Abstract
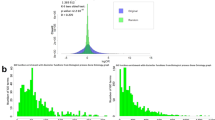
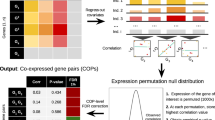
Chromosome correlation maps display correlations between the expression patterns of genes on the same chromosome. Using these maps, we show here that adjacent pairs of genes, as well as nearby non-adjacent pairs of genes, show correlated expression independent of their orientation. We present specific examples of adjacent pairs with highly correlated expression patterns, in which the promoter of only one of the two genes contains an upstream activating sequence (UAS) known to be associated with that expression pattern. Finally, we show that genes with similar functions tend to occur in adjacent positions along the chromosomes. Our results suggest that, in certain chromosomal expression domains, an UAS can affect the transcription of genes that are not immediately downstream from it.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cho, R.J. et al. A genome-wide transcriptional analysis of the mitotic cell cycle . Mol. Cell 2, 65–73 (1998).
Chu, S. et al. The transcriptional program of sporulation in budding yeast. Science 282, 699–705 ( 1998).
Roberts, C.J. et al. Signaling and circuitry of multiple MAPK pathways revealed by a matrix of global gene expression profiles. Science 287, 873–880 (2000).
Hughes, J.D., Estep, P.W., Tavazoie, S. & Church, G.M. Computational identification of cis-regulatory elements associated with groups of functionally related genes in Saccharomyces cerevisiae. J. Mol. Biol. 296, 1205–1214 ( 2000).
Zhu, J. & Zhang, M.Q. SCPD: a promoter database of the yeast Saccharomyces cerevisiae. Bioinformatics 15, 607–611 (1999).
Tavazoie, S., Hughes, J.D., Campbell, M.J., Cho, R.J. & Church, G.M. Systematic determination of genetic network architecture. Nature Genet. 22, 281–285 (1999).
Mewes, H.W. et al. MIPS: a database for genomes and protein sequences. Nucleic Acids Res. 28, 37–40 (2000).
Kraakman, L.S., Mager, W.H., Maurer, K.T., Nieuwint, R.T. & Planta, R.J. The divergently transcribed genes encoding yeast ribosomal proteins L46 and S24 are activated by shared RPG-boxes . Nucleic Acids Res. 17, 9693– 9706 (1989).
Nakao, J., Miyanohara, A., Toh-e, A. & Matsubsara, K. Saccharomyces cerevisiae PHO5 promoter region: location and function of the upstream activation site. Mol. Cell. Biol. 6, 2613–2623 (1986).
Osley, M.A., Gould, J., Kim, S., Kane, M.Y. & Hereford, L. Identification of sequences in a yeast histone promoter involved in periodic transcription. Cell 45, 537–544 (1986).
West, R.W. Jr, Yocum, R.R. & Ptashne, M. Saccharomyces cerevisiae GAL1-GAL10 divergent promoter region: location and function of the upstream activating sequence UASG. Mol. Cell. Biol. 4, 2467– 2478 (1984).
Nasmyth, K.A. repetitive DNA sequence that confers cell-cycle START (CDC28)-dependent transcription of the HO gene in yeast. Cell 42, 225– 235 (1985).
Koch, C., Schleiffer, A., Ammerer, G. & Nasmyth, K. Switching transcription on and off during the yeast cell cycle: Cln/Cdc28 kinases activate bound transcription factor SBF (Swi4/Swi6) at start, whereas Clb/Cdc28 kinases displace it from the promoter in G2. Genes Dev. 10, 129–141 ( 1996).
Felsenfeld, G. Chromatin unfolds. Cell 86, 13– 19 (1996).
Moore, D.S. & McCabe, G.P. Introduction to the Practice of Statistics (W.H. Freeman, New York, 1998).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Graber, J.H., Cantor, C.R., Mohr, S.C. & Smith, T.F. In silico detection of control signals: mRNA 3′-end-processing sequences in diverse species . Proc. Natl Acad. Sci. USA 96, 14055– 14060 (1999).
Acknowledgements
We thank J. Aach, W. Rindone, S. Tavazoie, J. Graber, K. Struhl and F. Winston for advice, suggestions and data files; and P. Sudarsanam, A. Dudley, T. Pilpel, A. Derti, P. Estep, M. Steffen, V. Badarinarayana, T. Wu and M. Bulyk for discussions and critical readings of the manuscript. B.A.C. was supported by a postdoctoral fellowship from the American Cancer Society (PF-98-159-01-MBC). This work was supported by the US Department of Energy (DE-FG02-87-ER60565), the Office of Naval Research and DARPA (N00014-97-1-0865), the Lipper Foundation and Hoechst Marion Roussel.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cohen, B., Mitra, R., Hughes, J. et al. A computational analysis of whole-genome expression data reveals chromosomal domains of gene expression. Nat Genet 26, 183–186 (2000). https://doi.org/10.1038/79896
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/79896
This article is cited by
-
Gene function prediction in five model eukaryotes exclusively based on gene relative location through machine learning
Scientific Reports (2022)
-
Evolutionary and functional patterns of shared gene neighbourhood in fungi
Nature Microbiology (2019)
-
Neighboring genes are closely related to whole genome duplications after their separation
Interdisciplinary Sciences: Computational Life Sciences (2019)
-
Global identification of Arabidopsis lncRNAs reveals the regulation of MAF4 by a natural antisense RNA
Nature Communications (2018)
-
An analysis of aging-related genes derived from the Genotype-Tissue Expression project (GTEx)
Cell Death Discovery (2018)