Abstract
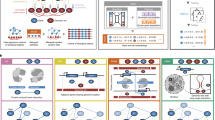
Technologies to measure whole-genome mRNA abundances1,2,3 and methods to organize and display such data4,5,6,7,8,9,10 are emerging as valuable tools for systems-level exploration of transcriptional regulatory networks. For instance, it has been shown that mRNA data from 118 genes, measured at several time points in the developing hindbrain of mice, can be hierarchically clustered into various patterns (or 'waves') whose members tend to participate in common processes5. We have previously shown that hierarchical clustering can group together genes whose cis-regulatory elements are bound by the same proteins in vivo6. Hierarchical clustering has also been used to organize genes into hierarchical dendograms on the basis of their expression across multiple growth conditions7. The application of Fourier analysis to synchronized yeast mRNA expression data has identified cell-cycle periodic genes, many of which have expected cis-regulatory elements8. Here we apply a systematic set of statistical algorithms, based on whole-genome mRNA data, partitional clustering and motif discovery, to identify transcriptional regulatory sub-networks in yeast—without any a priori knowledge of their structure or any assumptions about their dynamics. This approach uncovered new regulons (sets of co-regulated genes) and their putative cis-regulatory elements. We used statistical characterization of known regulons and motifs to derive criteria by which we infer the biological significance of newly discovered regulons and motifs. Our approach holds promise for the rapid elucidation of genetic network architecture in sequenced organisms in which little biology is known.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Velculescu, V.E. et al. Characterization of the yeast transcriptome. Cell 88, 243–251 ( 1997).
Lockhart, D.J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nature Biotechnol. 14, 1675– 1680 (1996).
DeRisi, J.L., Iyer, V.R. & Brown, P.O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680– 686 (1997).
Weinstein, J.N. et al. An information-intensive approach to the molecular pharmacology of cancer. Science 275, 343– 349 (1997).
Wen, X. et al. Large-scale temporal gene expression mapping of central nervous system development. Proc. Natl Acad. Sci. USA 95, 334–339 (1998).
Tavazoie, S. & Church, G.M. Quantitative whole-genome analysis of DNA-protein interactions by in vivo methylase protection in E. coli. Nature Biotechnol. 16, 566– 571 (1998).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863– 14868 (1998).
Spellman, P.T. et al. Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiase by microarray hybridization. Mol. Biol. Cell 9, 3273– 3297 (1998).
Holstege, F.C. et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95, 717–728 (1998).
Tamayo, P. et al. Interpreting patterns of gene expression with self-organizing maps: methods and application to hematopoietic differentiation. Proc. Natl Acad. Sci. USA 96, 2907– 2912 (1999).
Cho, R.J. et al. A genome wide transcriptional analysis of the mitotic cell cycle. Mol. Cell 2, 65–73 (1998).
Wodicka, L., Dong, H., Mittmann, M., Ho, M.H. & Lockhart, D.J. Genome-wide expression monitoring in Saccharomyces cerevisiae. Nature Biotechnol. 15, 1359 –1366 (1997).
Everitt, B. Cluster Analysis 122 (Heinemann, London, 1974).
Hartigan, J.A. Clustering Algorithms 351 (Wiley, New York, 1975).
Mewes, H.W. et al. Overview of the yeast genome. Nature 387, 7–65 (1997).
Roth, F.P., Hughes, J.D., Estep, P.W. & Church, G.M. Finding DNA-regulatory motifs within unaligned noncoding sequences clustered by whole-genome mRNA quantitation. Nature Biotechnol. 16, 949–945 (1998).
Schneider, T.D. & Stephens, R.M. Sequence logos: a new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100 (1990).
Berg, O.G. & von Hippel, P.H. Selection of DNA-binding sites by regulatory proteins. Statistical-mechanical theory and application to operators and promoters. J. Mol. Biol. 193, 723–750 ( 1987).
Koch, C. & Nasmyth, K. Cell cycle regulated transcription in yeast. Curr. Opin. Cell Biol. 6, 451– 459 (1994).
McInerny, C.J., Partridge, J.F., Mikesell, G.E., Creemer, D.P. & Breeden, L.L. A novel Mcm1-dependent element in the SWI4, CLN3, CDC6, and CDC47 promoters activates M/G1-specific transcription. Genes Dev. 11, 1277–1288 (1997).
Kuo, M. & Grayhack, E. A library of yeast genomic MCM1 binding sites contains genes involved in cell cycle control, cell wall and membrane structure, and metabolism. Mol. Cell. Biol. 14, 348–359 (1994).
Planta, R.J., Goncalves, M. & Mager, W.H. Global regulators of ribosome biosynthesis in yeast. Biochem. Cell Biol. 73, 825– 834 (1995).
Moskovina, E. et al. A search in the genome of Saccharomyces cerevisiae for genes regulated via stress response elements. Yeast 14, 1041–1050 (1998).
Thomas, D. & Surdin-Kerjan, Y. Metabolism of sulfur amino acids in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 61, 503–532 ( 1997).
Acknowledgements
We thank D. Lockhart and L. Wodicka for support, and B. Gewurz, V. Mootha, S. Tavazoie, M. Tavazoie and members of the Church lab, especially P. Estep, R. Mitra, B. Cohen, J. Johnson, M. Bulyk and J. Aach, for discussions and critical readings of the manuscript. This work was supported by the US Department of Energy (grant DE-FG02-87-ER60565), the office of Naval Research and DARPA (grant N00014-97-1-0865), the Lipper Foundation and Hoechst Marion Roussel.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tavazoie, S., Hughes, J., Campbell, M. et al. Systematic determination of genetic network architecture. Nat Genet 22, 281–285 (1999). https://doi.org/10.1038/10343
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/10343
This article is cited by
-
Temporal classification of short time series data
BMC Bioinformatics (2024)
-
Placental transcriptomic signatures of prenatal and preconceptional maternal stress
Molecular Psychiatry (2024)
-
Use of Euclidean distance to evaluate Pistia stratiotes and Eichhornia crassipes as organic fertilizer amendments in Capsicum annuum
Acta Physiologiae Plantarum (2024)
-
High-throughput time series expression profiling of Plasmopara halstedii infecting Helianthus annuus reveals conserved sequence motifs upstream of co-expressed genes
BMC Genomics (2023)
-
Oak stands along an elevation gradient have different molecular strategies for regulating bud phenology
BMC Plant Biology (2023)