Abstract
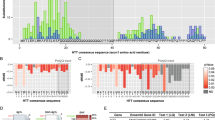
The Huntington's disease (HD) gene encodes a novel protein with as yet no known function. In order to identify the functionally important domains of this protein, we have cloned and sequenced the homologue of the HD gene in the pufferfish, Fugu rubripes. The Fugu HD gene spans only 23 kb of genomic DMA, compared to the 170 kb human gene, and yet all 67 exons are conserved. The first coding exon, the site of the disease–causing triplet repeat, is highly conserved. However, the glutamine repeat in Fugu consists of just four residues. We also show that gene order may be conserved over longer stretches of the two genomes. Our work describes a detailed example of sequence comparison between human and Fugu and illustrates the power of the pufferfish genome as a model system in the analysis of human genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Martin, J.B. & Gusella, J.F. Huntington's disease: pathogenesis and management. New Engl. J. Med. 315, 1267–1276 (1986).
The Hontington's Disease Collabrative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntingtons disease chromosomes. Cell 72, 971–983 (1993).
La Spada, A.R., Wilson, E.M., Luhbahn, D.B., Harding, A.E. & Fischbeck, K.H. Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature 352, 77–79 (1991).
Kremer, E.J. et al. Mapping of DNA instability at the fragile X to a trinucleotide repeat sequence p(CCG)n. Science 252, 1711–1714 (1991).
Verkerk, A.J.M.H. et al. Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell 65, 905–914 (1991).
Brook, J.D. et al. Molecular-basis of Myotonic-Dystrophy — expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a Protein Kinase family member. Cell 68, 799–808 (1992).
Caskey, C.T., Pizzuti, A., Fu, Y.H., Fenwick, R.G. & Nelson, D.L. Triplet repeat mutations in human disease. Science 256, 784–789 (1992).
Fu, Y.H. et al. An unstable triplet repeat in a gene related to Myotonic muscular-dystrophy. Science 255, 1256–1258 (1992).
Koide, R. et al. Unstable expansion of GAG repeat in hereditary dentatorubral-pallidoluysian atrophy (DRPLA). Nature Genet. 6, 9–13 (1994).
Orr, H.T. et al. Expansion of an unstable trinucleotide GAG repeat in spinocerebellar ataxia type 1. Nature Genet. 4, 221–226 (1993).
Mahadevan, M. et al. Myotonic dystrophy mutation — an unstable CTG repeat in the 3′ untranslated region of the gene. Science 255, 1253–1255 (1992).
Burke, J.R. et al. The Haw River-syndrome—dentatorubropallidoluysian atrophy (DRPLA) in an African-American family. Nature Genet. 7, 521–524 (1994).
Kawaguchi, Y. et al. GAG expansions in a novel gene for Machado-Joseph disease at chromosome 14q32.1. Nature Genet. 8, 221–228 (1994).
Ambrose, C.M. et al. Structure and expression of the Huntingtons disease gene—evidence against simple inactivation due to an expanded GAG repeat. Somat. Cell molec. Genet. 20, 27–38 (1994).
Barnes, G.T. et al. Mouse Huntingtons disease gene homolog (Hdh). Somat. Cell molec. Genet. 20, 87–97 (1994).
Lin, B.Y. et al. Sequence of the murine Huntington disease gene — evidence for conservation, and polymorphism in a triplet (CCG) repeat alternate splicing. Hum. molec. Genet. 3, 85–92 (1994).
Brenner, S. et al. Characterization of the pufferfish (Fugu) genome as a compact model vertebrate genome. Nature 366, 265–268 (1993).
Baxendale, S. et al. A cosmid contig and high resolution restriction map of the 2 megabase region containing the Huntingtons disease gene. Nature Genet. 4, 181 (1993).
Nizetic, D. et al. Construction, arraying, and high-density screening of large insert libraries of human chromosomes X and 21: their potential use as reference libraries. Proc. natl. Acad. Sci. U.S.A. 88, 3233–3237 (1991).
