Abstract
We took advantage of overlapping interstitial deletions at chromosome 8p11–p12 in two individuals with contiguous gene syndromes and defined an interval of roughly 540 kb associated with a dominant form of Kallmann syndrome, KAL2. We establish here that loss-of-function mutations in FGFR1 underlie KAL2 whereas a gain-of-function mutation in FGFR1 has been shown to cause a form of craniosynostosis. Moreover, we suggest that the KAL1 gene product, the extracellular matrix protein anosmin-1, is involved in FGF signaling and propose that the gender difference in anosmin-1 dosage (because KAL1 partially escapes X inactivation) explains the higher prevalence of the disease in males.
Similar content being viewed by others
Main
Kallmann syndrome involves hypogonadotropic hypogonadism and anosmia, a deficiency of the sense of smell. Anosmia is related to the absence or hypoplasia of the olfactory bulbs and tracts. Hypogonadism is due to deficiency in gonadotropin-releasing hormone (GnRH) and probably results from a failure of embryonic migration of neurons that synthesize GnRH. These cells normally migrate from the olfactory epithelium to the forebrain along the olfactory nerve pathway1. Familial cases of Kallmann syndrome have been reported2,3,4, which suggested X-chromosome linked, autosomal dominant, or, less often, recessive modes of inheritance of the disease. Two intriguing points have been noted: sporadic cases are much more numerous than familial cases, and the disease has a much higher prevalence in males (1:10,000) than in females (the sex ratio has been estimated at 5:1). The gene underlying the X-linked form of the disease (KAL1) has been identified1. But in many families with only males affected and presumably maternal transmission of the disease, no mutation has been found in the KAL1 coding region. Moreover, a molecular analysis of the gene in sporadic male cases rarely identified a mutation (ref. 4 and C.D., unpublished results). This suggests that mutations in the X-linked gene KAL1 do not account for the higher prevalence of the disease in males. Because the incomplete penetrance of the disease2,3 and the infertility in affected individuals impede linkage analysis, the genes underlying the autosomal forms of Kallmann syndrome have so far escaped identification.
By segregation analysis of polymorphic markers combined with fluorescence in situ hybridization (FISH) analysis, we determined that two de novo deletions of roughly 10.7 Mb and 11.6 Mb at chromosome 8p11.2–p12 in two individuals5 who are affected by different contiguous gene syndromes that both include Kallmann syndrome overlap by approximately 540 kb (see Supplementary Notes 1 and 2 and Supplementary Fig. 1 online). Three genes have been reported in this interval; from telomere to centromere, they are WHSC1L1 (OMIM 607083), FGFR1 (OMIM 136350) and the first two exons of TACC1 (OMIM 605301). FGFR1 encodes the fibroblast growth factor receptor 1 and was considered the best candidate to underlie Kallmann syndrome because of a possible functional interaction with the KAL1 gene product.
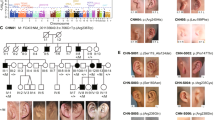
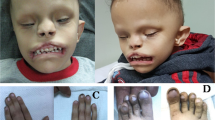
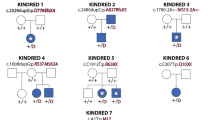
Southern-blot analysis of 43 individuals with familial or sporadic Kallmann syndrome using two genomic probes encompassing exons 4–5 and 16–18 of FGFR1 did not show a deletion of the gene in any of them (data not shown). We then determined the nucleotide sequence of the 18 coding exons and splice sites of FGFR1 in 129 unrelated individuals with Kallmann syndrome (91 males and 38 females). In four familial cases (Fig. 1) and eight sporadic cases, we detected heterozygous mutations of FGFR1: one nonsense mutation, two frameshift mutations, two donor splice–site mutations and seven missense mutations. In addition, we found that one individual, who was born to consanguineous parents and is affected by Kallmann syndrome with cleft palate, agenesis of the corpus callosum, unilateral hearing loss and fusion of the fourth and fifth metacarpal bones, was homozygous with respect to a deleterious missense mutation (Table 1 and Supplementary Fig. 2 online). From these results, we conclude that FGFR1 underlies one autosomal dominant form of Kallmann syndrome (KAL2). Moreover, cleft palate or lip and dental agenesis, two anomalies that are occasionally associated with Kallmann syndrome1,2,3 and were present in five individuals with mutations in FGFR1 (Table 1), can now be ascribed to this genetic form of the disease. We also found that bimanual synkinesia, identified in one family with KAL2 (Table 1), can be observed in autosomal Kallmann syndrome, whereas it had thus far been considered to be specific to the X-linked form of the disease.
