription factors implicated in human AML1,14,15,16,17, however, neither nonconditional nor conditional deletion of the gene encoding PU.1 induces leukemia3,4,18. Because haploinsufficient PU.1 expression also does not induce a spontaneous malignancy19, a narrow window of reduced PU.1 expression seems to be required for AML development. In addition, humans with AML have mutations in genes encoding transcription factors that lead to reduced function but generally not complete loss of function2,20,21,22,23,24. Taken together, these data suggest that retaining residual transcription factor function can be crucial for AML development.
The mechanism behind the development of neoplasia in PU.1-knockdown mice probably includes the failure of the mutant cells to express sufficient levels of essential growth factor receptors. Whereas the complete absence of PU.1 globally abrogates growth factor signaling, however, reduction of PU.1 expression to 20% of normal levels preserves specific cytokine response (e.g., G-CSF and perhaps others) as an essential requirement for cell survival. This may be an important step in the development of a leukemic stem cell25. Malignant transformation was accompanied by chromosomal abnormalities with recurring gain of chromosome 15, containing the gene Myc. Myc is frequently overexpressed in human neoplasia26 and induces tumorigenesis in mice27,28. Thus, reduced expression of PU.1 promotes recurring chromosomal changes, creating a secondary event that results in the subsequent outgrowth of leukemic cell clones.
In conclusion, we showed that graded reduction of physiological PU.1 expression below a critical level can induce AML. Therefore, it might be possible to change cancerous cell fate considerably with relatively small modulations that increase lineage-determining transcription factors above this critical level.
Methods
Generation of the targeting construct.
We subcloned a 3.3-kb BamHI-KpnI genomic fragment containing the proximal part of the −14-kb URE plus the downstream homology into the BamHI-KpnI sites of pPNT-loxPneo. To add the distal part of the URE, we released a 1.7-kb SalI-BamHI fragment from a pBluescript plasmid containing the entire URE as a 3.4-kb HindIII fragment and inserted it into the SalI-BamHI sites of the targeting construct. This causes the original neomycin gene and the proximal loxP site to drop out of the construct. We then released a loxP site from pPNT-loxPneo with XbaI and SmaI and inserted it into the NheI site at the very 3′ end of the URE by blunt-end ligation. To obtain the upstream homology, we amplified a 2.2-kb fragment by PCR from a genomic mouse Sfpi1 P1 clone5 using primers with NotI or XhoI site overhangs. We then subcloned this fragment into the NotI-XhoI sites of pBluescript. We released a FRT-PGKneo-FRT resistance cassette from pK-11 (ref. 29) and subcloned it into SacI-SalI sites of pBluescript. After inserting an XhoI site adaptor into the NotI site of the plasmid, we released FRT-PGKneo-FRT by SalI and XhoI and subcloned it into the XhoI site of the pBluescript that already contained the upstream homology. We released another loxP site from pPNT-loxPneo using SalI-XhoI and inserted it into the XhoI site downstream of FRT-PGKneo-FRT. Finally, we subcloned the entire fragment containing the upstream homology, the FRT-flanked PGKneo and the loxP site into NotI-SalI sites of pPNT that already contained the URE-loxP-downstream homology fragments. This procedure created the final targeting construct with the elements in the following order: 2.2-kb upstream homology, FRT-PGKneo-FRT-loxP, 3.4-kb URE-loxP, 1.7-kb downstream homology, PGKHSK thymidine kinase gene. We linearized this construct with NotI and used it to transfect embryonic stem cells.
Generation of mutant mice.
We electroporated 30 μg of linearized targeting construct into R1 embryonic stem cells and grew them under double selection on feeder layers treated with mitomycin C. We digested DNA of individual double-resistant clones with SspI and EcoRV and probed them with the external probe A and the neomycin cDNA as an internal probe. We identified homologous recombinant clones by a 4.7-kb band representing −14-kb UREloxP+neo in addition to the 9.7-kb wild-type band in Southern blots using probe A. We generated chimeric mice by standard injection procedures, bred chimeric males to C57/Bl6 females and determined germline transmission by agouti color and Southern blots from tail genomic DNA. To remove the URE, we bred conditional mutants to CMV-cre deleter mice6. We identified heterozygous offspring with the proper deletions by Southern blotting. Incorporation of the neomycin cassette into the Sfpi1 locus (−14-kb UREloxP+neo) did not cause changes in expression of PU.1 or any other phenotypic changes before deletion of the URE. Furthermore, Southern-blot assays of cells derived from mice with AML had a pattern similar to that shown in Figure 1, indicating that the structure of the URE was not altered after transformation (data not shown). Mouse experiments were approved by the Beth Israel Deaconess Medical Center Institutional Animal Care and Use Committee (Protocol # 115-2001).
