Abstract
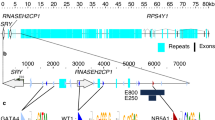
SRY, the mammalian Y-chromosomal sex-determining gene1,2,3, encodes a protein characterized by a DNA-binding and -bending domain referred to as the HMG box4,5. Despite the pivotal role of this gene, only the HMG box region has been conserved through evolution6,7, suggesting that SRY function depends solely on the HMG box and therefore acts as an architectural transcription factor8. In mice (genus Mus) Sry also includes a large CAG trinucleotide repeat region encoding a carboxy-terminal glutamine-rich domain that acts as a transcriptional trans-activator in vitro9. The absence of this or any other potential trans-activating domain in other mammals1,9, however, has raised doubts as to its biological relevance. To test directly whether the glutamine-rich region is required for Sry function in vivo, we created truncation mutations of the Mus musculus musculus Sry gene and tested their ability to induce testis formation in XX embryos using a transgenic mouse assay. Sry constructs that encode proteins lacking the glutamine-rich region were unable to effect male sex determination, in contrast to their wild-type counterparts. We conclude that the glutamine-rich repeat domain of the mouse Sry protein has an essential role in sex determination in vivo, and that Sry may act via a fundamentally different biochemical mechanism in mice compared with other mammals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Sinclair, A.H. et al. A gene from the human sex-determining region encodes a protein with homology to a conserved DNA-binding motif. Nature 346, 240–244 (1990).
Gubbay, J. et al. A gene mapping to the sex-determining region of the mouse Y chromosome is a member of a novel family of embryonically expressed genes. Nature 346, 245–250 (1990).
Koopman, P., Gubbay, J., Vivian, N., Goodfellow, P. & Lovell-Badge, R. Male development of chromosomally female mice transgenic for Sry. Nature 351, 117–121 (1991).
Ferrari, S. et al. SRY, like HMG1, recognises sharp angles in DNA. EMBO J. 11, 4497–4509 (1992).
Harley, V.R. et al. DNA binding activity of recombinant SRY from normal males and XY Females. Science 255, 453–456 (1992).
Tucker, P.K. & Lundrigan, B.L. Rapid evolution of the sex determining locus in Old World mice and rats. Nature 364, 715–717 (1993).
Whitfield, L.S., Lovell-Badge, R. & Goodfellow, P.N. Rapid sequence evolution of the mammalian sex determining gene SRY. Nature 364, 713–715 (1993).
Grosschedl, R., Giese, K. & Pagel, J. HMG domain proteins: architectural elements in the assembly of nucleoprotein structures. Trends Genet. 10, 94–100 (1994).
Dubin, R.A. & Ostrer, H. Sry is a transcriptional activator. Mol. Endocrinol. 8, 1182–1192 (1994).
Gubbay, J. et al. Inverted repeat structure of the Sry locus in mice. Proc. Natl Acad. Sci. USA 89, 7953–7957 (1992).
Miller, K.E., Lundrigan, B.L. & Tucker, P.K. Length variations of CAG repeats in Sry across populations of Mus domesticus. Mamm. Genome 6, 206–208 (1995).
Albrecht, K.H. & Eicher, E.M. DNA sequence analysis of Sry alleles (Subgenus Mus) implicates misregulation as the cause of C57BL/6J-YPOS sex reversal and defines the SRY functional unit. Genetics 147, 1267–1277 (1997).
Koopman, P., Münsterberg, A., Capel, B., Vivian, N. & Lovell-Badge, R. Expression of a candidate sex-determining gene during mouse testis differentiation. Nature 348, 450–452 (1990).
Pontiggia, A. et al. Sex-reversing mutations affect the architecture of SRY-DNA complexes. EMBO J. 13, 6115–6124 (1994).
Kamachi, Y., Cheah, K.S. & Kondoh, H. Mechanism of regulatory target selection by the SOX high-mobility-group domain proteins as revealed by comparison of SOX1/2/3 and SOX9. Mol. Cell. Biol. 19, 107–120 (1999).
