Abstract
An organism's lifespan is modulated by environmental conditions. When nutrients are abundant, the metabolism of many organisms shifts to growth or reproduction at the expense of longer lifespan, whereas a scarcity of nutrients reverses this shift1,2,3. These correlations suggest that organisms respond to environmental changes by altering their metabolism to promote either reproduction and growth or long life. The only previously reported signaling mechanism involved in this response is the nutrient-responsive insulin/insulin-like growth factor-1 receptor pathway1. Here we report another pathway that controls the length of yeast lifespan. Commitment to cell growth activates the Slt2p MAP kinase pathway, which phosphorylates the transcriptional silencing protein Sir3p, resulting in a shorter lifespan. Elimination of the Sir3p phosphorylation site at Ser275 extended lifespan by 38%. Lifespan extension occurs by a mechanism that is independent of suppressing rDNA recombination. Thus, Slt2p is an enzymatic regulator of silencing function that couples commitment to cell growth and shorter lifespan.
Similar content being viewed by others
Main
Silencing in yeast is a chromatin-based phenomenon mediated by four Sir proteins that silence transcription at the silent-mating-type cassettes (HMRa and HMLα), the telomeres and the array of ribosomal RNA genes (the rDNA; refs. 4,5). Each locus has a unique set of DNA-binding proteins that recruit some or all of the Sir proteins. The size of the Sir protein pool limits the extent of silencing: telomere silencing is limited by the amount of available Sir3p6 whereas rDNA silencing is limited by the availability of Sir2p7. Telomeres are thought to be a reservoir for silencing proteins that can be redistributed to the silent-mating-type cassettes8 and rDNA7.
Sir3p is phosphorylated, and this modification correlates with greater telomere silencing9. We designed a yeast strain in which we could examine the distribution of silencing by simultaneously monitoring silencing at a silent-mating-type cassette, the telomeres and the rDNA array (Fig. 1a). We used this strain to show that single deletions of one of two paralogs, ZDS1 or ZDS2, caused opposite changes in phosphorylation of Sir3p, lifespan and silencing10. The ZDS1 deletion (zds1Δ) caused less Sir3p phosphorylation and longer lifespan, whereas the ZDS2 deletion caused more Sir3p phosphorylation and shorter lifespan10. These data suggested that phosphorylation of Sir3p might shorten yeast lifespan. We therefore used silencing as a reporter to identify the kinase that controls the redistribution of silencing and lifespan by screening for mutants with the zds1Δ silencing phenotype.
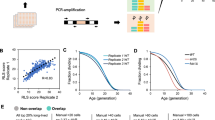
a, The three different reporter genes in the triple silencer strain10. Expression of TRP1 in the hmrΔE locus can be monitored by growth on medium lacking tryptophan, where more growth equals less silencing. Expression of CAN1 in the rDNA locus is monitored using negative selection against CAN1 expression, so that more growth equals more silencing. Expression of URA3 placed adjacent to the right telomere of chromosome V (indicated by the black dot) is monitored by negative selection, so that more growth equals more silencing. Bent arrows show the direction of transcription of each reporter gene. b, A schematic of the Slt2p MAP kinase pathway. c, The level of Slt2p pathway signaling controls the distribution of silencing function. Ten-fold serial dilutions of single yeast colonies bearing the indicated mutations were spotted onto different media to monitor silencing. Representative colonies from three or more assays are shown. Complete medium (com) shows the total number of cells spotted and the other media (FOA, 5-fluoro-orotic acid medium; can, canavanine medium) show the extent of silencing. Arrows to the right indicate the relative level of silencing at each locus compared to wild-type cells, with up arrows showing more silencing and down arrows showing less silencing. Because the CAN1 reporter in the rDNA locus represents a single gene in an array of 100–200 genes, small changes in growth are considered to represent a larger change in silencing compared with the hmr and telomere reporters, which comprise the entire locus. The opposite responses seen for hmr and telomere silencing may be due to their having different DNA-binding proteins that recruit the Sir proteins. The bck1Δ and slt2Δ deletions and the kinase-defective slt2K54R mutation block signaling through the Slt2p MAPK pathway, whereas the sac7Δ mutation enhances signaling.