Northern-blot assays.
We isolated RNA from bone marrow cells using Trizol Reagent (Molecular Research Center) according to the manufacturer's protocol. We detected Sfpi1 (PU.1) RNA with a labeled 431-bp mouse Sfpi1 5′ cDNA fragment, Myc RNA with a probe containing exons 2 and 3 of the mouse gene and Gapd RNA with a 1.3-kb PstI fragment of the rat cDNA. Reduction of PU.1 expression in Northern (Fig. 1c) blots was determined using Imagequant densitometry software (Amersham).
Histologic and immunohistologic methods.
We carried out hematoxylin and eosin staining or immunohistochemical analysis for myeloperoxidase on 5-μm thick formalin-fixed, paraffin-embedded tissue sections. We removed the paraffin from the slides, pretreated them with 1.0 mM EDTA, pH 8.0 (Zymed), in a steam pressure cooker (Decloaking Chamber, BioCare Medical) following the manufacturer's instructions and washed them in distilled water. All further steps were done at room temperature in a hydrated chamber. We pretreated slides with Peroxidase Block (DAKO USA) for 5 min to quench endogenous peroxidase activity and then with a 1:5 dilution of goat serum in 50 mM Tris-Cl, pH 7.4, for 20 min to block nonspecific binding sites. We applied primary rabbit antibody to myeloperoxidase (DAKO) at a 1:2,500 dilution in 50 mM Tris-Cl, pH 7.4, with 3% goat serum for 1 h. We washed slides in 50 mM Tris-Cl, pH 7.4, and applied goat antibody to rabbit conjugated with horseradish peroxidase (Envision detection kit, DAKO) for 30 min. After further washing, we developed the immunoperoxidase staining using a DAB chromogen kit (DAKO) following the manufacturer's instructions and counterstained with hematoxylin. Blood films and cytospins were fixed with methanol and stained with Wright-Giemsa staining solutions.
Flow cytometry.
We analyzed single-cell suspensions from spleens and bone marrow by flow cytometry using unlabeled antibodies or antibodies conjugated to phycoerythrin, fluorescein isothiocyanate, Texas red or allophycocyanin (all from Pharmigen) against the following cell-surface molecules: Mac-1/CD11b (M1/70), CD3 (KT31.1), CD4 (GK1.5), CD8 (53-6.7), B220 (6B2), Gr-1 (8C5), TER119, CD19 (1D3), IgM (R6-60.2), IL-7Ra chain (A7R34), FcRII/IIIg (2.4G2), CD34 (RAM34), Sca-1 (E13-161-7) and c-Kit (2B8). We analyzed total bone marrow and spleen samples on a FACSscan cytometer (Becton Dickinson) according to standard protocols. Gates on viable cells were set according to the exclusion of propidium iodide staining. Flow cytometric analysis and sorting methods for HSCs and myeloid precursors using a double laser (488 nm/350 nm Enterprise II + 647nm Spectrum) high-speed cell sorter (Moflo-MLS, Cytomation) were previously described7.
Colony-forming assays.
We carried out in vitro clonogenic assays in Methocult M3234 (Stem Cell Technologies) supplemented with cytokines as indicated. We scored cells at 4–7 d. For myeloid precursor and HSC single-cell clonogenic assays, we used an automatic cell deposition unit system to sort cells into 96-well plates (Becton Dickinson). We cultured these highly purified cell populations in Methocult H4100 (Stem cell Technologies) supplemented with 20% fetal bovine serum, 1% bovine serum albumin, 2 mM L-glutamine and 50 mM 2-mercaptoethanol. We purchased recombinant human IL-6 and mouse M-CSF, G-CSF and GM-CSF from Stem Cell Technologies and mouse SCF, IL-3, IL-11 and TPO and human erythropoietin from R&D Systems.
Western-blot assays.
We extracted total cell lysates as described30. We resolved proteins by SDS-PAGE and electrotransferred them to a nitrocellulose membrane (Bio-Rad Laboratories). We used polyclonal rabbit antibody to PU.1 (Santa Cruz), polyclonal rabbit antibody to c-Myc (Santa Cruz), polyclonal rabbit antibody to phosphorylated c-Myc (Thr58/Ser62; Cell Signaling) and monoclonal mouse antibody to actin (Sigma). We detected immunoreactive proteins using HPRT-conjugated antibodies to rabbit or mouse (Santa Cruz) and the ECL system (Amersham). Reduction of PU.1 expression in Western blots was determined using Imagequant densitometry software (Amersham).
