Abstract
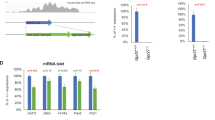
High mobility group 1 (HMG1) protein is an abundant component of all mammalian nuclei, and related proteins exist in all eukaryotes1,2. HMG1 binds linear DNA with moderate affinity and no sequence specificity3, but bends the double helix significantly on binding through the minor groove4. It binds with high affinity to DNA that is already sharply bent, such as linker DNA at the entry and exit of nucleosomes5,6,7; thus, it is considered a structural protein of chromatin. HMG1 is also recruited to DNA by interactions with proteins required for basal and regulated transcription8,9,10,11,12,13 and V(D)J recombination14,15. Here we generate mice harbouring deleted Hmg1. Hmg1–/– pups are born alive, but die within 24 hours due to hypoglycaemia. Hmg1-deficient mice survive for several days if given glucose parenterally, then waste away with pleiotropic defects (but no alteration in the immune repertoire). Cell lines lacking Hmg1 grow normally, but the activation of gene expression by the glucocorticoid receptor (GR, encoded by the gene Grl1) is impaired. Thus, Hmg1 is not essential for the overall organization of chromatin in the cell nucleus, but is critical for proper transcriptional control by specific transcription factors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bianchi, M.E. & Beltrame, M. Flexing DNA: HMG-box proteins and their partners. Am. J. Hum. Genet. 63, 1573 –1577 (1998).
Bustin, M. & Reeves, R. High mobility group chromosomal proteins: Architectural components that facilitate chromatin function. Prog. Nucleic Acid Res. Mol. Biol. 54, 35– 100 (1996).
Bianchi, M.E., Beltrame, M. & Paonessa, G. Specific recognition of cruciform DNA by nuclear protein HMG1. Science 243, 1056– 1059 (1989).
Pil, P.M., Chow, C.S. & Lippard, S.J. High-mobility-group 1 protein mediates DNA bending as determined by ring closures. Proc. Natl Acad. Sci. USA 90, 9465–9469 (1993).
Travers, A.A., Ner, S.S. & Churchill, M.E.A. DNA chaperones: a solution to a persistence problem? Cell 77, 167–169 (1994).
Ura, K., Nightingale, K. & Wolffe, A.P. Differential association of HMG1 and linker histones B4 and H1 with dinucleosomal DNA: structural transitions and transcriptional repression. EMBO J. 15, 4959– 4969 (1996).
Falciola, L. et al. High mobility group 1 (HMG1) protein is not stably associated with the chromosomes of somatic cells. J. Cell Biol. 137, 19–26 (1997).
Shykind, B.M., Kim, J. & Sharp, P.A. Activation of the TFIID-TFIIA complex with HMG-2. Genes Dev. 9, 1354–1365 (1995).
Sutrias-Grau, M., Bianchi, M.E. & Bernués, J. HMG1 interacts with the core domain of human TBP and interferes with TFIIB within the pre-initiation complex. J. Biol. Chem. 274, 1628–1634 ( 1999).
Boonyaaratanakornkit, V. et al. High mobility group chromatin proteins -1 and -2 functionally interact with steroid hormone receptors to enhance their DNA binding in vitro and transcriptional activity in mammalian cells. Mol. Cell. Biol. 18, 4471–4487 ( 1998).
Jayaraman, L. et al. High mobility group protein-1 (HMG1) is a unique activator of p53. Genes Dev. 12, 462– 472 (1998).
Zwilling, S., König, H. & Wirth, T. High mobility group protein 2 functionally interacts with the POU domains of octamer transcription factors. EMBO J. 14, 1198–1208 ( 1995).
Zappavigna, V., Falciola, L., Helmer Citterich, M., Mavilio, F. & Bianchi, M.E. HMG1 cooperates with HOX proteins in DNA binding and transcriptional activation. EMBO J. 15, 4981–4991 (1996).
Agrawal, A. & Schatz, D.G. RAG1 and RAG2 form a stable post-cleavage synaptic complex with DNA containing signal ends in V(D)J recombination. Cell 89, 43–53 ( 1997).
Kwon, J., Imbalzano, A.N., Matthews, A. & Oettinger, M.A. Accessibility of nucleosomal DNA to V(D)J cleavage is modulated by RSS positioning and HMG1. Mol. Cell 2, 829– 839 (1998).