The only kinase identified in this screen was Slt2p, a mitogen-activated protein kinase (MAPK) in the protein kinase C signaling pathway (Fig. 1b). The properties of Slt2p can explain the previous observations regarding Sir3p phosphorylation. First, heat shock has been shown to increase both Sir3p phosphorylation9 and Slt2p kinase activity11. Second, induction of the Fus3p or Kss1p MAPK cascades increases Sir3p phosphorylation9 and induces polarized cell growth that activates Slt2p11. Because commitment to a new cell cycle induces new polarized growth and also activates Slt2p12, the Slt2p kinase could link commitment to growth with shortened lifespan if phosphorylation of Sir3p was controlled by Slt2p.
Blocking signaling at different steps in the Slt2p pathway (Fig. 1b) should give the same silencing phenotype if this MAPK cascade controls silencing. We found that eliminating different pathway components with the slt2Δ and bck1Δ deletions or substituting Slt2p with the kinase-defective Slt2p-K54R all gave the same silencing phenotype (Fig. 1c). In contrast, constitutively activating the Slt2p MAPK pathway by eliminating Sac7p (ref. 13; Fig. 1b) had the opposite effect on silencing (Fig. 1c). Slt2p was the only MAPK whose elimination changed the relative levels of silencing at all three loci (see Supplementary Fig. 1 online). These data suggested that Slt2p is the kinase that phosphorylates Sir3p to control silencing and shorten lifespan.
Slt2p was shown to be an in vivo Sir3p kinase by several criteria. First, cells lacking Slt2p (slt2Δ) had much lower levels of the slower migrating Sir3p phosphorylated band (Fig. 2a). Second, Slt2p interacted with Sir3p in vivo (Fig. 2b). This analysis showed that only a fraction of Slt2p associated with Sir3p, consistent with the fact that Slt2p also binds and phosphorylates the SBF and Rlm1p transcription factors14,15. Third, Slt2p phosphorylated the N-terminal region of Sir3p whereas the kinase-defective Slt2p-K54R did not (Fig. 2c).
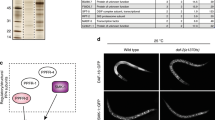
a, Western-blot analysis using antibodies against Sir3p (α-Sir3p) on extracts from cells with the indicated genotypes are shown (wt, wild-type). The arrow indicates the slower migrating Sir3p band that represents phosphorylated protein9. The lower band in the sir3Δ extract is a cross-reacting band unrelated to Sir3p. The sir3-4A lane shows extracts from cells that produce the mutant Sir3p protein Sir3p-4A. b, Western-blot analysis of Sir3p immunoprecipitations (IP). Extracts from slt2Δ cells bearing either vector or an SLT2:HA fusion construct14 were immunoprecipitated with antibody against Sir3p (α-Sir3p) and then analyzed by probing a western blot with antibodies against the hemagglutinin epitope tag (HA) on Slt2:HAp. The slt2Δ sir3Δ SLT2 lane shows that precipitation of Slt2p requires the presence of Sir3p. IgG denotes the band from the immunoprecipitating antibody. c, Slt2p immunoprecipitates can phosphorylate Sir3p in vitro. Cells bearing an slt2Δ allele and a plasmid expressing either SLT2:HA or slt2K54R:HA were heat-shocked at 39 °C for 10 min to activate the Slt2p kinase14, extracts were made and the hemagglutinin-tagged Slt2p proteins were immunoprecipitated with antibodies against hemagglutinin (α-HA) on Protein A Sepharose. The presence of Slt2:HAp and Slt2-K54R:HAp in the extracts was confirmed by western blotting (top row). Purified GST–Sir3p, bearing amino acids 1–439 of Sir3p, was added to the immunoprecipitates in a kinase reaction in the presence of γ-[32P]-ATP. The Coomassie-stained gel shows the amounts of GST–Sir3p added to each assay, and the PhosphorImager scan of the same gel shows the relative levels of [32P] incorporation.
