Abstract
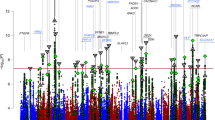
We conducted a combined genome-wide association study (GWAS) of 7,481 individuals with bipolar disorder (cases) and 9,250 controls as part of the Psychiatric GWAS Consortium. Our replication study tested 34 SNPs in 4,496 independent cases with bipolar disorder and 42,422 independent controls and found that 18 of 34 SNPs had P < 0.05, with 31 of 34 SNPs having signals with the same direction of effect (P = 3.8 × 10−7). An analysis of all 11,974 bipolar disorder cases and 51,792 controls confirmed genome-wide significant evidence of association for CACNA1C and identified a new intronic variant in ODZ4. We identified a pathway comprised of subunits of calcium channels enriched in bipolar disorder association intervals. Finally, a combined GWAS analysis of schizophrenia and bipolar disorder yielded strong association evidence for SNPs in CACNA1C and in the region of NEK4-ITIH1-ITIH3-ITIH4. Our replication results imply that increasing sample sizes in bipolar disorder will confirm many additional loci.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Change history
21 June 2012
In the version of this article initially published, there were errors in the consortium membership list and the associated affiliations and in the acknowledgements and contributions sections. These errors and their corrections are detailed below by section. Consortium members: Janice M. Fullerton was omitted from the membership list and has now been added with affiliations 76 and 77. Phil H. Lee was listed incorrectly as Phil L. Hyoun. Fan Meng was listed incorrectly as Fan Guo Meng, and the associated affiliation has been changed from 51 to 54. Robert Thompson was assigned affiliation 50; the correct affiliation is 54. Marian Hamshere and Valentina Moskvina were assigned affiliation 26; the correct affiliation for both is 22. Richard Day was assigned affiliation 47; the correct affiliation is 46. Jun Li was assigned affiliation 24; the correct affiliation is 48. In addition to the affiliation originally listed for Sebastian Zöllner and Peng Zhang, both have now also been assigned affiliation 4. Howard Endenberg has now also been assigned affiliation 12, and Shaun Purcell has now also been assigned affiliation 1. Consortium affiliations: Affiliation 46 was originally given as the University of Dundee School of Medicine, Nethergate, Dundee, UK. The correct affiliation is the Division of Neuroscience, University of Dundee, Ninewells Hospital & Medical School, Dundee, UK. Affiliation 47 was originally given as the School of Neurology, Neurobiology and Psychiatry, Royal Victoria Infirmary, Newcastle upon Tyne, UK. The correct affiliation is the Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA. This affiliation was also listed out of order and has now been changed to affiliation 48. Affiliation 76 was originally given as the Prince of Wales Medical Institute, Sydney, Australia. The correct affiliation is Neuroscience Research Australia, Sydney, Australia. Acknowledgments: The Stanley Foundation for Medical Research was listed as a source of funding. The correct name is the Stanley Medical Research Institute, and the Merck Genome Research Institute has also been added as a source of support. Contributions: In four instances, the contribution of Sven Cichon was indicated with the incorrect spelling S. Chichon instead of S. Cichon. Manuel A. Ferreira (M.A.F.) was incorrectly listed as a contributor to manuscript preparation. Manuel A. Ferreira (M.A.F.) was listed as a contributor to primary study data at the NIMH/Pritzke; the correct contributor was Matthew Flickinger (M.F.). In the section listing contributors to replication data, Neuroscience Research Australia was named incorrectly as the Prince of Wales Medical Institute, and Janice M. Fullerton (J.M.F.) has been added as a contributor at this site and at the University of New South Wales. The errors detailed above have been corrected in the HTML and PDF versions of the article. In addition, the subsections of the contributions section detailing the individuals contributing to primary study data and replication data were omitted from the original HTML version of the article, and this error has now been corrected.
29 August 2012
An Erratum to this paper has been published: https://doi.org/10.1038/ng0912-1072a
References
Craddock, N. & Sklar, P. Genetics of bipolar disorder: successful start to a long journey. Trends Genet. 25, 99–105 (2009).
Cardno, A.G. et al. Heritability estimates for psychotic disorders: the Maudsley twin psychosis series. Arch. Gen. Psychiatry 56, 162–168 (1999).
Craddock, N. & Jones, I. Genetics of bipolar disorder. J. Med. Genet. 36, 585–594 (1999).
Craddock, N., O'Donovan, M.C. & Owen, M.J. The genetics of schizophrenia and bipolar disorder: dissecting psychosis. J. Med. Genet. 42, 193–204 (2005).
Baum, A.E. et al. A genome-wide association study implicates diacylglycerol kinase eta (DGKH) and several other genes in the etiology of bipolar disorder. Mol. Psychiatry 13, 197–207 (2007).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Sklar, P. et al. Whole-genome association study of bipolar disorder. Mol. Psychiatry 13, 558–569 (2008).
