Abstract
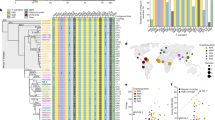
Binary polymorphisms associated with the non-recombining region of the human Y chromosome (NRY) preserve the paternal genetic legacy of our species that has persisted to the present, permitting inference of human evolution, population affinity and demographic history1. We used denaturing high-performance liquid chromatography (DHPLC; ref. 2) to identify 160 of the 166 bi-allelic and 1 tri-allelic site that formed a parsimonious genealogy of 116 haplotypes, several of which display distinct population affinities based on the analysis of 1062 globally representative individuals. A minority of contemporary East Africans and Khoisan represent the descendants of the most ancestral patrilineages of anatomically modern humans that left Africa between 35,000 and 89,000 years ago.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Hammer, M.F. & Zegura, S.L. The role of the Y chromosome in human evolutionary studies. Evol. Anthropol. 5, 116–134 (1996).
Oefner, P.J. & Underhill, P.A. DNA mutation detection using denaturing high-performance liquid chromatography. Current Protocols in Human Genetics. Suppl 19, 7.10.1– 7.10.12 (Wiley & Sons, New York, 1998).
Underhill, P.A. et al. Detection of numerous Y chromosome biallelic polymorphisms by denaturing high performance liquid chromatography (DHPLC). Genome Res. 7, 996–1005 (1997).
Shen, P. et al. Population genetic implications from sequence variation in four Y chromosome genes. Proc. Natl Acad. Sci. USA 97, 7354–7359 (2000).
Hammer, M.F. et al. Out of Africa and back again: nested cladistic analysis of human Y chromosome variation. Mol. Biol. Evol. 15, 427–441 (1998).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism Genetics 123, 585–595 (1989).
Nachman, M.W. Y chromosome variation of mice and men. Mol. Biol. Evol. 15, 1744–1750 (1998).
Jaruzelska, J., Zietkiewicz, E. & Labuda, D. Is selection responsible for the low level of variation in the last intron of the ZFY locus? Mol. Biol. Evol. 16, 1633–1640 (1999).
Wyckoff, G.J., Wang, W. & Wu, C.I. Rapid evolution of male reproductive genes in the descent of man. Nature 403, 304–309 ( 2000).
Jorde, L.B. et al. The distribution of human genetic diversity: a comparison of mitochondrial, autosomal, and Y-chromosome data. Am. J. Hum. Genet. 66, 979–988 ( 2000).
Thomson, R. et al. Recent common ancestry of human Y chromosomes: Evidence from DNA sequence data. Proc. Natl Acad. Sci. USA 97, 7360–7365 (2000).
Pritchard, J.K., Seielstad, M.T., Perez-Lezaun, A. & Feldman, M.W. Population growth of human Y chromosomes: a study of Y chromosome microsatellites. Mol. Biol. Evol. 16, 1791–1798 (1999).
Klein, R.G. The Human Career: Human Biological and Cultural Origins (University of Chicago Press, Illinois, 1999).
Quintana-Murci, L. et al. Genetic evidence of an early exit of Homo sapiens sapiens from Africa through eastern Africa. Nature Genet. 23 , 437–441 (1999).
Rogers, A.R. Genetic evidence for a Pleistocene population explosion. Evolution 49, 608–615 ( 1995).
Vollrath, D. et al. The human Y chromosome: a 43-interval map based on naturally occurring deletions. Science 258, 52– 59 (1992).
Hammer, M.F. & Horai, S. Y chromosomal DNA variation and the peopling of Japan. Am. J. Hum. Genet. 56, 951–962 (1995).
Seielstad, M.T. et al. Construction of human Y-chromosome haplotypes using a new polymorphic A to G transition. Hum. Mol. Genet. 3, 2159–2161 (1994).
Hammer, M.F. et al. The geographic distribution of human Y chromosome variation . Genetics 145, 787–805 (1997).
Zerjal, T. et al. Genetic relationships of Asians and northern Europeans, revealed by Y-chromosomal DNA analysis. Am. J. Hum. Genet. 60 , 1174–1183 (1997).
Bergen, A.W. et al. An Asian-native American paternal lineage identified by RPS4Y resequencing and by microsatellite haplotyping. Ann. Hum Genet. 63, 63–80 ( 1999).
Bianchi, N.O. et al. Origin of Amerindian Y-chromosomes as inferred by the analysis of six polymorphic markers. Am. J. Phys. Anthropol. 102, 79–89 (1997).
Diamond, J. Guns, Germs, and Steel (Norton, New York, 1999).
Macaulay, V. et al. The emerging tree of west Eurasian mtDNAs: a synthesis of control-region sequences and RFLPs. Am. J. Hum. Genet. 64, 232–249 (1999).
Jin, L. et al. Distribution of haplotypes from a chromosome 21 region distinguishes multiple prehistoric human migrations. Proc. Natl Acad. Sci. USA 96, 3796–3800 ( 1999).
Cavalli-Sforza, L.L., Menozzi, P. & Piazza, A. The History and Geography of Human Genes (Princeton University Press, Princeton, New Jersey, 1994).
Acknowledgements
We thank the 1,062 men who donated DNA; R.G. Klein, J. Mountain and M. Ruhlen for helpful discussions; D. Vollrath, R. Hyman and F.S. Dietrich for Y-specific cosmid sequences; and J. Block, D. Soergel, K. Prince, C. Edmonds and A. Rojas for technical help. A.W. Bergen made the RPS4YC711T marker (M130) information available to us before its publication. This work was supported in part by the NIH, NIHGR and L.S.B. Leakey Foundation.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
Underhill, P., Shen, P., Lin, A. et al. Y chromosome sequence variation and the history of human populations. Nat Genet 26, 358–361 (2000). https://doi.org/10.1038/81685
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/81685
This article is cited by
-
Parallel signatures of Mycobacterium tuberculosis and human Y-chromosome phylogeography support the Two Layer model of East Asian population history
Communications Biology (2023)
-
Y-chromosome target enrichment reveals rapid expansion of haplogroup R1b-DF27 in Iberia during the Bronze Age transition
Scientific Reports (2022)
-
Origin and diffusion of human Y chromosome haplogroup J1-M267
Scientific Reports (2021)
-
Genetic Predisposition and its Heredity in the Context of Increased Prevalence of Dermatophytoses
Mycopathologia (2021)
-
New insights on intercontinental origins of paternal lineages in Northeast Brazil
BMC Evolutionary Biology (2020)