These data and analysis with the NetPhos program identified Ser275, Ser282, Ser289 and Ser295 as the most likely sites of Sir3p phosphorylation. We converted all four serine residues to alanine and expressed this mutated Sir3p (called Sir3p-4A) in wild-type cells as the only functional Sir3 protein. Sir3p-4A migrated as the lower molecular weight form of Sir3p (Fig. 2a). Therefore, some or all of the four serines must have been required for production of the phosphorylated Sir3p band. Slt2p has also recently been identified by others as a kinase that phosphorylates Sir3p in this cluster of serines16.
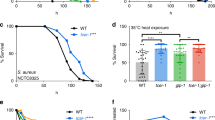
We used the sir3-4A allele in lifespan assays to test our hypothesis that phosphorylation of Sir3p shortens lifespan. Yeast cells undergo asymmetric divisions to produce a larger mother cell and smaller daughter cell, and yeast aging is most frequently measured as a replicative lifespan, that is, the number of times a mother cell can divide before it dies17,18. The sir3-4A cells had a median cell lifespan that was 24% longer than that of wild-type cells (Fig. 3a). This lifespan extension is of the same magnitude as that seen in yeast and rodents as a result of caloric restriction3,19,20 and in worms as a result of overexpression of sir-2.1 protein, an ortholog of yeast Sir2p21. Thus, elimination of Sir3p phosphorylation significantly extended yeast lifespan.
Groups of 40–42 cells were assayed for lifespan, and the fraction of cells alive at each generation was plotted as percent survival against the number of generations that cells had grown. a, The sir3-4A mutation extends lifespan. The median (and maximum) lifespans for wild-type (wt; the W303 strain) were 21 (33), for sir3-4A were 26 (41), for slt2Δ were 21 (30) and for slt2Δ sir3-4A were 19 (30). The 24% extension in lifespan in sir3-4A cells relative to the wild-type strains was significant (P < 0.0001 by log-rank test), whereas the slt2Δ and slt2Δ sir3-4A results were not significantly different from the wild-type results (P > 0.70 and P > 0.15, respectively). b, Sir3p residues Ser275 and Ser282 control lifespan. All strains shown in b and c bear the sir3Δ:kanMX deletion and HIS3-sir3*-his3 single-copy insertion, where sir3* is one of the wild-type or mutated alleles shown. The SIR3 wild-type strain in b and c is therefore different from the wild-type strain in a. c, Lifespan lengths and P values for the different sir3 phosphorylation mutants analyzed in b and in Supplementary Figure 2 online. P values are from log-rank tests for comparison with wild-type lifespan.
To determine the role of Ser275, Ser282, Ser289 and Ser295 in lifespan control, we analyzed two classes of mutations. In the first class, we individually changed each of the four serine codons in the wild-type SIR3 gene to alanine. If phosphorylation of a particular serine shortens lifespan, then cells expressing a sir3 allele that blocks phosphorylation at that serine residue should have a longer lifespan. Changing Ser275 to alanine (the sir3-S275A allele) extended lifespan by 38%, and changing Ser282 to alanine caused a change of borderline significance (Fig. 3b,c).
In the second class of mutations, we individually reverted each mutated alanine codon in the sir3-4A allele back to serine. If phosphorylation of one residue alone (without phosphorylation of the others) controls lifespan, then cells expressing this mutation should have a shorter lifespan. Reverting Ala282 to serine (the sir3-3A282S allele) shortened lifespan to the length seen in the wild type length, and changing Ala289 to serine caused a change of borderline significance (Fig. 3b,c). Analysis of both classes of mutations, therefore, indicates that phosphorylation of Sir3p at Ser275 and Ser282 shortens yeast lifespan.
Cells lacking Slt2p (slt2Δ cells) did not have the extended lifespan of sir3-4A and sir3-S275A cells (Fig. 3a). The slt2Δ mutation perturbs both cell-cycle progression and the cytoskeleton by interfering with the activation of the transcription factors SBF and Rlm1p11. To test whether these global changes were abrogating the lifespan extension caused by eliminating Sir3p phosphorylation, we determined the lifespan of the slt2Δ sir3-4A double mutant. The slt2Δ sir3-4A cells had nearly the same lifespan as wild-type and slt2Δ cells. Thus, the slt2Δ mutation blocked the lifespan extension associated with the sir3-4A mutation.
