Abstract
Population stratification refers to differences in allele frequencies between cases and controls due to systematic differences in ancestry rather than association of genes with disease. It has been proposed that false positive associations due to stratification can be controlled by genotyping a few dozen unlinked genetic markers. To assess stratification empirically, we analyzed data from 11 case-control and case-cohort association studies. We did not detect statistically significant evidence for stratification but did observe that assessments based on a few dozen markers lack power to rule out moderate levels of stratification that could cause false positive associations in studies designed to detect modest genetic risk factors. After increasing the number of markers and samples in a case-cohort study (the design most immune to stratification), we found that stratification was in fact present. Our results suggest that modest amounts of stratification can exist even in well designed studies.
Similar content being viewed by others
Main
There has been much debate1,2,3,4 but limited data5,6,7 about the impact of population stratification on case-control association studies. Systematic differences in the ancestry of cases and controls are one source of false positive associations8,9, but the fraction of published associations that is attributable to stratification is unknown10. It has been argued that the effects of stratification can be eliminated simply by carefully matching cases and controls according to self-reported ancestry and geographical origin2. Recently, empirical methods to detect stratification based on genotypes at unlinked markers have been described11. The largest application of such methods involved genotyping 44 unlinked markers in four case-control studies5. Stratification was detected in one study, although the signal was no longer apparent after more stringent matching of cases and controls based on the birthplaces of the individuals' grandparents. This has been interpreted as evidence that stratification may be less of a concern than originally anticipated.
We assessed stratification empirically by analyzing data from 24–48 unlinked single-nucleotide polymorphisms (SNPs) in 11 association studies spanning a range of disease states and self-reported ancestries and three different epidemiological designs. These studies included seven ongoing studies in our laboratory and reanalysis of data from the four studies previously reported5. We assessed stratification first by testing for statistically significant evidence of differentiation between cases and controls using the method of Pritchard and Rosenberg11 and second by estimating the magnitude of stratification consistent with the data using Genomic Control12,13.
None of the 11 studies showed significant evidence of stratification after correcting for multiple hypothesis testing, consistent with previous studies5,6 (Table 1). Comparing cases and controls from different studies with the same self-reported ancestry (European American), we found no significant evidence of stratification in nine pairwise comparisons using 33–43 SNPs (Supplementary Table 1 online).
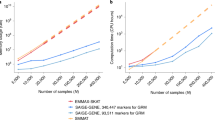
We next applied the method of Genomic Control12,13 to estimate quantitatively the amount of stratification consistent with the data for each of the 11 studies. Genomic Control is conceptually simple: the method examines the distribution of association statistics (χ2) between unlinked genetic variants typed in cases and controls. The statistic at a candidate allele being tested for association can then be compared with the genome-wide distribution of statistics for markers that are probably unrelated to disease to assess whether the candidate allele stands out. In the absence of stratification, association between unlinked genetic variants and disease should follow a χ2 distribution with 1 degree of freedom12,13. In the presence of stratification, the distribution of association statistics should be inflated by a value termed λ, which becomes larger with increasing of sample size (Fig. 1).
(a) Stratification inflates χ2 association statistics by a factor λ, which changes depending on the sample size. Scenario 1 corresponds to gross stratification; scenarios 2 and 3 correspond to the range of stratification estimated in the African American prostate cancer study; and scenario 4 corresponds to no stratification. (b) Comparison of the nominal P values with those corrected for stratification shows that stratification that is difficult to detect in a study of hundreds of cases and controls can cause many false positive signals in a study of thousands of samples.
We estimated stratification for each of the 11 data sets and report the inflation of association statistics that would be expected in a study of 1,000 cases and 1,000 controls, called λ1000. (It is simple to extrapolate from λ1000 to the inflation factor due to stratification for any sample size12.) Consistent with the fact that the 11 data sets showed no significant evidence for stratification, the confidence intervals for λ1000 overlapped 1 in every study. Nevertheless, we found that the confidence intervals were sufficiently broad that substantial levels of stratification could not be excluded. For example, the 95th percentile upper bound on λ1000 in the studies averaged 7.9 (Table 1 and Fig. 2).
