Abstract
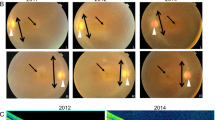
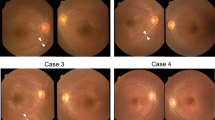
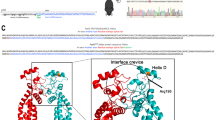
Retinitis pigmentosa (RP) comprises a clinically and genetically heterogeneous group of diseases that afflicts approximately 1.5 million people worldwide. Affected individuals suffer from a progressive degeneration of the photoreceptors, eventually resulting in severe visual impairment. To isolate candidate genes for chorioretinal diseases, we cloned cDNAs specifically or preferentially expressed in the human retina and the retinal pigment epithelium (RPE) through a novel suppression subtractive hybridization (SSH) method1,2. One of these cDNAs (RET3C11) mapped to chromosome 1q31–q32.1, a region harbouring a gene involved in a severe form of autosomal recessive RP characterized by a typical preservation of the para-arteriolar RPE (RP12; ref. 3). The full-length cDNA encodes an extracellular protein with 19 EGF-like domains, 3 laminin A G-like domains and a C-type lectin domain. This protein is homologous to the Drosophila melanogaster protein crumbs (CRB), and denoted CRB1 (crumbs homologue 1). In ten unrelated RP patients with preserved para-arteriolar RPE, we identified a homozygous AluY insertion disrupting the ORF, five homozygous missense mutations and four compound heterozygous mutations in CRB1. The similarity to CRB suggests a role for CRB1 in cell-cell interaction and possibly in the maintenance of cell polarity in the retina. The distinct RPE abnormalities observed in RP12 patients suggest that CRB1 mutations trigger a novel mechanism of photoreceptor degeneration.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
den Hollander, A.I. et al. Isolation and mapping of novel candidate genes for retinal disorders using suppression subtractive hybridization. Genomics 58, 240–249 (1999).
Diatchenko, L. et al. Suppression subtractive hybridization: a method for generating differentially regulated or tissue-specific cDNA probes and libraries. Proc. Natl Acad. Sci. USA 93, 6025–6030 (1996).
van Soest, S. et al. Integrated genetic and physical map of the 1q31–q32.1 region, encompassing the RP12 locus, the F13B and HF1 genes and the EEF1AL11 and RPL30 pseudogenes. Cytogenet. Cell Genet. 84, 22–27 (1999).
Heckenlively, J.R. Preserved para-arteriole retinal pigment epithelium (PPRPE) in retinitis pigmentosa. Br. J. Ophthalmol. 66, 26–30 (1982).
van den Born, L.I. et al. Autosomal recessive retinitis pigmentosa with preserved para-arteriolar retinal pigment epithelium. Am. J. Ophthalmol. 118, 430–439 (1994).
von Heijne, G. Signal sequences. The limits of variation. J. Mol. Biol. 184, 99–105 (1985).
Doolittle, R.F., Feng, D.F. & Johnson, M.S. Computer-based characterization of epidermal growth factor precursor. Nature 307, 558–560 (1984).
Rothberg, J.M. & Artavanis-Tsakonas, S. Characterization of a conserved carboxy-terminal sequence in secreted proteins and a motif implicated in extracellular protein interactions. J. Mol. Biol. 227, 367–370 (1992).
Sasaki, M., Kleinman, H.K., Huber, H., Deutzmann, R. & Yamada, Y. Laminin, a multidomain protein. J. Biol. Chem. 263, 16536–16544 (1988).
Drickamer, K. Two distinct classes of carbohydrate-recognition domains in animal lectins. J. Biol. Chem. 263, 9557–9560 (1988).
Tepass, U., Theres, C. & Knust, E. crumbs encodes an EGF-like protein expressed on apical membranes of Drosophila epithelial cells and required for organization of epithelia. Cell 61, 787–799 (1990).
Kere, J. et al. Mapping human chromosomes by walking with sequence-tagged sites from end fragments of yeast artificial chromosome inserts. Genomics 14, 241–248 (1992).
van Soest, S. et al. Assignment of a gene for autosomal recessive retinitis pigmentosa (RP12) to chromosome 1q31-q32.1 in a inbred and genetically heterogeneous disease population. Genomics 22, 499–504 (1994).
Batzer, M.A. et al. Standardized nomenclature for Alu repeats. J. Mol. Evol. 42, 3–6 (1996).