The silencing phenotype of slt2Δ also differed from the phenotypes of the sir3 phosphorylation mutants (Figs. 1c and 4). Thus, Slt2p may have targets other than Sir3p that affect silencing (for example, Slt2p phosphorylates SBF, which controls the rate of cell-cycle progression, and slower cell-cycle progression alters silencing22). The nine different sir3 mutants showed complex effects on silencing that did not correlate with lifespan. For example, sir3-S275A and sir3-3A275S had extended lifespans but different silencing phenotypes (Figs. 3c and 4). These data suggest that combinations of phosphoserine residues may affect aging and silencing differently. If so, the cell could potentially regulate silencing and lifespan independently to respond to different environmental conditions.
One known cause of yeast aging is the production of autonomously replicating plasmids by rDNA recombination17, a process suppressed by Sir2p23. Sir2p also forms a complex with Sir3p and other proteins at telomeres and silent-mating-type cassettes. Sir3p-4A and Sir3p-S275A might release Sir2p from some chromosomal sites, thereby freeing Sir2p to repress rDNA recombination and extend lifespan. We therefore determined the rDNA recombination frequencies in wild-type, slt2Δ, sir3-4A and sir3-S275A cells. We found that the median recombination frequencies in wild-type, sir3-4A and sir3-275A cells were similar (Fig. 5). These data were in marked contrast with the order-of-magnitude differences seen in long-lived yeast strains in which rDNA recombination is suppressed10,24. Thus, the sir3-4A and sir3-S275A mutations must extend life-span by a mechanism independent of the formation of rDNA circles. One way these sir3 mutants might lengthen lifespan is by silencing rDNA circles so that they do not titrate transcription factors away from essential genes.
a, The system for measuring rDNA recombination frequencies. Recombination within the rDNA repeats can excise the ADE2–CAN1 construct as an episome, which is rapidly lost from progeny cells to produce Ade2− canavanine-resistant (CanR) cells. The fraction of Ade2− cells in a colony is a function of both the rate of rDNA recombination and the point during colony growth at which the recombination event occurs. Events that occur early in the growth of a colony produce a large fraction of Ade2− cells, whereas events that occur later produce a smaller fraction of Ade2− cells. By analyzing several colonies, one can obtain a median frequency that is directly related to the rate of rDNA recombination, using the standard procedures of fluctuation analysis10,29. b, rDNA recombination frequencies (frequency of Ade2− cells per 104 cells) in different strains. Each dot represents a value determined for a single colony, and the bars show the median recombination frequency for all five colonies tested. The wild-type (the W303 strain), slt2Δ, sir3-4A and sir3-S275A strains are the same strains analyzed in Figures 1 and 3.
The slt2Δ cells had higher levels of rDNA recombination than did wild-type cells (Fig. 5b), which could explain why slt2Δ cells did not have an extended lifespan (Fig. 3b): the lifespan of slt2Δ cells is the net result of lifespan shortening due to more rDNA recombination and lifespan lengthening due to lack of Sir3p phosphorylation. This explanation is consistent with the wild-type lifespan of the sir3-4A slt2Δ double mutant; because elimination of the Slt2p MAPK already prevents Sir3p phosphorylation, further blocking of Sir3p phosphorylation with the sir3-4A mutation cannot further extend lifespan.
The results of this work identify the Slt2p MAPK cascade as the pathway that phosphorylates Sir3p to control cell lifespan and the distribution of silencing at different loci. Because Slt2p is activated on commitment to cell growth11,12, our results suggest that rapid cell growth continuously induces the Slt2p MAPK pathway, which phosphorylates Sir3p to shorten lifespan (Fig. 6).