The upper bound on the level of stratification can be obtained from the figures as the point where the likelihood drops to 4.5% of its maximum, which is a log-likelihood criterion for a 95th percentile upper bound (one-sided test). With the handful of SNPs genotyped initially (24–48), the likelihood distribution is broad. Although no studies show significant stratification, all are consistent with levels of stratification that could produce notable numbers of false positives. Increasing the number of SNPs and samples can tighten the estimate of population stratification. This is shown for the African American and European American prostate cancer studies, for which we provide both an initial estimate of stratification based on the noncoding SNPs and a more precise estimate based on an expanded sample size and inclusion of missense SNPs.
We increased power to detect stratification by increasing the number of SNPs and samples examined in one of the 11 studies that initially showed no significance evidence for stratification, the African American prostate cancer study (P < 0.75). For the follow-up, we approximately quadrupled the number of markers and increased the sample size by a factor of 5–6 (474 prostate cancer cases and 476 cohort controls). The new markers consisted of a collection of missense SNPs, which we treated as being in the same class as the noncoding SNPs, because, within the limits of our resolution (Table 2), they showed the same levels of population differentiation (with sufficient power, such differences can probably be detected14). The new markers also included a second set of SNPs chosen for their large allele frequency differences between west Africans and Europeans15, which makes them particularly powerful for detecting stratification16.
In this expanded data set we found significant evidence of stratification (P < 0.0001). When we restricted the analysis to 469 cases and 268 controls in whom all markers were successfully typed, the result was still significant (P < 0.01; Table 1). We then removed from the analysis 40 cases and 48 controls who reported that either they or their parents had some non-African American ancestry2,5, because a small number of individuals with misclassified ancestry might disproportionately affect the result. The evidence for stratification was stronger in this subset (P < 0.005; Table 1). Notably, the Genomic Control estimate of stratification (removing SNPs that had been specifically chosen to have large differences in frequency across populations17) was λ1000 = 1.5, with a 95th percentile upper bound of 3.34. This indicates that an observation of χ2 = 19.5, expected only once by chance in a scan of 100,000 SNPs, would instead be seen 31 times (effective χ2 = 19.5/1.5 = 13) due to this level of stratification. At the 95% upper confidence limit of our estimate (λ1000 = 3.34), 1,568 false positives would be expected due to stratification.
The observation of population stratification in African Americans with prostate cancer is not entirely unexpected. People of west African descent are thought to have a higher genetic risk for prostate cancer than those of European descent18, and hence African Americans with prostate cancer, who are known to have ancestry from both populations15, might be expected to have more African ancestry, on average, than controls. Population stratification was also observed in a separate study of African Americans with prostate cancer9. Our analysis strengthens this result, in that our sample was prospectively collected in a population-based cohort19, considered to be the optimal epidemiological design to minimize systematic differences between cases and controls (as opposed to the case-control design).
We also followed up with a study of European Americans with prostate cancer (approximately doubling the number of SNPs to 79 and quadrupling the number of samples to 391 cases and 456 cohort controls). In this study, we did not find statistically significant evidence for stratification (P < 0.10). The 95th percentile upper bound on stratification from Genomic Control, however, was similar to that in the study of African Americans (λ1000 = 3.03; Fig. 2). Much more data will be needed from many studies before it is possible to assess whether matching cases and controls solely on the basis of their self-reported ancestry, in a population such as European Americans without recent mixture, is adequate to take into account population stratification.
Our data indicate that genotyping a few dozen markers cannot rule out modest levels of population stratification that could generate false positives in an association study designed to detect alleles of weak effect—even in the setting of a prospectively collected cohort study. Stratification is probably most problematic in populations whose ancestors recently mixed due to intercontinental migrations and for diseases that have different prevalence rates across these ancestral populations11,13 (such as hypertension, obesity, diabetes and autoimmunity). Because the importance of stratification grows with sample size12,13, however, it seems possible that, even for diseases whose incidence rates are not currently known to vary across populations, stratification could exist. Thus, our study argues that stratification cannot be excluded based on either first principles or published empirical data. We suggest instead that investigators continue to monitor for stratification. In addition to presenting nominal P values, investigators should also report the range of values consistent with the Genomic Control estimate of stratification in the samples based on genotyping unlinked markers. Alternatively, investigators could present a P value corrected for the full range of possible values of λ1000, using the full Bayesian approach to Genomic Control12.