Ferreira, M.A. et al. Collaborative genome-wide association analysis supports a role for ANK3 and CACNA1C in bipolar disorder. Nat. Genet. 40, 1056–1058 (2008).
Scott, L.J. et al. Genome-wide association and meta-analysis of bipolar disorder in individuals of European ancestry. Proc. Natl. Acad. Sci. USA 106, 7501–7506 (2009).
Smith, E.N. et al. Genome-wide association study of bipolar disorder in European American and African American individuals. Mol. Psychiatry 14, 755–763 (2009).
Djurovic, S. et al. A genome-wide association study of bipolar disorder in Norwegian individuals, followed by replication in Icelandic sample. J. Affect. Disord. 126, 312–316 (2010).
Cichon, S. et al. Genome-wide association study identifies genetic variation in neurocan as a susceptibility factor for bipolar disorder. Am. J. Hum. Genet. 88, 372–381 (2011).
Schulze, T.G. et al. Two variants in Ankyrin 3 (ANK3) are independent genetic risk factors for bipolar disorder. Mol. Psychiatry 14, 487–491 (2009).
Psychiatric Genome-wide Association Study Consortium. A framework for interpreting genome-wide association studies of psychiatric disorders. Mol. Psychiatry 14, 10–17 (2009).
Cichon, S. et al. Genomewide association studies: history, rationale, and prospects for psychiatric disorders. Am. J. Psychiatry 166, 540–556 (2009).
Browning, S.R. & Browning, B.L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
International Schizophrenia Consortium. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Zhang, D. et al. Singleton deletions throughout the genome increase risk of bipolar disorder. Mol. Psychiatry 14, 376–380 (2009).
Levy, N.A. & Janicak, P.G. Calcium channel antagonists for the treatment of bipolar disorder. Bipolar Disord. 2, 108–119 (2000).
Kempton, M.J. et al. Effects of the CACNA1C risk allele for bipolar disorder on cerebral gray matter volume in healthy individuals. Am. J. Psychiatry 166, 1413–1414 (2009).
Krug, A. et al. Effect of CACNA1C rs1006737 on neural correlates of verbal fluency in healthy individuals. Neuroimage 49, 1831–1836 (2010).
Bigos, K.L. et al. Genetic variation in CACNA1C affects brain circuitries related to mental illness. Arch. Gen. Psychiatry 67, 939–945 (2010).
Erk, S. et al. Brain function in carriers of a genome-wide supported bipolar disorder variant. Arch. Gen. Psychiatry 67, 803–811 (2010).
Nyegaard, M. et al. CACNA1C (rs1006737) is associated with schizophrenia. Mol. Psychiatry 15, 119–121 (2010).
Curtis, D. et al. Case-case genome-wide association analysis shows markers differentially associated with schizophrenia and bipolar disorder and implicates calcium channel genes. Psychiatr. Genet. 21, 1–4 (2011).
Green, E.K. et al. The bipolar disorder risk allele at CACNA1C also confers risk of recurrent major depression and of schizophrenia. Mol. Psychiatry 15, 1016–1022 (2010).
Gomez-Ospina, N., Tsuruta, F., Barreto-Chang, O., Hu, L. & Dolmetsch, R. The C terminus of the L-type voltage-gated calcium channel Ca(V)1.2 encodes a transcription factor. Cell 127, 591–606 (2006).
Lichtenstein, P. et al. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. Lancet 373, 234–239 (2009).
The Schizophrenia Psychiatric Genome-Wide Association Study (GWAS) Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. advance online publication, doi:10.1038/ng.940 (18 September 2011).
Acknowledgements
We would like to recognize the contribution of thousands of subjects without whom this work would not be possible. T. Lehner (National Institute of Mental Health (NIMH)) was instrumental in initiating and planning the overall project. D. Posthuma and the Dutch Genetic Cluster Computer provided invaluable computational resources. We also thank the PGC schizophrenia group for allowing us to perform the combined analyses of six loci before publication. This work was supported by many grants from the US National Institutes of Health (NIH) (MH078151, MH081804, MH059567 supplement, MH59553, MH080372 and 1U54RR025204). Other sources of support include: the Genetic Association Information Network (GAIN), the NIMH Intramural Research Program, the Tzedakah Foundation, the American Philosophical Society, the Stardust foundation, the National Library of Medicine, the Stanley Medical Research Institute, the Merck Genome Research Institute and the Wellcome Trust, the Pritzker Neuropsychiatric Disorders Research Fund L.L.C., GlaxoSmithKline, as well as grants for individual studies (see the Supplemental Note for a full list of Acknowledgements). The TOP Study was supported by grants from the Research Council of Norway (167153/V50, 163070/V50 and 175345/V50), the South-East Norway Health Authority (123-2004) and the EU (ENBREC). Additional acknowledgments can be found in the Supplementary Note.