When nutrients are available, the yeast cell commits to another cell cycle, which causes changes in the cell wall that result in the activation of the Slt2p MAPK cascade. Activated Slt2p then phosphorylates SBF to induce the transcription of genes required for cell-cycle progression11 and phosphorylates Rlm1p to induce the transcription of genes required for remodeling the cell wall15. Slt2p also phosphorylates Sir3p, which results in shorter lifespan and redistribution of silencing function. Under conditions of rapid growth, mother cells continuously bud and remodel their cell walls, and so Slt2p is continuously activated. In a state of growth quiescence, Slt2p is not activated and unphosphorylated Sir3p accumulates owing to either new synthesis or dephosphorylation, which leads to extended lifespan.
Methods
Yeast and recombinant DNA methods.
The triple silencer and sir3Δ complete ORF deletion yeast strains have been described10. We used MATα strains. For ORF deletions we used the PCR method and the kanMX G418 resistance marker in pRS400, noting that the orientation of the kanMX marker is opposite to that reported in the GenBank database. For all lifespan and silencing experiments, we used cells with each sir3 allele cloned into pRS303 (ref. 25) and integrated into the HIS3 locus in single copy in the sir3Δ:kanMX strain. We constructed the sir3-4A allele by overlap PCR to convert the four serine codons (TCN) to the corresponding four alanine codons (GCN) in the internal PstI–HindIII fragment, cloning this fragment in place of the same wild-type SIR3 fragment and sequencing the PCR-generated DNA. We then cloned the sir3-4A allele into pRS303. We similarly constructed the individual sir3-S#A mutations using the SIR3 gene as a template to generate the internal EagI–BspDI fragment of SIR3. We constructed the individual sir3-3A#S mutations in the same way, using the sir3-4A allele as template. We confirmed all PCR-generated DNA fragments by DNA sequencing both before and after substitution for the same fragment of the SIR3-pRS303 vector, and we also sequenced the entire sir3-3A289S ORF. Oligonucleotide sequences and plasmid maps of these constructions are available on request. We inserted fragments into the yeast genome and screened for single-copy integrants by Southern blotting following standard procedures. We carried out spot tests for silencing assays by growing cells overnight on yeast extract peptone dextrose (YEPD) plates at 30 °C, streaking cells for single colonies on synthetic complete medium, growing them for 40–42 h at 30 °C, picking three independent colonies from each strain and spotting 5 μl of 10-fold serial dilutions of each colony onto different media. We constructed the glutathione S-tranferase/Sir3p fusion protein (GST–Sir3p) by cloning a PCR fragment encoding the first 439 amino acids of Sir3p (from the ATG to the first EcoRI site in the SIR3 ORF, bp 1,930) into pGEX-6P-1 (Amersham Pharmacia). We purified the fusion protein according to the instructions from the manufacturer. We also used this fusion to affinity-purify antibodies against Sir3p raised in rabbits (N.R., K.W.R., E. Stone and L. Pillus, unpublished data) for use in immunoprecipitations and western blotting.
Identification of potential Sir3p phosphorylation sites.
We used the NetPhos program26 (v. 2.0) and the knowledge that Slt2p is a MAPK that uses Ser-Pro and Thr-Pro sites as substrates to search for potential phosphorylation sites in Sir3p. We identified only four sites with high scores: Ser275, Ser282, Ser289 and Ser295 had respective scores of 0.995, 0.426, 0.941 and 0.997.
Immunoprecipitations and kinase assays.
We carried out immunoprecipitations and western blotting following previously published protocols12. For kinase assays, we used immunoprecipitated proteins isolated from cells that had been heat shocked at 39 °C and followed previously described reaction conditions12 using 0.6 μg of affinity-purified GST–Sir3p.
Mutant isolation.
We subjected the triple silencer strain CCFY100 (ref. 10) to Tn3 transposon mutagenesis27 and screened over 35,000 transposon insertions for greater hmr silencing (poor growth on medium lacking tryptophan), lower telomere silencing (poor growth on 5-fluoro-orotic acid medium) and greater rDNA silencing (better growth on canavanine medium). We confirmed the growth phenotypes of positive mutants by spot tests. We identified the mutated genes by inverse PCR using primers in the Tn3 transposon by the Gottschling lab method (see URLs). A more complete description of the mutant isolation will be published elsewhere.
Lifespan analyses.