Our data show that stratification cannot be excluded as a possibility in real case-control studies, but that there is no need to abandon case-control and case-cohort studies in favor of family-based designs (such as transmission disequilibrium tests). Two powerful approaches are available to detect and correct for stratification20. The first clusters samples based on multilocus genotypes (e.g., STRUCTURE21) to identify individuals with different ancestries. This provides a way to adjust for ancestry as a covariate in the association analysis7,21. Genomic Control, on the other hand, makes a quantitative estimate of the degree of stratification and uses it to adjust for any stratification that might be present. The two methods are not mutually exclusive: STRUCTURE can be used first to identify and eliminate samples that contribute unduly to stratification, and a smaller Genomic Control correction can then be made in the final study.
How many SNPs need to be used in an assessment of stratification? This question must be viewed in relation to the magnitude of genetic effects under study. Given a substantial magnitude of effect and a highly significant P value, only a few dozen markers probably need to be genotyped to rule out gross stratification as an explanation for the positive association11,22 (Table 3). In contrast, if the results point to more modest influences on disease, such as the risk due to variation in CTLA4 on autoimmune thyroid disease and type 1 diabetes23, it may be necessary to genotype a larger number of markers to rule out modest amounts of stratification. Genotyping more than 340 markers can bring the conservative 95th percentile upper bound on the level of stratification to within 10% of the true value (Table 3). Fortunately, as the number of SNPs tested in association studies grows larger (to survey the genome for risk-associated alleles of increasingly modest effect), the bounds on the estimate of stratification should become increasingly precise with no additional effort, as all the markers in a study can be used to assess and adjust for stratification12,13.
Methods
Clinical samples.
We obtained all samples for the new data collections with permission of the principal investigators and with approval of the Institutional Review Boards of the Massachusetts General Hospital, the Cleveland Clinic, SUNY/Upstate Medical University, the University of Hawaii and the University of Southern California. Informed consent was obtained from all subjects by the institutions responsible for the collections. Citations provide additional detail on the ascertainment of cases and controls.
GeneQuest coronary artery disease study24.
We randomly selected 83 cases and 80 controls, all European Americans. The cases were from Cleveland, and controls were identified by random digit phone dialing in Atlanta, Georgia, USA.
Multiethnic Cohort prostate cancer study.
The Multiethnic Cohort19 is an ongoing (n = 215,251) study focusing on the effects of diet, genes and environment on the risk of cancer. The cohort samples include four main ethnic groups in Los Angeles and Hawaii. For European Americans, we randomly selected 110 incident cases and 97 cohort controls; for African Americans, we selected 90 incident cases and 69 cohort controls; for Japanese Americans, we selected 121 incident cases and 106 cohort controls; and for Hispanic Americans, we selected 142 incident cases and 124 cohort controls. We followed up in the study of African American prostate samples by genotyping all the missense and ancestry-informative SNPs described below. We genotyped an expanded sample of 469 African-American incident cases and 268 cohort controls from the cohort for all the SNPs and genotyped an additional 5 cases and 208 cohort controls for 31 of the SNPs that had high allele frequency differences across populations before the DNA for these samples ran out.
Bipolar disorder in European Americans.
We obtained 93 DNA samples from Massachusetts General Hospital from individuals with diagnoses of bipolar disorder 1 or bipolar disorder 2. As controls, we used GeneQuest samples (both this and the coronary artery disease study are examples where cases are matched to controls only using self-reported ancestry.)
Schizophrenia.
