Abstract
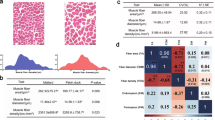
An exceptional muscle development commonly referred to as ‘double-muscled’ (Fig. 1) has been seen in several cattle breeds and has attracted considerable attention from beef producers. Double-muscled animals are characterized by an increase in muscle mass of about 20%, due to general skeletal-muscle hyperplasia—that is, an increase in the number of muscle fibers rather than in their individual diameter1. Although the hereditary nature of the double-muscled condition was recognized early on, the precise mode of inheritance has remained controversial; monogenic (dominant and recessive), oligogenic and polygenic models have been proposed2. In the Belgian Blue cattle breed (BBCB)4, segregation analysis performed both in experimental crosses3 and in the outbred population suggested an autosomal recessive inheritance. This was confirmed when the muscular hypertrophy (mh) locus was mapped 3.1 cM from microsatellite TGLA44 on the centromeric end of bovine chromosome 2 (ref. 5). We used a positional candidate approach to demonstrate that a mutation in bovine MSTN, which encodes myostatin, a member of the TGFβ superfamily, is responsible for the double-muscled phenotype. We report an 11-bp deletion in the coding sequence for the bioactive carboxy-termihal domain of the protein causing the muscular hypertrophy observed in Belgian Blue cattle.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Hanset, R. The major gene of muscular hypertrophy. in The Belgian Blue cattle breed. in Breeding for Disease Resistance in Farm Animals (eds Owen, J. B. & Axford, R.F.E.) 467–478 (C.A.B. International, Oxford, 1991).
Ménissier, F. Present state of knowledge about the genetic determination of muscular hypertrophy or the double muscled trait in cattle. in Current Topics in Veterinary Medicine and Animal Science, vol 16: Muscle Hyertrophy of Genetic Origin and Its Use to Improve Beef Production (eds King, J.W.B. & Ménissier, F.) 387–428 (Martinus Nijhoff, Norwell, Massachusetts, 1982).
Hanset, R. & Michaux, C. On the genetic determinism of muscular hypertrophy in the Belgian White and Blue cattle breed. I. Experimental data. Genet. Sel. Evol. 17, 359–368 (1985).
Hanset, R. & Michaux, C. On the genetic determinism of muscular hypertrophy in the Belgian White and Blue cattle breed: II. Population data. Genet Sel. Evol. 17, 369–386 (1985).
Charlier, C. et al. The mh gene causing double-muscling in cattle maps to bovine chromosome 2. Mamm. Genome 6, 788–792 (1995).
Solinas-Toldo, S., Lengauer, C. & Fries, R. Comparative genome map of man and cattle. Genomics 27, 489–496 (1995).
Fisher, S.R., Beever, J.E. & Lewin, H.A. Genetic mapping of COL3AI to bovine chromosome 2. Mamm. Genome 8, 76–77 (1996).
O'Brien, S.J. et al. Anchored reference loci for comparative genome mapping in mammals. Nature Genet. 3, 103–112 (1993).
Lyons, A.L. et al. Comparative anchor tagged sequences (CATS) for integrative mapping of mammalian genomes. Nature Genet. 15, 47–56 (1996).
Kappes, S.M. et al. A second-generation linkage map of the bovine genome. Genome Res. 7, 235–249 (1997).
McPherron, A.C., Lawler, A.M. & Lee, S.-J. Regulation of skeletal muscle mass in mice by a new TGFβ superfamily member. Nature 387, 83–90 (1997).
Walter, M.A., Spillett, D.J., Thomas, P., Weissenbach, J. & Goodfellow, P.N. A method for constructing radiation hybrid maps of whole genomes. Nature Genet. 7, 22–28 (1994).
Hudson, T.G. et al. An STS-based map of the human genome. Science 270, 1945–1954 (1995), with supplementary data from the Whitehead Institute/MIT Center for Genome Research, Human Genetic Mapping Project, data release 11.9 (May 1997).
McPherron, A.C. & Lee, S.-J. The transforming growth factor β superfamily. in Growth Factors and Cytokines in Health and Disease, vol 1B (eds LeRoith, D. & Bondy, C.) 357–393 (Jai Publishing, Greenwich, Connecticut, 1996).
Dunner, S. et al. Towards interbreed IBD fine mapping of the mh locus: double-muscling in the Asturiana de los Valles breed involves the same locus as in the Belgian Blue cattle breed. Mamm. Genome 8, 430–435 (1997).
Georges, M. et al. Mapping quantitative trait loci controlling milk production by exploiting progeny testing. Genetics 139, 907–920 (1995).
Lathrop, M. & Lalouel, J.M. Easy calculations of lodscores and genetic risk on small computers. Am. J. Hum. Genet. 36, 460–465 (1984).
Cottingham, R.W. Jr., Idury, R.M. & Schäffer, A.A. Faster sequential genetic linkage computations. Am. J. Hum. Genet. 53, 252–263 (1993).
Libert, F., Lefort, A., Okimoto, R., Womack, J. & Georges, M. Construction of a bovine genomic library of large yeast artificial chromosome clones. Genomics 18, 270–276 (1993).
Hunter, K.W. et al. Toward the construction of integrated physical and genetic maps of the mouse genome using interspersed repetitive sequence PCR (IRS–PCR) genomics. Genome Res. 6, 290–299 (1996).
Lenstra, J.A., van Boxtel, J.A.F., Zwaagstra, K.A. & Schwerin, M. Short interspersed nuclear element (SINE) sequences of the Bovidae. Anim. Genet. 24, 33–39 (1993).
Cornélis, F. et al. Identification of a CA repeat at the TCRA locus using yeast artificial chromosomes: a general method for generating highly polymorphic markers at chosen loci. Genomics 13, 820–825 (1992).
Chirgwin, J.M., Przybyla, A.E., MacDonald, R.J. & Rutter, W.J. Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry 18, 5294–5299 (1979).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Grobet, L., Royo Martin, L., Poncelet, D. et al. A deletion in the bovine myostatin gene causes the double–muscled phenotype in cattle. Nat Genet 17, 71–74 (1997). https://doi.org/10.1038/ng0997-71
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0997-71
This article is cited by
-
Sequence-based GWAS meta-analyses for beef production traits
Genetics Selection Evolution (2023)
-
Sequenced-based GWAS for linear classification traits in Belgian Blue beef cattle reveals new coding variants in genes regulating body size in mammals
Genetics Selection Evolution (2023)
-
Fast, multiplexable and efficient somatic gene deletions in adult mouse skeletal muscle fibers using AAV-CRISPR/Cas9
Nature Communications (2023)
-
Comparison of nutritional quality, flesh quality, muscle cellularity, and expression of muscle growth-related genes between wild and recirculating aquaculture system (RAS)-farmed black rockfish (Sebastes schlegelii)
Aquaculture International (2023)
-
Characterisation of Single Nucleotide Polymorphisms and Haplotypes of MSTN Associated with Growth Traits in European Sea Bass (Dicentrarchus labrax)
Marine Biotechnology (2023)