Abstract
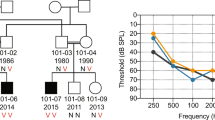
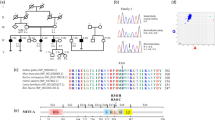
DFNA9 is an autosomal dominant, nonsyndromic, progressive sensorineural hearing loss with vestibular pathology. Here we report three missense mutations in human COCH (previously described as Coch5b2), a novel cochlear gene, in three unrelated kindreds with DFNA9. All three residues mutated in DFNA9 are conserved in mouse and chicken Coch, and are found in a region containing four conserved cysteines with homology to a domain in factor C, a lipopolysaccharide-binding coagulation factor in Limulus polyphemus. COCH message, found at high levels in human cochlear and vestibular organs, occurs in the chicken inner ear in the regions of the auditory and vestibular nerve fibres, the neural and abneural limbs adjacent to the cochlear sensory epithelium and the stroma of the crista ampullaris of the vestibular labyrinth. These areas correspond to human inner ear structures which show histopathological findings of acidophilic ground substance in DFNA9 patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cohen, M.M. & Gorlin, R.J. Epidemiology, etiology, and genetic patterns. in Hereditary Hearing Loss and Its Syndromes (eds Gorlin, R.J., Toriello, H.V. & Cohen, M.M.) 9–21 (Oxford University Press, New York, 1995).
Petit, C. Genes responsible for human hereditary deafness: symphony of a thousand. Nature Genet. 14, 385–391 (1996).
Van Camp, G., Willems, P.J. & Smith, R.J.H. Nonsyndromic hearing impairment: unparalleled heterogeneity. Am. J. Hum. Genet. 60, 758– 764 (1997).
Liu, X.-Z. et al. Mutations in the myosin VIIA gene cause nonsyndromic recessive deafness. Nature Genet. 16, 188– 190 (1997).
Weil, D. et al. The autosomal recessive isolated deafness, DFNB2, and the Usher 1B syndrome are allelic defects of the myosin VIIA gene. Nature Genet. 16, 191–193 (1997).
Liu, X.-Z. et al. Autosomal dominant nonsyndromic deafness caused by a mutation in the myosin VIIA gene. Nature Genet. 17, 268–269 (1997).
Wang, A. et al. Association of unconventional myosin MYO15 mutations with human nonsyndromic deafness DFNB3 [see comments]. Science 280, 1447–1451 (1998).
Kelsell, D.P. et al. Connexin 26 mutations in hereditary non-syndromic sensorineural deafness. Nature 387, 80– 83 (1997).
Scott, D.A., Kraft, M.L., Stone, E.M., Sheffield, V.C. & Smith, R.J.H. Connexin mutations and hearing loss. Nature 391, 32 (1998).
Kelley, P.M. et al. Novel mutations in the connexin 26 gene (GJB2) that cause autosomal recessive (DFNB1) hearing loss. Am. J. Hum. Genet. 62, 792–799 (1998).
Denoyelle, F. et al. Connexin 26 gene linked to a dominant deafness. Nature 393, 319–320 (1998).
Lynch, E.D. et al. Nonsyndromic deafness DFNA1 associated with mutation of a human homolog of the Drosophila gene diaphanous. Science 278, 1315–1318 (1997).
Li, X.C. et al. A mutation in PDS causes nonsyndromic recessive deafness. Nature Genet. 18, 215– 217 (1998).
Vahava, O. et al. Mutation in transcription factor POU4F3 associated with inherited progressive hearing loss in humans. Science 279, 1950–1954 (1998).
Verhoeven, K. et al. Mutations in the human α-tectorin gene cause autosomal dominant nonsyndromic hearing impairment. Nature Genet. 19, 60–62 (1998).
Robertson, N.G., Khetarpal, U., Gutiérrez-Espeleta, G.A., Bieber, F.R. & Morton, C.C. Isolation of novel and known genes from a human fetal cochlear cDNA library using subtractive hybridization and differential screening. Genomics 23, 42–50 (1994).