Sharp, P.A. Speculations on RNA splicing. Cell 23, 643–646 (1981).
Radley, E., Alderton, R.P., Kelly, A., Trowsdale, J. & Beck, S. Genomic organization of HLA-dma and HLA-dmb — comparison of the gene organization of all 6 class II families in the human major histocompatibility complex. J. biol. Chem. 269, 18834–18838 (1994).
Brown, N.P., Whittaker, A.J., Newell, W.R., Rawlings, C.J. & Beck, S. A novel technique for the identification and analysis of multigene families by comparing intron/exon boundaries. J. molec. Biol. (in the press).
Stine, O.C. et al. Correlation between the onset age of Huntington's disease and length of the trinucleotide repeat in IT-15. Hum. molec. Genet. 2, 1547–1549 (1993).
Barron, L.H. et al. A study of the Huntingtons disease associated trinucleotide repeat in the Scottish population. J. med. Genet. 30, 1003–1007 (1993).
Andrew, S.E. et al. The relationship between trinucleotide (CAG) repeat length and clinical features of Huntingtons disease. Nature Genet. 4, 398–403 (1993).
Rubinsztein, D.C. et al. Mutational bias provides a model for the evolution of Huntingtons disease and predicts a general increase in disease prevalence. Nature Genet. 7, 525–530 (1994).
Perutz, M.F., Johnson, T., Suzuki, M. & Finch, J.T. Glutamine repeats as polar zippers — their possible role in inherited neurodegenerative diseases. Proc. natn. Acad. Sci. U.S.A. 91, 5355–5358 (1994).
Lin, B.Y. et al. Differential 3′ polyadenylation of the Huntington disease gene results in 2 messenger RNA species with variable tissue expression. Hum. molec. Genet. 2, 1541–1545 (1993).
Lin, B. et al. Structural analysis of the 5′ region of mouse and human Huntington Disease genes reveals conservation of putative promoter region and di- and trinucleotide polymorphisms. Genomics 25, 707–715 (1995).
Jurka, J. & Smith, T. A fundamental division in the Alu family of repeated sequences. Proc. natn. Acad. Sci. U.S.A. 85, 4775–4778 (1988).
Beck, S. et al. DNA sequence analysis of 66 kb of the human MHC class II region encoding a cluster of genes for antigen processing. J. molec. Biol. 228, 433–441 (1992).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. molec. Biol. 215, 403–410 (1990).
Snell, R.G. et al. The isolation of cDNAs within the Huntington disease region by hybridization of yeast artificial chromosomes to a cDNA library. Hum. molec. Genet. 2, 305–309 (1993).
Taylor, S.A.M. et al. Cloning of the alpha-adducin gene from the Huntingtons-disease candidate region of chromosome 4 by exon amplification. Nature Genet. 2, 223–227 (1992).
Grosson, C.L.S. et al. Synteny conservation of the Huntingtons disease gene and surrounding loci on mouse chromosome 5. Mamm. Genome 5, 424–428 (1994).
Nasir, J. et al. The murine homologs of the Huntington disease gene (Hdh) and the alpha-adducin gene (ADD1) map to mouse chromosome 5 within a region of conserved synteny with human chromosome 4p16.3. Genomics 22, 198–201 (1994).
Ambrose, C. et al. A novel G protein-coupled receptor kinase cloned from 4p16.3. Hum. molec. Genet. 1, 697–703 (1992).
Koop, B.F. & Hood, L. Striking sequence similarity over almost 100 kilobases of human and mouse T-cell receptor DNA. Nature Genet. 7, 48–53 (1994).
Shehee, W.R. et al. Nucleotide sequence of the BALB/c mouse β-globin complex. J. molec. Biol. 205, 41–62 (1989).