Vidaud, D. et al. Haemophilia B due to a de novo insertion of a human-specific Alu subfamily member within the coding region of the factor IX gene. Eur. J. Hum. Genet. 1, 30–36 (1993).
Shapiro, M.B. & Senapathy, P. RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression. Nucleic Acids Res. 15, 7155–7174 (1987).
Appella, E., Weber, I.T. & Blasi, F. Structure and function of epidermal growth factor-like regions in proteins. FEBS Lett. 231, 1–4 (1988).
Knust, E. et al. EGF homologous sequences encoded in the genome of Drosophila melanogaster, and their relation to neurogenic genes. EMBO J. 6, 761–766 (1987).
Wodarz, A., Hinz, U., Engelbert, M. & Knust, E. Expression of crumbs confers apical character on plasma membrane domains of ectodermal epithelia of Drosophila. Cell 82, 67–76 (1995).
Nakayama, M. et al. Identification of high-molecular weight proteins with multiple EGF-like motifs by motif-trap screening. Genomics 51, 27–34 (1998).
Ushkaryov, Y.A., Petrenko, A.G., Geppert, M. & Südhof, T.C. Neurexins: synaptic cell surface proteins related to the →-latrotoxin receptor and laminin. Science 257, 50–56 (1992).
Kallunki, P. & Tryggvason, K. Human basement membrane heparan sulfate proteoglycan core protein: a 467-kD protein containing multiple domains resembling elements of the low density lipoprotein receptor, laminin, neural cell adhesion molecules, and epidermal growth factor. J. Cell Biol. 116, 559–571 (1992).
Joseph, D.R. & Baker, M.E. Sex hormone-binding globulin, androgen-binding protein, and vitamin K-dependent protein S are homologous to laminin A, merosin, and Drosophila crumbs protein. FASEB J. 6, 2477–2481 (1992).
Patthy, L. A family of laminin-related proteins controlling ectodermal differentiation in Drosophila. FEBS Lett. 298, 182–184 (1992).
Dryja, T.P. & Li, T. Molecular genetics of retinitis pigmentosa. Hum. Mol. Genet. 4, 1739–1743 (1995).
van Soest, S., Westerveld, A., de Jong, P.T.V.M., Bleeker-Wagemakers, E.M. & Bergen, A.A.B. Retinitis pigmentosa: defined from a molecular point of view. Surv. Ophthalmol. 43, 321–334 (1999).
Pearson, W.R. & Lipman, D.J. Improved tools for biological sequence comparison. Proc. Natl Acad. Sci. USA 85, 2444–2448 (1988).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Dunn, K.C., Aotaki-Keen, A.E., Putkey, F.R. & Hjelmeland, L.M. ARPE-19, a human retinal pigment epithelial cell line with differentiated properties. Exp. Eye Res. 62, 155–169 (1996).
Davis, A.A. et al. A human retinal pigment epithelial cell line that retains epithelial characteristics after prolonged culture. Invest. Ophthalmol. Vis. Sci. 36, 955–964 (1995).
Acknowledgements
We thank R. Roepman and H. Yntema for their help with northern-blot analysis and cDNA library screening; M. van Schooneveld and D. Bessant for ascertaining patients with RP12; and P. de Jong for his continuous interest in this project. This work was supported by The Foundation Fighting Blindness, Inc., USA and The British Retinitis Pigmentosa Society.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
den Hollander, A., ten Brink, J., de Kok, Y. et al. Mutations in a human homologue of Drosophila crumbs cause retinitis pigmentosa (RP12). Nat Genet 23, 217–221 (1999). https://doi.org/10.1038/13848
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/13848
This article is cited by
-
Genotype-phenotype associations in CRB1 bi-allelic patients: a novel mutation, a systematic review and meta-analysis
BMC Ophthalmology (2024)
-
Der Crumbs-Komplex – von Epithelpolarität zu retinaler Degeneration
BIOspektrum (2023)
-
Dissecting the role of EYS in retinal degeneration: clinical and molecular aspects and its implications for future therapy
Orphanet Journal of Rare Diseases (2021)
-
Podocyte-specific Crb2 knockout mice develop focal segmental glomerulosclerosis
Scientific Reports (2021)
-
Unraveling the genetic cause of hereditary ophthalmic disorders in Arab societies from Israel and the Palestinian Authority
European Journal of Human Genetics (2020)