Real-time PCR.
We extracted RNA with Trizol, treated it with DNase I, reverse-transcribed it and then amplified it using an AbiPrism 7700 Sequence Detector (Applied Biosystems) with the following parameters: 48 °C (30 min), 95 °C (10 min) followed by 40 cycles of 95 °C (15 s), 60 °C (1 min). Primer and probe sequences are available on request.
SKY analysis.
We carried out cytogenetic and SKY analysis on spleen or bone marrow cells as described previously11.
Retroviral transduction of mouse bone marrow cells.
We transduced bone marrow cells with MIG and MIG-PU.1 retroviruses as described9. We cultured cells in IL-3, IL-6 and SCF or IL-3 alone for 2 d and then infected them by resuspension in high-titer cell-free retroviral supernatants for 4 h in the presence of 24 μg ml−1 of polybrene. We washed infected cells three times in complete medium and then cultured them for 2 d in complete medium containing IL-3, IL-6 and SCF or IL-3 alone to allow retroviral integration and expression. We washed infected cells three times before plating them in methylcellulose to assess colony-forming potential.
Note: Supplementary information is available on the Nature Genetics website.
References
Orkin, S.H. Diversification of haematopoietic stem cells to specific lineages. Nat. Rev. Genet. 1, 57–64 (2000).
Tenen, D.G. Disruption of differentiation in human cancer: AML shows the way. Nat. Rev. Cancer 3, 89–101 (2003).
Scott, E.W., Simon, M.C., Anastai, J. & Singh, H. The transcription factor PU.1 is required for the development of multiple hematopoietic lineages. Science 265, 1573–1577 (1994).
McKercher, S.R. et al. Targeted disruption of the PU.1 gene results in multiple hematopoietic abnormalities. EMBO J. 15, 5647–5658 (1996).
Li, Y. et al. Regulation of the PU.1 gene by distal elements. Blood 98, 2958–2965 (2001).
Schwenk, F., Baron, U. & Rajewsky, K. A cre-transgenic mouse strain for the ubiquitous deletion of loxP-flanked gene segments including deletion in germ cells. Nucleic Acids Res. 23, 5080–5081 (1995).
Akashi, K., Traver, D., Miyamoto, T. & Weissman, I.L. A clonogenic common myeloid progenitor that gives rise to all myeloid lineages. Nature 404, 193–197 (2000).
Iwama, A. et al. Use of RDA analysis of knockout mice to identify myeloid genes regulated in vivo by PU.1 and C/EBPα. Nucleic Acids Res. 26, 3034–3043 (1998).
DeKoter, R.P., Walsh, J.C. & Singh, H. PU.1 regulates both cytokine dependent proliferation and differentiation of granulocyte/macrophage progenitors. EMBO J. 17, 4456–4468 (1998).
Anderson, K.L. et al. Myeloid development is selectively disrupted in PU.1 null mice. Blood 91, 3702–3710 (1998).
Le Beau, M.M., Bitts, S., Davis, E.M. & Kogan, S.C. Recurring chromosomal abnormalities in leukemia in PML-RARA transgenic mice parallel human acute promyelocytic leukemia. Blood 99, 2985–2991 (2002).
DeKoter, R.P. & Singh, H. Regulation of B lymphocyte and macrophage development by graded expression of PU.1. Science 288, 1439–1441 (2000).
Dahl, R. et al. Regulation of macrophage and neutrophil cell fates by the PU.1:C/EBPalpha ratio and granulocyte colony-stimulating factor. Nat. Immunol. 4, 1029–1036 (2003).
Iwasaki-Arai, J. et al. C/EBPα deficiency in hematopoiesis induces accumulation of non-malignant myeloblasts mimicking acute myelogenous leukemia. Blood 100, 61a (2002).
Okuda, T., van Deursen, J., Hiebert, S.W., Grosveld, G. & Downing, J.R. AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis. Cell 84, 321–330 (1996).
Zhang, D.-E. et al. Absence of granulocyte colony-stimulating factor signaling and neutrophil development in CCAAT enhancer binding protein α-deficient mice. Proc. Natl. Acad. Sci. USA 94, 569–574 (1997).