We obtained samples from 149 cases diagnosed with schizophrenia according to the criteria of the Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition, and 152 matched controls as part of a study of schizophrenia in the Portuguese population. Samples were descended from continental Portugal (83% of cases, 87% of controls), the Azore islands (13% of cases, 3% of controls) or the Madeira islands (3% of cases, 10% of controls). Some individuals were from Fall River, Rhode Island, but in each of these cases, all four grandparents were from the Azore islands.
For comparison of noncoding to missense SNPs, we studied 50 African American, 88 European American and 42 Asian American population samples. These were identical to those previously studied25, except that the 88 European American samples were replaced by the parents of the 44 samples resequenced previously26.
Choice of markers.
The physical and genetic map positions, along with flanking sequences, of all SNPs used in this study are available from the authors on request.
We obtained noncoding SNPs (67) from the SNP Consortium website. They were identified by comparing a single sequencing read from a diverse panel of individuals with the publicly available genome sequence27. The SNPs were evenly spaced throughout the autosomes, each at least 20 Mb from the others. In practice, only 34–48 of these SNPs genotyped successfully and were of high enough frequency in any study to use in our analysis (the expected number of reference and variant alleles based on the allele frequency and sample size was ≥5).
We identified missense SNPs (100) from a database of SNPs in coding regions of genes28, obtained as part of an effort to catalog SNPs in genes of interest for disease. We used only genes that were not designated in the database or in a published meta-analysis10 as having any relationship with prostate cancer, coronary artery disease, asthma or atopy. We excluded from the study those SNPs with a minor allele frequency <10% in a multiethnic screening panel. SNPs were chosen to be at least 1 Mb away from each other and from all the noncoding SNPs.
We obtained ancestry-informative SNPs (101) with high allele frequency differences comparing European and African Americans by combining data from ref. 15 with unpublished data from our own laboratory. These SNPs were all chosen to be at least 20 Mb from each other. The average frequency difference comparing west Africans and Europeans was 67%.
Genotyping.
The genotypes collected for this study are available from the authors to the extent that is consistent with the informed consent provided by the study participants. We used matrix-associated laser desorption ionization–time of flight mass spectrometry (MALDI-TOF)29 with 5 ng of DNA per multiplex genotyping reaction to genotype most SNPs in this study. The PCR protocol is described elsewhere25. Error rates with the Sequenom MassARRAY system have been estimated to be ∼0.4% at our laboratory25, although the discrepancy rate in the present data set suggests closer to 0.25% (215 conflicts out of 42,766 genotypes, each done at least in duplicate).
Elimination of poorly performing SNPs.
We removed all SNPs from our analysis that showed Hardy Weinberg P values of <0.01 in at least two of the three diversity samples (CEPH, East Asian and African American). We also excluded SNPs from the analysis if the combined Hardy-Weinberg P value, over all populations excluding African Americans and Hispanic Americans, was <0.01. To calculate the P value, we summed the χ2 values for the Hardy-Weinberg test over all n populations for which the statistic could be calculated and assessed significance using a χ2 distribution with n degrees of freedom. (We excluded African Americans and Hispanic Americans from the Hardy-Weinberg assessment because different levels of population mixture across individuals in these groups can produce a deficiency of heterozygotes, even with accurate genotyping.) We also excluded from analysis SNPs for those studies in which the genotyping success rates were <75% in either cases or controls25. We also eliminated from analysis SNPs that showed discrepancy rates of >3% in duplicate genotypes.
Detection of population stratification.
We calculated χ2 association statistics for all k SNPs in a study, including only those for which the expected number of allele counts (based on the combined frequency in the two population samples) was at least 5. We then summed the values and assessed significance using a χ2 distribution with k degrees of freedom11.
Quantitative assessment of population stratification.
For each SNP in each study for which at least 40% of the cases and controls had been successfully genotyped, we calculated χ2 values for all SNPs for which the expected number of allele counts (based on the combined frequency in the two population samples) was at least 5.