We determined lifespan as previously described10,28, except we did the analyses in Figure 3a using YEPD plates containing 1 M sorbitol to mitigate the lysis defect of slt2Δ cells and did the analyses in Figure 3b using YEPD plates. We analyzed survival curves with GraphPad Prism (v. 3.0). For the assays in each panel of Figure 3, we determined the lifespans of the mutants and wild type in the same experiment at the same time.
Determination of rDNA recombination frequency.
We used an assay based on determining the fraction of ade2− cells per colony, identifying the ade2− cells by their ability to form red colonies on YEPD plates. We streaked yeast cells on YEPD plates at room temperature and incubated them at 30 °C for 19–22 h. We picked a single test colony with a sterile Pasteur pipet and resuspended it in 1 ml sterile water. We spun down these cells, resuspended them in 0.1 ml sterile water and determined the number of cells per colony by hemocytometer counting. We immediately picked five new single colonies, resuspended them separately in 1 ml sterile water and made a ten-fold dilution for each suspension. We plated volumes equivalent to 200 cells (based on the test colony) on eight YEPD plates. We spread an additional eight plates with one-fifth the number of cells to allow for variations in the number of cells per colony. We incubated plates for 3 d at 30 °C and 4 d at room temperature to allow colony color to develop. We counted the set of eight plates bearing 100–300 colonies per plate, which allows the clear visualization of completely red colonies, to determine the fraction of red Ade2− colonies. After all of the plates were counted, we picked 50 red and 50 white colonies from each strain (10 colonies from each of the 5 colonies tested) and checked them for the appropriate genotypes. All red colonies were Ade− and strongly resistant to canavanine; all white colonies were Ade+ and weakly resistant to canavanine; and all other auxotrophic markers behaved as expected.
URLs.
NetPhos is available at http://www.cbs.dtu.dk/services/NetPhos/, and information from the Gottschling laboratory is available at http://www.fhcrc.org/labs/gottschling/misc/ipcr.html.
GenBank accession numbers.
Note: Supplementary information is available on the Nature Genetics website.
References
Kenyon, C. A conserved regulatory system for aging. Cell 105, 165–168 (2001).
Carey, J.R., Liedo, P., Muller, H.G., Wang, J.L. & Vaupel, J.W. Dual modes of aging in Mediterranean fruit fly females. Science 281, 996–998 (1998).
Sohal, R.S. & Weindruch, R. Oxidative stress, caloric restriction, and aging. Science 273, 59–63 (1996).
Laurenson, P. & Rine, J. Silencers, silencing and heritable transcriptional states. Microbiol. Rev. 56, 543–560 (1992).
Smith, J.S. & Boeke, J.D. An unusual form of transcriptional silencing in yeast ribosomal DNA. Genes Dev. 11, 241–254 (1997).
Renauld, H. et al. Silent domains are assembled continuously from the telomere and are defined by promoter distance and strength, and by SIR3 dosage. Genes Dev. 7, 1133–1145 (1993).
Smith, J.S., Brachmann, C.B., Pillus, L. & Boeke, J.D. Distribution of a limited Sir2 protein pool regulates the strength of yeast rDNA silencing and is modulated by Sir4p. Genetics 149, 1205–1219 (1998).
Buck, S.W. & Shore, D. Action of a RAP1 carboxy-terminal silencing domain reveals an underlying competition between HMR and telomeres in yeast. Genes Dev. 9, 370–384 (1995).
Stone, E.M. & Pillus, L. Activation of an MAP kinase cascade leads to Sir3p hyperphosphorylation and strengthens transcriptional silencing. J. Cell Biol. 135, 571–583 (1996).
Roy, N. & Runge, K.W. Two paralogs involved in transcriptional silencing that antagonistically control yeast life span. Curr. Biol. 10, 111–114 (2000).
Gustin, M.C., Albertyn, J., Alexander, M. & Davenport, K. MAP kinase pathways in the yeast Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 62, 1264–1300 (1998).
Madden, K. & Snyder, M. Cell polarity and morphogenesis in budding yeast. Annu. Rev. Microbiol. 52, 687–744 (1998).