Ichikawa, M. et al. AML-1 is required for megakaryocytic maturation and lymphocytic differentiation, but not for maintenance of hematopoietic stem cells in adult hematopoiesis. Nat. Med. 10, 299–304 (2004).
Duprez, E. et al. PU.1 deficiency blocks the production of Common Myeloid and Common Lymphoid Progenitors in both adult and fetal liver myelopoiesis. Blood 100, 61a (2002).
Scott, E.W. et al. PU.1 functions in a cell-autonomous manner to control the differentiation of multipotential lymphoid-myeloid progenitors. Immunity 6, 437–447 (1997).
Osato, M. et al. Biallelic and heterozygous point mutations in the runt domain of the AML1/PEBP2alphaB gene associated with myeloblastic leukemias. Blood 93, 1817–1824 (1999).
Pabst, T., Mueller, B.U., Harakawa, N., Zhang, D.-E. & Tenen, D.G. AML1-ETO downregulates the granulocytic differentiation factor C/EBPα in t(8;21) myeloid leukemia. Nat. Med. 7, 444–451 (2001).
Pabst, T. et al. Dominant negative mutations of CEBPA, encoding CCAAT/Enhancer Binding Protein-α (C/EBPα), in acute myeloid leukemia. Nat. Genet. 27, 263–270 (2001).
Mueller, B.U. et al. Heterozygous PU.1 mutations are associated with acute myeloid leukemia. Blood 100, 998–1007 (2002).
Wechsler, J. et al. Acquired mutations in GATA1 in the megakaryoblastic leukemia of Down Syndrome. Nat. Genet. 32, 148–152 (2002).
Passegue, E., Jamieson, C.H., Ailles, L.E. & Weissman, I.L. Normal and leukemic hematopoiesis: are leukemias a stem cell disorder or a reacquisition of stem cell characteristics? Proc. Natl. Acad. Sci. USA 100 Suppl 1, 11842–11849 (2003).
Marcu, K.B., Bossone, S.A. & Patel, A.J. myc function and regulation. Annu. Rev. Biochem. 61, 809–860 (1992).
Adams, J.M. et al. The c-myc oncogene driven by immunoglobulin enhancers induces lymphoid malignancy in transgenic mice. Nature 318, 533–538 (1985).
Felsher, D.W. & Bishop, J.M. Reversible tumorigenesis by MYC in hematopoietic lineages. Mol. Cell 4, 199–207 (1999).
Meyers, E.N., Lewandoski, M. & Martin, G.R. An Fgf8 mutant allelic series generated by Cre- and Flp-mediated recombination. Nat. Genet. 18, 136–141 (1998).
Kobayashi, S. et al. Calpain-mediated X-linked inhibitor of apoptosis degradation in neutrophil apoptosis and its impairment in chronic neutrophilic leukemia. J. Biol. Chem. 277, 33968–33977 (2002).
Acknowledgements
We thank K. Geary, J. Fields, S. Warner and the Dartmouth Transgenic Facility for assistance in generating and keeping the described mice; R. Cruz and A. Radhakrishnan for technical assistance with the SKY analysis; V. Petkova and T. Dayaram for help with the real-time PCRs; S. Dymecki and G. Martin (pK11-FRT-PGKneo-FRT), K.P. Knobeloch (pPNT-loxPneo) and H. Singh (MIG and MIG-PU.1) for constructs and advice; and J.D. Griffin, D.G. Gilliland, A.T. Look and S. Koschmieder for comments on the manuscript. This work was supported by DFG (German research foundation) research fellowships to F.R. and K.W. and by National Institutes of Health grants to M.M.L. and D.G.T.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Rosenbauer, F., Wagner, K., Kutok, J. et al. Acute myeloid leukemia induced by graded reduction of a lineage-specific transcription factor, PU.1. Nat Genet 36, 624–630 (2004). https://doi.org/10.1038/ng1361
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1361
This article is cited by
-
FANCA deficiency promotes leukaemic progression by allowing the emergence of cells carrying oncogenic driver mutations
Oncogene (2023)
-
PU.1-c-Jun interaction is crucial for PU.1 function in myeloid development
Communications Biology (2022)
-
CircSPI1 acts as an oncogene in acute myeloid leukemia through antagonizing SPI1 and interacting with microRNAs
Cell Death & Disease (2021)
-
GSK-3: a multifaceted player in acute leukemias
Leukemia (2021)
-
An order independent algorithm for inferring gene regulatory network using quantile value for conditional independence tests
Scientific Reports (2021)