We carried out a likelihood analysis to estimate the level of stratification consistent with the data in each study. Defining cj as the association statistic observed at marker j genotyped in nj cases and mj controls and f as the χ2 distribution with 1 degree of freedom, the likelihood of a given inflation factor due to stratification is simply

a consequence of the fact that the χ2 distribution scales with the inflation factor12. The likelihood at all K markers is then

To estimate a likelihood distribution for the level of stratification, we define a reference sample size (we use nref = 1,000 cases and mref = 1,000 controls). We then use an equation derived in ref. 12 and confirmed by simulation as in ref. 13 to relate this to the inflation factor applicable to nj cases and mj controls. The inflation factor should be different from marker to marker because it scales with sample size:

In this paper we abbreviate λ1000,1000 as λ1000.
Substituting equation 3 into equation 2 allows us to obtain a likelihood distribution for λ1000. The maximum likelihood estimate for λ1000 is simply the value for which L is maximized, with the requirement that λ1000 ≥ 1. We obtained the likelihood surfaces shown in Figure 2 by plotting the values of L for different λ1000, normalizing by the maximum likelihood (set equal to 1 in Fig. 2). We obtained the upper bound on λ1000 by picking the value such that the likelihood ratio 2log10(Lmax/L) = 2.7; that is, the point for which the likelihood was 4.5% of the maximum, corresponding roughly to a P < 0.05 cutoff (one-sided test).
To test for a difference in the distribution of χ2 values between missense and noncoding SNPs (Table 2), we compared the random African American, European American and Asian American population samples in our study. For each SNP for which at least 70% both sample sets had been successfully genotyped, we randomly dropped samples until we had the same number at all sites. We then calculated χ2 values and used a Mann-Whitney U test to assess whether the empirical distributions of statistics at missense and noncoding SNPs were distinguishable.
URL.
The SNP Consortium website is available at http://snp.cshl.org.
Note: Supplementary information is available on the Nature Genetics website.
References
Thomas, D.C. & Witte, J.S. Point: population stratification: a problem for case-control studies of candidate-gene associations? Cancer Epidemiol. Biomarkers Prev. 11, 505–512 (2002).
Wacholder, S., Rothman, N. & Caporaso, N. Counterpoint: bias from population stratification is not a major threat to the validity of conclusions from epidemiological studies of common polymorphisms and cancer. Cancer Epidemiol. Biomarkers Prev. 11, 513–520 (2002).
Ziv, E. & Burchard, E.G. Human population structure and genetic association studies. Pharmacogenomics 4, 431–441 (2003).
Cardon, L.R. & Palmer, L.J. Population stratification and spurious allelic association. Lancet 361, 598–604 (2003).
Ardlie, K.G., Lunetta, K.L. & Seielstad, M. Testing for population subdivision and association in four case-control studies. Am. J. Hum. Genet. 71, 304–311 (2002).
Schork, N.J. et al. The future of genetic case-control studies. Adv. Genet. 42, 191–212 (2001).
Hoggart, C.J. et al. Control of confounding of genetic associations in stratified populations. Am. J. Hum. Genet. 72, 1492–1504 (2003).
Knowler, W.C., Williams, R.C., Pettitt, D.J. & Steinberg, A.G. Gm3;5,13,14 and type 2 diabetes mellitus: an association in American Indians with genetic admixture. Am. J. Hum. Genet. 43, 520–526 (1988).
Kittles, R.A. et al. CYP3A4-V and prostate cancer in African Americans: causal or confounding association because of population stratification? Hum. Genet. 110, 553–560 (2002).
Lohmueller, K.E., Pearce, C.L., Pike, M., Lander, E.S. & Hirschhorn, J.N. Meta-analysis of genetic association studies supports a contribution of common variants to susceptibility to common disease. Nat. Genet. 33, 177–182 (2003).
Pritchard, J.K. & Rosenberg, N.A. Use of unlinked genetic markers to detect population stratification in association studies. Am. J. Hum. Genet. 65, 220–228 (1999).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Reich, D.E. & Goldstein, D.B. Detecting association in a case-control study while correcting for population stratification. Genet. Epidemiol. 20, 4–16 (2001).
Akey, J.M., Zhang, G., Zhang, K., Jin, L. & Shriver, M.D. Interrogating a high-density SNP map for signatures of natural selection. Genome Res. 12, 1805–1814 (2002).