Lindsay, S. & Bird, A.P. Use of restriction enzymes to detect potential gene sequences in mammalian DNA. Nature 327, 336–338 (1987).
Cross, S., Kovarik, P., Schmidtke, J. & Bird, A. Non-methylated islands in fish genomes are GC poor. Nucl. Acids Res. 19, 1469–1474 (1991).
Marshall, H. et al. A conserved retinoic acid response element required for early expression of the homeobox gene hoxb-1. Nature 370, 567–571 (1994).
Speek, M., Raff, J.W., Harrison-Lavoie, K., Little, P.F. & Glover, D.M., Smart2, a cosmid vector with a phage lambda origin for both systematic chromosome walking and P-element-mediated gene transfer in. Drosophila. Gene 64, 173–177 (1988).
Weis, J.H., Tan, S.S., Martin, B.K. & Wittwer, C.T. Detection of rare mRNAs via quantitative RT-PCR. Trends Genet. 8, 263–264 (1992).
Gaudette, M.F. & Crain, W.R. A simple method for quantifying specific mRNAs in small numbers of early mouse embryos. Nucl. Acids Res. 19, 1879–1884 (1991).
Bates, G.P. et al. Defined physical limits of the Huntington disease gene candidate region. Am. J. hum. Genet. 49, 7–16 (1991).
Bankier, A.T., Weston, K.M. & Barrel, B.G. Random cloning and sequencing by the M13/dideoxynucleotide chain termination method. Meth. Enzym. 155, 51–93 (1987).
Beck, S. & Alderton, R.P. A strategy for the amplification, purification, and selection of M13 templates for large-scale DNA-sequencing. Anal. Biochem. 212, 498–505 (1993).
Sanger, F., Nicklen, S. & Coulson, A.R. DNA sequencing with chain-terminating inhibitors. Proc. natn. Acad. Sci. U.S.A. 74, 5463–5467 (1977).
Alderton, R.P. & Beck, S. Automated DNA hybridisation. Anal. Biochem. 218, 98–102 (1994).
Dear, S. & Staden, R. A sequence assembly and editing program for efficient management of large projects. Nucl. Acids Res. 19, 3907–3911 (1991).
Staden, R. in Nucleic Acids and Sequence Analysis: a practical approach. (eds Bishop, M.J. & Rawlings, C.J.). 173–217 (IRL Press Oxford, 1987).
Wilson, R. et al. 2.2 Mb of contiguous nucleotide sequence from chromosome III of C. elegans. Nature 368, 32–38 (1994).
Pearson, W.L. & Lipman, D.J. Improved tools for biological sequence analysis. Proc. natl. Acad. Sci. U.S.A. 85, 2444–2448 (1988).
Sternberg, M.J.E. PROMOT: a FORTRAN program to scan protein sequences against a library of known motifs. Comp. Applic. Biosci. 7, 257–260 (1991).
Fuchs, R. MacPattern — Protein pattern searching on the apple Macintosh Comp. Applic. Biosci. 7, 105–106 (1991).
Jurka, J., Walichiewicz, J. & Milosavljevic, A. Prototypic sequences for human repetitive DNA. J. molec. Evol. 35, 286–291 (1992).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Baxendale, S., Abdulla, S., Elgar, G. et al. Comparative sequence analysis of the human and pufferfish Huntington's disease genes. Nat Genet 10, 67–76 (1995). https://doi.org/10.1038/ng0595-67
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0595-67
This article is cited by
-
Fish genomics and its impact on fundamental and applied research of vertebrate biology
Reviews in Fish Biology and Fisheries (2022)
-
Huntington’s disease: the coming of age
Journal of Genetics (2018)
-
What is a gene for?
Biology & Philosophy (2016)
-
Evidence for dynamic and multiple roles for huntingtin in Ciona intestinalis
Invertebrate Neuroscience (2013)
-
Huntingtin gene evolution in Chordata and its peculiar features in the ascidian Ciona genus
BMC Genomics (2006)