Filled symbols indicate clinically affected individuals with both hypogonadism and anosmia. Half-filled symbols indicate individuals with anosmia only. Open symbols containing a black dot indicate unaffected carriers of the mutation. N indicates absence of the mutation. In family 1, individual IV2 (who was 16 years old) underwent spontaneous puberty but suffers from anosmia; individuals III2, IV1 and IV2 have multiple dental agenesis. In family 2, individuals II1 and II2 both have bimanual synkinesia. In family 3, individual III1 has a cleft lip and individual III2 has both a cleft lip and a cleft palate. Notably, non-penetrance of the disease in some mutation carriers can simulate recessive transmission of Kallmann syndrome in certain families, for example, families 1, 2 and 4. Diagonal line indicates deceased individual; question mark indicates unknown clinical status.
FGF signaling is involved in a variety of developmental processes including the formation, growth and shaping of tissues and organs. Extracellular interaction between FGF, the FGF receptor and heparan sulfate proteoglycans (HSPGs) is required for receptor dimerization and resulting autophosphorylation of several tyrosine residues in the intracellular domain. The phosphotyrosines either stimulate protein tyrosine kinase activity of the receptor or serve as docking sites for downstream signaling molecules. A dominant gain-of-function mutation of FGFR1 underlies a form of craniosynostosis6. This mutation is believed to result in enhanced affinity for certain FGF ligands7. In contrast, KAL2 results from loss-of-function mutations in FGFR1, as shown by the presence of a single allele in the individuals carrying a chromosomal deletion at 8p11.2. Moreover, some of the mutations associated with KAL2 could hinder the formation of functional receptor dimers by a dominant negative effect. Whatever the molecular mechanism may be, our results indicate that olfactory bulb development in humans is crucially sensitive to reduced dosage of FGFR1. Dominant gain-of-function and loss-of-function mutations in FGFR1 cause different developmental disorders: premature fusion of skull bone sutures (craniosynostosis) and failed morphogenesis of the olfactory bulbs (Kallmann syndrome), respectively. Given the phenotype associated with gain-of-function mutation of FGFR1, it is noteworthy that delayed closure of calvarial sutures has not been reported in Kallmann syndrome.
The implication of FGFR1 and KAL1 in the same developmental disease raises the possibility that the gene products functionally interact. KAL1 encodes anosmin-1, a locally restricted protein of roughly 100 kDa in embryonic extracellular matrices8. Several arguments suggest that anosmin-1 is involved in FGF signaling through FGFR1. First, anosmin-1 binds to HSPGs9, and HSPGs are important in the dimerization of the binary FGF–FGFR complex10. Also, the axon-branching phenotype in some neurons in Caenorhabditis elegans that mis- or overexpress the nematode gene kal1 was no longer present in mutant worms lacking heparan 6O-sulfotransferase, an enzyme involved in HSPG biosynthesis11,12. Second, KAL1 and FGFR1 are coexpressed at different sites during embryonic development. KAL1 is expressed in the presumptive olfactory bulbs8, and Fgfr1 expression in the rostral forebrain is required for initial olfactory bulb evagination in the mouse13. In addition, cleft palate or cleft lip was present in several individuals with KAL2 (Table 1), and a high arched palate, which can be regarded as a mild anomaly of palatal fusion, is a common feature of KAL1. Finally, the bimanual synkinesia that we report in one family with KAL2 has been observed in 75% of KAL1 cases.