Martin, H., Rodriguez-Pachon, J.M., Ruiz, C., Nombela, C. & Molina, M. Regulatory mechanisms for modulation of signaling through the cell integrity Slt2-mediated pathway in Saccharomyces cerevisiae. J. Biol. Chem. 275, 1511–1519 (2000).
Madden, K., Sheu, Y.J., Baetz, K., Andrews, B. & Snyder, M. SBF cell cycle regulator as a target of the yeast PKC-MAP kinase pathway. Science 275, 1781–1784 (1997).
de Nobel, H., van Den Ende, H. & Klis, F.M. Cell wall maintenance in fungi. Trends Microbiol. 8, 344–345 (2000).
Ai, W., Bertram, P.G., Tsang, C.K., Chan, T.F. & Zheng, X.F. Regulation of subtelomeric silencing during stress response. Mol. Cell 10, 1295–1305 (2002).
Sinclair, D., Mills, K. & Guarente, L. Aging in Saccharomyces cerevisiae. Annu. Rev. Microbiol. 52, 533–560 (1998).
Jazwinski, S.M. Longevity, genes, and aging. Science 273, 54–59 (1996).
Lin, S.J., Defossez, P.A. & Guarente, L. Requirement of NAD and SIR2 for life-span extension by calorie restriction in Saccharomyces cerevisiae. Science 289, 2126–2128 (2000).
Jiang, J.C., Jaruga, E., Repnevskaya, M.V. & Jazwinski, S.M. An intervention resembling caloric restriction prolongs life span and retards aging in yeast. FASEB J. 14, 2135–2137 (2000).
Tissenbaum, H.A. & Guarente, L. Increased dosage of a sir-2 gene extends lifespan in Caenorhabditis elegans. Nature 410, 227–230 (2001).
Laman, H., Balderes, D. & Shore, D. Disturbance of normal cell cycle progression enhances the establishment of transcriptional silencing in Saccharomyces cerevisiae. Mol. Cell. Biol. 15, 3608–3617 (1995).
Gottlieb, S. & Esposito, R.E. A new role for a yeast transcriptional silencer gene, SIR2, in regulation of recombination in ribosomal DNA. Cell 56, 771–776 (1989).
Kaeberlein, M., McVey, M. & Guarente, L. The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes Dev. 13, 2570–2580 (1999).
Sikorski, R.S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989).
Blom, N., Gammeltoft, S. & Brunak, S. Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J. Mol. Biol. 294, 1351–1362 (1999).
Burns, N. et al. Large-scale analysis of gene expression, protein localization, and gene disruption in Saccharomyces cerevisiae. Genes Dev. 8, 1087–1105 (1994).
Park, P.U., McVey, M. & Guarente, L. Separation of mother and daughter cells. Methods Enzymol. 351, 468–477 (2002).
Lea, D.E. & Coulson, C.A. The distribution of the numbers of mutants in bacterial populations. J. Genet. 49, 264–284 (1949).
Acknowledgements
We thank J. Franco and X. Wang for technical assistance, M. Snyder for providing plasmids, D. Sinclair for advice on lifespan analysis and D. Stacey and R. Wellinger for helpful comments on the manuscript. This work was supported by grants from the US National Institute of Health to K.W.R.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Ray, A., Hector, R., Roy, N. et al. Sir3p phosphorylation by the Slt2p pathway effects redistribution of silencing function and shortened lifespan. Nat Genet 33, 522–526 (2003). https://doi.org/10.1038/ng1132
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1132
This article is cited by
-
Signalling through the yeast MAPK Cell Wall Integrity pathway controls P-body assembly upon cell wall stress
Scientific Reports (2019)
-
PMT1 deficiency enhances basal UPR activity and extends replicative lifespan of Saccharomyces cerevisiae
AGE (2015)
-
Transcriptional outcome of telomere signalling
Nature Reviews Genetics (2014)
-
Rtt107/Esc4 binds silent chromatin and DNA repair proteins using different BRCT motifs
BMC Molecular Biology (2006)
-
The SIT4 gene, which encodes protein phosphatase 2A, is required for telomere function in Saccharomyces cerevisiae
Current Genetics (2005)