Südbeck, P. & Scherer, G. Two independent nuclear localization signals are present in the DNA-binding high-mobility group domains of SRY and SOX9. J. Biol. Chem. 272, 27848–27852 (1997).
Coward, P. et al. Polymorphism of a CAG repeat within Sry correlates with B6.YDom sex reversal. Nature Genet. 6, 245–250 (1994).
Eicher, E.M., Shown, E.P. & Washburn, L.L. Sex reversal in C57BL/6J-YPOS mice corrected by an Sry transgene. Philos. Trans. R. Soc. Lond. B Biol. Sci. 350, 263–269 (1995).
Reddy, P.S. & Housman, D.E. The complex pathology of trinucleotide repeats. Curr. Opin. Cell Biol. 9, 364–372 (1997).
Perutz, M.F., Johnson, T., Suzuki, M. & Finch, J.T. Glutamine repeats as polar zippers: Their possible role in inherited neurodegenerative diseases. Proc. Natl Acad. Sci. USA 91, 5355–5358 (1994).
Tanaka, M., Clouston, W.M. & Herr, W. The Oct-2 glutamine-rich and proline-rich activation domains can synergize with each other or duplicates of themselves to activate transcription. Mol. Cell. Biol. 14, 6046–6055 (1994).
Lau, Y.-F.C. & Zhang, J. Sry interactive proteins: implications for the mechanisms of sex determination. Cytogenet. Cell. Genet. 80, 128–132 (1998).
Latchman, D.S. Eukaryotic Transcription Factors (Academic Press, London, 1998).
Krawczak, M. & Cooper, D.N. The Human Gene Mutation Database. Trends Genet. 13, 121–122 (1997).
Poulat, F. et al. The human testis determining factor SRY binds a nuclear factor containing PDZ protein interaction domains. J. Biol. Chem. 14, 7167–7172 (1997).
Mann, J.R. & McMahon, A.P. Factors influencing frequency production of transgenic mice. in Guide to Techniques in Mouse Development (eds Wassarman, P.M. & DePamphilis, M.L.) 771–781 (Academic Press, San Diego, 1993).
Burgoyne, P.S., Tam, P.P.L. & Evans, E.P. Retarded development of XO conceptuses during early pregnancy in the mouse. J. Reprod. Fertil. 68, 387–393 (1983).
Chomczynski, P. & Sacchi, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162, 156–159 (1987).
Jeske, Y.W.A., Bowles, J., Greenfield, A. & Koopman, P. Expression of a linear Sry transcript in the mouse genital ridge. Nature Genet. 10, 480–482 (1995).
Schreiber, E., Matthias, P., Muller, M.M. & Schaffner, W. Rapid detection of octamer binding proteins with 'mini-extracts' prepared from a small number of cells. Nucleic Acids Res. 17, 6419 (1989).
Acknowledgements
We thank R. Lovell-Badge for providing plasmid L741 and for helpful discussions; C. Nagamine for comments on the manuscript; anonymous referees for constructive suggestions; R. Behringer, R. Krumlauf, A. Greenfield, S. Wheatley, Y. Kanai, M. Azuma-Kanai, D. Pennisi and M. Hargrave for advice and encouragement; and L. Kelly and A. Hardacre for supply of mice. This work was supported by an NHMRC Postdoctoral Research Fellowship to J. Bowles and research grants from the NHMRC and the Australian Research Council.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bowles, J., Cooper, L., Berkman, J. et al. Sry requires a CAG repeat domain for male sex determination in Mus musculus. Nat Genet 22, 405–408 (1999). https://doi.org/10.1038/11981
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/11981
This article is cited by
-
Reduced Activity of SRY and its Target Enhancer Sox9-TESCO in a Mouse Species with X*Y Sex Reversal
Scientific Reports (2017)
-
Aberrant activation of the human sex-determining gene in early embryonic development results in postnatal growth retardation and lethality in mice
Scientific Reports (2017)
-
Production of Sry knockout mouse using TALEN via oocyte injection
Scientific Reports (2013)
-
Mammalian sex determination—insights from humans and mice
Chromosome Research (2012)
-
SRY protein function in sex determination: thinking outside the box
Chromosome Research (2012)