Abstract
The evolution of complexity relies on changes that result in new gene functions. Here we show that the unique morphological and functional features of the excretory duct cell in C. elegans result from the gain of expression of a single gene. Our results show that innovation can be achieved by altered expression of a transcription factor without coevolution of all target genes.
Similar content being viewed by others
Main
Differences in gene expression patterns contribute to the structural and functional differences between species1,2. These differences can result from loss of a particular regulatory element from the gene in one species or gain of the element in the other. From an evolutionary standpoint, only changes that result in a gain of expression serve to increase genomic and species complexity. Although differences in gene expression patterns have been described previously (e.g., refs. 3–6), evolutionary gain of gene expression has not been conclusively shown, to our knowledge.
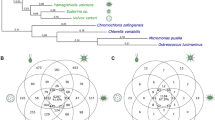
The excretory system in nematodes mediates osmotic and ionic regulation7,8. C. elegans and C. briggsae have different morphologies of part of the excretory system, the excretory duct cell, including different positions of the duct opening9. It was not known whether this difference resulted from loss of a feature in C. briggsae or from gain of a feature in C. elegans. We characterized several strains representing six species and found that C. elegans is unique among Caenorhabditis species with respect to duct position (Fig. 1a). In an assay of excretory system function, C. elegans also showed a greater ability to survive when challenged with high salt (Fig. 1b). Thus, C. elegans has innovations in both structure and function of the excretory system.
(a) Excretory duct position in Caenorhabditis species. Bars (mean ± s.e.m.) show the distance between the excretory duct opening and the base of the posterior pharynx bulb (see Fig. 2c). Species are grouped according to a reported molecular phylogeny11. Our analysis of lin-48 coding and noncoding DNA gives a similar phylogeny for the four species analyzed (data not shown). (b) Salt tolerance in Caenorhabditis species. Percent survival of L3 worms after 24 h on NGM plates made with 500 mM (rather than 50 mM) sodium chloride. (a,b) *P < 0.001 by treatment (strains and species other than C. elegans N2) versus control (C. elegans N2). Data were analyzed using one-way ANOVA and two-sample t-test with Bonferroni adjustment. (c) Alignment of the lin-48 proximal upstream regulatory region from C. briggsae, C. remanei, C. sp. CB5161 and C. elegans. Black arrows indicate the position of the CES-2-response sites identified in the C. elegans gene9. Sequences were aligned using ClustalW 1.8 (ref. 12) and shaded with Boxshade 3.21. (d) lin-48:gfp transgenes from C. elegans9 and C. briggsae9 and comparable ones from C. remanei and C. sp. CB5161 were tested in C. elegans. The percentages of cells expressing green fluorescent protein (GFP; black bar), cells expressing very low but detectable levels of GFP (gray bar) and cells not expressing GFP (white bar) are indicated for each construct. n, number of worms scored. Transgenes were generated and assayed as described9.
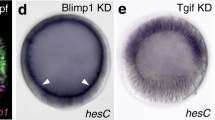
The zinc-finger gene lin-48 is expressed in the C. elegans excretory duct cell and is necessary for normal duct cell features9 (Fig. 2), but it is not expressed in this cell in C. briggsae. One difference between the two species is the presence of enhancers in C. elegans lin-48 that are required for correct expression of lin-48, including sites that respond to the bZip transcription factor CES-2 (ref. 9). We isolated lin-48 from two additional Caenorhabditis species and did not find the CES-2 enhancer sequences that are present in C. elegans lin-48 (Fig. 1c). We made lin-48:gfp reporter transgenes and found that lin-48 from the other species was expressed at lower levels than C. elegans lin-48:gfp in the excretory duct cell (Fig. 1d). In contrast, lin-48 cDNA from each of the four species rescued the structural and functional defects in C. elegans lin-48 mutants (Fig. 2a,b). These results indicate that changes have occurred in the regulation of lin-48 expression but not in the function of LIN-48 protein.