Parra, E.J. et al. Estimating African American admixture proportions by use of population-specific alleles. Am. J. Hum. Genet. 63, 1839–1851 (1998).
Pfaff, C.L., Kittles, R.A. & Shriver, M.D. Adjusting for population structure in admixed populations. Genet. Epidemiol. 22, 196–201 (2002).
Reich, D.E. & Goldstein, D.B. Response to Pfaff et al.: Adjusting for population structure in admixed populations. Genet. Epidemiol. 22, 196–201 (2002).
Bunker, C.H. et al. High prevalence of screening-detected prostate cancer among Afro-Caribbeans: the Tobago prostate cancer survey. Cancer Epidemiol. Biomarkers Prev. 11, 726–729 (2002).
Kolonel, L.N. et al. A multiethnic cohort in Hawaii and Los Angeles: baseline characteristics. Am. J. Epidemiol. 151, 346–357 (2000).
Pritchard, J.K. & Donnelly, P. Case-control studies of association in structured or admixed populations. Theor. Popul. Biol. 60, 227–237 (2001).
Pritchard, J.K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Siddiqui, A. et al. Association of multidrug resistance in epilepsy with a polymorphism in the drug-transporter gene ABCB1. N. Engl. J. Med. 348, 1442–1448 (2003).
Ueda, H. et al. Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature 423, 506–511 (2003).
Topol, E.J. et al. Single nucleotide polymorphisms in multiple novel thrombospondin genes may be associated with familial premature myocardial infarction. Circulation 104, 2641–2644 (2001).
Gabriel, S.B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002).
Reich, D.E. et al. Linkage disequilibrium in the human genome. Nature 411, 199–204 (2001).
Sachidanandam, R. et al. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409, 928–933 (2001).
Cargill, M. et al. Characterization of single-nucleotide polymorphisms in coding regions of human genes. Nat. Genet. 22, 231–238 (1999).
Tang, K. et al. Chip-based genotyping by mass spectrometry. Proc. Natl. Acad. Sci. USA 96, 10016–10020 (1999).
Acknowledgements
We thank A. Villapakkam for assistance in genotyping and data checking and K. Ardlie, D. Goldstein and C. Haiman for discussions. M.L.F. is supported by a Department of Defense Health Disparity training grant and a Postdoctoral Fellowship for Physicians from the Howard Hughes Medical Institute. D.A. is a Clinical Scholar in Translational Research from the Burroughs Wellcome Fund, as well as a Charles E. Culpeper Medical Scholar. J.N.H. and D.R. are recipients of Career Development Awards from the Burroughs Welcome Fund. T.L.P. is supported by a Canadian Institutes of Health Research Postdoctoral Fellowship and is a NARSAD Young Investigator. This work was supported in part by a grant from the Functional Genomics Program at the Whitehead Institute/MIT Center for Genome Research (supported by Affymetrix, Millennium Pharmaceuticals and Bristol Myers Squibb) and by a grant from the US National Institutes of Health to B.H. and L.K.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
D.A. is a paid consultant to Genomics Collaborative, which provided the previously published data set (Am. J. Hum. Genet. 71, 304–311; 2002) that was reanalyzed in this paper.
Supplementary information
Rights and permissions
About this article
Cite this article
Freedman, M., Reich, D., Penney, K. et al. Assessing the impact of population stratification on genetic association studies. Nat Genet 36, 388–393 (2004). https://doi.org/10.1038/ng1333
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1333
This article is cited by
-
An overview of DNA methylation-derived trait score methods and applications
Genome Biology (2023)
-
Using residual regressions to quantify and map signal leakage in genomic prediction
Genetics Selection Evolution (2023)
-
Hybrid autoencoder with orthogonal latent space for robust population structure inference
Scientific Reports (2023)
-
Proteomic association with age-dependent sex differences in Wisconsin Card Sorting Test performance in healthy Thai subjects
Scientific Reports (2023)
-
Children’s Dopaminergic Genotype is Associated with Maternal Reports of Household Chaos during Middle Childhood
Journal of Child and Family Studies (2023)