Vaccari, T., Beltrame, M., Ferrari, S. & Bianchi, M.E. Hmg4, a new member of the Hmg1/2 gene family. Genomics 49, 247–252 ( 1998).
Reichardt, H.M. et al. DNA binding of the glucocorticoid receptor is not essential for survival. Cell 93, 531– 541 (1998).
Cole, T.J. et al. Targeted disruption of the glucocorticoid receptor gene blocks adrenergic chromaffin cell development and severely retards lung maturation. Genes Dev. 9, 1608–1621 (1995).
Ruppert, S. et al. Two genetically defined trans-acting loci coordinately regulate overlapping sets of liver-specific genes. Cell 61, 895–904 (1990).
Ferrari, S., Ronfani, L., Calogero, S. & Bianchi, M.E. The mouse gene coding for high mobility group 1 protein (HMG1). J. Biol. Chem. 269, 28803–28808 (1994).
Chomczynski, P. & Sacchi, N. Single-step method for RNA isolation by acid guanidinium thiocyanate-phenol-choloroform extraction. Anal. Biochem. 162, 156– 159 (1987).
Yoo-Warren, H. et al. Identification of a DNA clone to phosphoenolpyruvate carboxykinase (GTP) from rat cytosol. Alterations in phosphoenolpyruvate carboxykinase RNA levels detectable by hybridization. J. Biol. Chem. 256, 10224–10227 (1981).
Noda, C., Tomomura, M., Nakamura, T. & Ichihara, A. Molecular cloning of DNA complementary to mRNA of rat liver serine dehydratase. Biochem. Biophys. Res. Commun. 132, 232 –239 (1985).
Wagner, B.L., Norris, J.D., Knotts, T.A., Weigel, N.L. & McDonnell, D.P. The nuclear corepressors NCoR and SMRT are key regulators of both ligand- and 8-bromo-cyclic AMP-dependent transcriptional activity of the human progesterone receptor. Mol. Cell. Biol. 18, 1369–1378 (1998).
Reue, K., Leff, T. & Breslow, J.L. Human apolipoprotein CIII gene expression is regulated by positive and negative cis-acting elements and tissue-specific protein factors. J. Biol. Chem. 263, 6857– 6864 (1988).
Swat, W., Ignatowicz, L. & Kisielow, P. Detection of apoptosis of immature CD4+8+ thymocytes by flow cytometry. J. Immunol. Methods 137, 79–87 (1991).
Acknowledgements
We thank G. Bouvier and F. Watrin for generating ES mutant clones; A. Plück and the EMBL Transgenic Facility for generating Hmg1 +/- mice on a pure 129Sv genetic background; M. Pezzo for mouse blood tests; A. De Luigi and M.G. De Simoni for corticosterone measurements; L. Ronfani for control experiments; T. Ravasi for establishing cell lines; F. Duranti for personal support; and D. Edwards, F. Barbetti, L. Falqui, P. Dellabona, G. Ferrari, F. Tronche and G. Schütz for help with reagents and suggestions. S.C. was a recipient of fellowships from Universita' di Milano and Centro Italiano Biotecnologie. Supported by Associazione Italiana Ricerca sul Cancro, Ministero dell'Universita' e della Ricerca Scientifica and the European Union (grant FMRX-CT97-0109).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Calogero, S., Grassi, F., Aguzzi, A. et al. The lack of chromosomal protein Hmg1 does not disrupt cell growth but causes lethal hypoglycaemia in newborn mice. Nat Genet 22, 276–280 (1999). https://doi.org/10.1038/10338
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/10338
This article is cited by
-
The multifunctional protein HMGB1: 50 years of discovery
Nature Reviews Immunology (2023)
-
eQTL mapping in fetal-like pancreatic progenitor cells reveals early developmental insights into diabetes risk
Nature Communications (2023)
-
The Seasonal and Stage-Specific Expression Patterns of HMGB2 Suggest Its Key Role in Spermatogenesis in the Chinese Soft-Shelled Turtle (Pelodiscus sinensis)
Biochemical Genetics (2022)
-
Role of High Mobility Group Box 1 in Cardiovascular Diseases
Inflammation (2022)
-
HMGB1 from Astrocytes Promotes EAE by Influencing the Immune Cell Infiltration-Associated Functions of BMECs in Mice
Neuroscience Bulletin (2022)