(a) Excretory duct position in transgenic worms. Data presented as in Figure 1. Bars grouped in bracket correspond to transgene-bearing lin-48(sa469) worms. The source of each cDNA is indicated by the two-letter species abbreviation, except C. sp. CB5161 is indicated as CB. Duct distance was significantly shorter in C. elegans N2 and transgene-rescued worms than in lin-48(sa469) mutants (P < 0.001). Under the control of B0303.11 promoter (Ce_pB0303.11), C. briggsae lin-48 cDNA altered the C. briggsae duct to a morphology similar to that of C. elegans wild-type worms (P = 0.56, t-test). (b) Salt tolerance in transgenic worms. C. elegans lin-48(sa469) mutants were sensitive to salt and could be rescued by expression of lin-48 cDNA. In contrast, transgenic C. briggsae worms were as sensitive to salt as were C. briggsae AF16 wild-type worms. (a,b) *P < 0.001 by treatment (strains and species other than C. elegans N2) versus control (C. elegans N2). (c) Diagram of the excretory system. The large excretory cell nucleus (labeled with asterisk) is posterior to the excretory duct cell nucleus (open arrowhead). The excretory duct connects to an opening in the ventral cuticle (filled arrow). The distance between the duct and the base of the posterior pharynx bulb is indicated with a bracket. In the photos, the excretory duct cell nucleus is out of the plane of focus. Excretory system in (d) C. elegans wild-type worm, (e) C. elegans lin-48 mutant, (f) C. briggsae wild-type worm and (g) C. briggsae worm expressing ectopic lin-48 in the excretory duct cell. Left, anterior; top, dorsal.
The genetic and comparative analysis shows C. elegans has unique excretory system features that result from the gain of expression of a transcription factor. However, a transcription factor can mediate cellular effects only if target genes respond. Are naive excretory duct cells able to respond appropriately to LIN-48 when introduced de novo? To test this, we used a promoter that acts in the excretory duct cell in both C. elegans and C. briggsae (Ce_pB0303.11; ref. 10 and data not shown). Under control of this promoter, lin-48 cDNA rescued excretory duct features of C. elegans lin-48 mutants (Fig. 2a,b). Notably, it also conferred C. elegans-like morphology on otherwise wild-type C. briggsae worms. This result indicates that genes downstream of lin-48 that mediate duct cell morphogenesis can respond to LIN-48 in C. briggsae without any further change or coevolution. In contrast, transgenic C. briggsae worms were still sensitive to high salt. Thus, the role of lin-48 in duct morphogenesis is separable from its role in salt tolerance. We propose that this is due to different LIN-48 target genes that mediate the different excretory duct cell features. Our results show that specific, gain-of-function changes in a single regulatory gene between species can result in structural and functional innovation.
References
Tautz, D. Curr. Opin. Genet. Dev. 10, 575–579 (2000).
Levine, M. & Tjian, R. Nature 424, 147–151 (2003).
Sucena, E. & Stern, D.L. Proc. Natl. Acad. Sci. USA 97, 4530–4534 (2000).
Wittkopp, P.J., Vaccaro, K. & Carroll, S.B. Curr. Biol. 12, 1547–1556 (2002).
Sucena, E., Delon, I., Jones, I., Payre, F. & Stern, D.L. Nature 424, 935–938 (2003).
Gompel, N. & Carroll, S.B. Nature 424, 931–935 (2003).
Nelson, F.K. & Riddle, D.L. J. Exp. Zool. 231, 45–56 (1984).
Nelson, F.K., Albert, P.S. & Riddle, D.L. J. Ultrastrastruct. Res. 82, 156–171 (1983).
Wang, X. & Chamberlin, H.M. Genes Dev. 16, 2345–2349 (2002).
Lynch, A.S., Briggs, D. & Hope, I.A. Nat. Genet. 11, 309–313 (1995).
Stein, L.D. et al. PLoS Biol. 1, E45 (2003).
Jeanmougin, F., Thompson, J.D., Gouy, M., Higgins, D.G. & Gibson, T.J. Trends Biochem. Sci. 23, 403–405 (1998).
Acknowledgements
We thank A. Uttam for Ce_plin-48:Ce_cDNA, members of the laboratory of H.M.C. for comments on the manuscript, the Caenorhabditis Genetics Center and D. Fitch for strains and W. Deng and L. Liu for data analysis advice. This work is supported by the US National Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Wang, X., Chamberlin, H. Evolutionary innovation of the excretory system in Caenorhabditis elegans. Nat Genet 36, 231–232 (2004). https://doi.org/10.1038/ng1301
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1301
This article is cited by
-
A toolkit for rapid gene mapping in the nematode Caenorhabditis briggsae
BMC Genomics (2010)
-
Comparative analysis of function and interaction of transcription factors in nematodes: Extensive conservation of orthology coupled to rapid sequence evolution
BMC Genomics (2008)
-
The world of a worm: a framework for Caenorhabditis evolution
EMBO reports (2006)
-
A novel p21WAF1/CIP1 transcript is highly dependent on p53 for its basal expression in mouse tissues
Oncogene (2004)
-
Making a better worm
Nature Genetics (2004)