Regarding the low prevalence of the X-linked form of the disease4, the proposal that KAL1 and FGFR1 functionally interact provides a possible explanation for the higher prevalence of Kallmann syndrome in males. The explanation is based on the assumption that the local concentration of anosmin-1 is important in FGF signaling and the fact that KAL1 partially escapes X inactivation in females14. Accordingly, females are expected to synthesize a higher amount of anosmin-1 than do males; in some cases, this could be enough to maintain FGF signaling above the critical threshold in the context of FGFR1 haploinsufficiency. Indeed, in four of five families in whom we were able to follow the transmission of the FGFR1 mutation, the mutation was transmitted by the unaffected mother (Table 1). Moreover, the probable pseudoautosomal location of Kal1 in rodents can account for the observation that male mice heterozygous with respect to a null allele at the Fgfr1 locus are viable and fertile13.
Note: Supplementary information is available on the Nature Genetics website.
References
Hardelin, J.-P. Mol. Cell. Endocrinol. 179, 75–81 (2001).
Santen, R.J. & Paulsen, C.A. J. Clin. Endocrinol. Metab. 36, 47–54 (1972).
White, B.J., Rogol, A.D., Brown, K.S., Lieblich, J.M. & Rosen, S.W. Am. J. Med. Genet. 15, 417–436 (1983).
Oliveira, L.M.B. et al. J. Clin. Endocrinol. Metab. 86, 1532–1538 (2001).
Vermeulen, S. et al. Am. J. Med. Genet. 108, 315–318 (2002).
Muenke, M. et al. Nat. Genet. 8, 269–274 (1994).
Anderson, J., Burns, H., Enriquez-Harris, P., Wilkie, A. & Heath, J. Hum. Mol. Genet. 7, 1475–1483 (1998).
Hardelin, J.-P. et al. Dev. Dyn. 215, 26–44 (1999).
Soussi-Yanicostas, N. et al. J. Cell Sci. 109, 1749–1757 (1996).
Pellegrini, L. Curr. Opin. Struct. Biol. 11, 629–634 (2001).
Bülow, H.E., Berry, K.L., Topper, L.H., Peles, E. & Hobert, O. Proc. Natl. Acad. Sci. USA 99, 6346–6351 (2002).
Soussi-Yanicostas, N. et al. Cell 109, 217–228 (2002)
Hébert, J.M., Partanen, J., Rossant, J. & McConnell, S.K. Development 130, 1101–1111 (2003).
Franco, B. et al. Nature 353, 529–536 (1991).
Plotnikov, A.N., Schlessinger, J., Hubbard, S.R. & Mohammadi, M. Cell 98, 641–650 (1999).
Acknowledgements
We thank the individuals with Kallmann syndrome and their families for their participation in this study, the Service de séquençage de l'Institut Cochin and J. Weissenbach at the Centre National de Séquençage for sequencing facilities, B. Menten and E. Hermanci for expert technical assistance, J. Hébert and S. McConnell for sharing results before publication and S. Cure for critical reading of the manuscript. This study was supported by European Community, Human Frontiers Science Program (Génétique des Déficits Sensoriels, Paris, France) and Fund for Scientific Research-Flanders (Belgium).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Dodé, C., Levilliers, J., Dupont, JM. et al. Loss-of-function mutations in FGFR1 cause autosomal dominant Kallmann syndrome. Nat Genet 33, 463–465 (2003). https://doi.org/10.1038/ng1122
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1122
This article is cited by
-
Mutation spectrum of Kallmann syndrome: identification of five novel mutations across ANOS1 and FGFR1
Reproductive Biology and Endocrinology (2023)
-
Rare structural variants, aneuploidies, and mosaicism in individuals with Mullerian aplasia detected by optical genome mapping
Human Genetics (2023)
-
How human genetic context can inform pathogenicity classification: FGFR1 variation in idiopathic hypogonadotropic hypogonadism
Human Genetics (2023)
-
Whole-Exome Sequencing Analysis of Idiopathic Hypogonadotropic Hypogonadism: Comparison of Varicocele and Nonobstructive Azoospermia
Reproductive Sciences (2023)
-
Clinical, hormonal, and genetic characteristics of 25 Chinese patients with idiopathic hypogonadotropic hypogonadism
BMC Endocrine Disorders (2022)