Robertson, N.G. et al. Mapping and characterization of a novel cochlear gene in human and in mouse: a positional candidate gene for a deafness disorder, DFNA9. Genomics 46, 345– 354 (1997).
Manolis, E.N. et al. A gene for non-syndromic autosomal dominant progressive postlingual sensorineural hearing loss maps to chromosome 14q12–13. Hum. Mol. Genet. 5, 1047–1050 (1996).
Khetarpal, U., Schuknecht, H.F., Gacek, R.R. & Holmes, L.B. Autosomal dominant sensorineural hearing loss: pedigrees, audiologic and temporal bone findings in two kindreds. Arch. Otolaryngol. Head Neck Surg. 117, 1032–1042 (1991).
Khetarpal, U. Autosomal dominant sensorineural hearing loss: further temporal bone findings. Arch. Otolaryngol. Head Neck Surg. 119, 106–108 (1993).
Halpin, C., Khetarpal, U. & McKenna, M. Autosomal dominant progressive sensorineural hearing loss in a large North American Family. Am. J. Audiol. 5, 105–111 (1996).
Colombatti, A. & Paolo, B. The superfamily of proteins with von Willebrand factor type A-like domains: one theme common to components of extracellular matrix, hemostasis, cellular adhesion, and defense mechanisms. Blood 77, 2305– 2315 (1991).
Colombatti, A., Bonaldo, P. & Doliana, R. Type A modules: interacting domains found in several non-fibrillar collagens and in other extracellular matrix proteins. Matrix 13, 297–306 (1993).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Muta, T. et al. Limulus factor C: an endotoxin-sensitive serine protease zymogen with a mosaic structure of complement-like, epidermal growth factor-like, and lectin-like domains. J. Biol. Chem. 266, 6554–6561 (1991).
Iwanaga, S., Miyata, T., Tokunaga, F. & Muta, T. Molecular mechanism of hemolymph clotting system in Limulus. Thrombosis Res. 68, 1–32 (1992).
Nakamura, T. et al. Intracellular serine-protease zymogen, factor C, from horseshoe crab hemocytes: its activation by synthetic lipid A analogues and acidic phospholipids. Eur. J. Biochem. 176, 89– 94 (1988).
Schaeren-Wiemers, N. & Gerfin-Moser, A. A single protocol to detect transcripts of various types and expression levels in neural tissue and cultured cells: in situ hybridization using digoxigenin-labeled cRNA probes. Histochemistry 100, 431– 440 (1993).
Benson, D.W. et al. Missense mutation in the pore region of HERG causes familial long QT syndrome. Circulation 93, 1791– 1795 (1996).
Acknowledgements
The authors are grateful to the three families for their collaboration in this study. We thank S. Blacklow, S. Maxwell, B. McDonough, M. Miri and H. Niimura for their invaluable assistance. This work was supported by NIH grants DC03402 (C.C.M.) and DC00317 (A.J.H.) and by Howard Hughes Medical Institute (A.J.H., C.E.S. and J.G.S.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Robertson, N., Lu, L., Heller, S. et al. Mutations in a novel cochlear gene cause DFNA9, a human nonsyndromic deafness with vestibular dysfunction. Nat Genet 20, 299–303 (1998). https://doi.org/10.1038/3118
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/3118
This article is cited by
-
AudioGene: refining the natural history of KCNQ4, GSDME, WFS1, and COCH-associated hearing loss
Human Genetics (2022)
-
COCH-related autosomal dominant nonsyndromic hearing loss: a phenotype–genotype study
Human Genetics (2022)
-
First Report of Bilateral External Auditory Canal Cochlin Aggregates (“Cochlinomas”) with Multifocal Amyloid-Like Deposits, Associated with Sensorineural Hearing Loss and a Novel Genetic Variant in COCH Encoding Cochlin
Head and Neck Pathology (2020)
-
Novel loss-of-function mutations in COCH cause autosomal recessive nonsyndromic hearing loss
Human Genetics (2020)
-
A systematic review of hearing and vestibular function in carriers of the Pro51Ser mutation in the COCH gene
European Archives of Oto-Rhino-Laryngology (2019)