Genotype data from this manuscript for the 10,257 samples can be obtained from the Center on Collaborative Genetic Studies of Mental Disorders in accordance with NIMH data release policies (http://zork.wustl.edu/nimh/). Genotype data from the WTCCC sample can be obtained from https://www.wtccc.org.uk/info/access_to_data_samples.shtml. Genotype data from the BOMA-Bipolar Study can be obtained by contacting S. Cichon directly (sven.cichon@uni-bonn.de).
Author information
Authors and Affiliations
Consortia
Contributions
Manuscript preparation: P. Sklar. (chair), O.A.A., S. Cichon, N.C., H.J.E., J.R.K., J.I.N., S.M.P., M.R., S. Ripke, L.J.S. Analysis group: S.M.P. (chair), D.B., A.C., N.C., M.F., W.G., J.R.K., M. Mattingsdal, A. McQuillin, P.K., S. Ripke, L.J.S., P. Sklar, T.F.W. PGC central analysis group: S. Ripke (chair), M.D., F.D., P.A.H., D.L. Management group: J.R.K. (co-chair), P. Sklar. (co-chair), O.A.A., D.B., M. Burmeister, S. Cichon, N.C., T.A.G., M.L.H., S.M.P., P. Muglia, J.I.N., L.J.S., E.N.S., P.P.Z.
Primary study data was contributed from the following investigators: BiGS (Bipolar Genome Study) –University of California San Diego: J.R.K., T.A.G., C.M.N., R.M., P.D.S.; Scripps Translational Science Institute: N.J.S., E.N.S., C.S.B.; Indiana University: J.I.N., H.J.E., T.F., D.L.K.; University of Chicago: E.S.G., C.L., J.A.B.; Rush University Medical Center: W.A.S.; Howard University: W.B.L., E.A.N., M. Hipolito; University of Iowa: W.C.; Washington University: J.R.; University of California San Francisco: W. Byerley; National Institute of Mental Health: F.J.M., T.G.S.; University of Pennsylvania: W. Berrettini, F.W.L.; Johns Hopkins Hospital: J.B.P., P.P.Z., P.B. Mahon; University of Michigan: M.G.M., S.Z., P.Z.; The Translational Genomics Research Institute: D.W.C., S. Szelinger; Portland Veterans Affairs Medical Center: T.B.B.; Central Institute of Mental Health, Mannheim: R.B., S.M., M.R., T.G.S., J. Strohmaier, S.H.W.; GlaxoSmithKline, Institute of Psychiatry, Centre for Addiction & Mental Health, University of Dundee: F.T., P. Muglia, A.F., P. McGuffin, J. Strauss, W.X., J.L.K., J.B.V., K. Matthews, R.D.; Massachusetts General Hospital: M.A.F., C.O.'D., R.P., S.M.P., S. Raychaudhuri, P.L.H., D.R., P. Sklar, J.W.S.; NIMH/Pritzker: L.J.S., M.F., M. Burmeister, J. Li, W.G., P.K., D.A., R.C.T., F.G.M., A.F.S., W.E.B., J.D.B., E.G.J., S.J.W., R.M.M., H.A., M. Boehnke; Stanley Center Broad Institute: K.C., J.M., E. Scolnick; TOP Study Group: O.A.A., S.D., M. Mattingsdal, I.M., G.M.; Trinity College Dublin: A.C., M.G., D.M., E.Q.; University of Bonn: S. Cichon, T.W.M., F.A.D., M. Mattheisen, J. Schumacher, W.M., M. Steffens, T.F.W., P.P., M.M.N.; University College London: A.A., N.B., H.G., R.K., J. Lawrence, A. McQuillin; University of Edinburgh: D.B., K. McGhee, A. McIntosh, A.W.M., W.J.M., B.S.P.; Wellcome Trust Case Control Consortium Bipolar Disorder Group—Aberdeen: G.B., D.St.C.; Birmingham: S. Caesar, K.G.-S., L.J.; Cardiff: C.F., E.K.G., D.G., M.L.H., P.A.H., I.R.J., G.K., V.M., I.N., M.C.O., M.J.O., N.C.; London: D.A.C., A.E., A.F., R.W., P. McGuffin; Newcastle: A.H.Y., I.N.F. Replication data contributed by the following investigators: Central Institute of Mental Health, Mannheim: R.B., S.M., M.R., T.G.S., J. Strohmaier, S.H.W.; deCODE genetics: H.S., Þ.Þ., S. Steinberg, Ó.G., K.S.; FaST: S.E.B., K.C., J.M., V.N., S.M.P., P. Sklar, J.W.S.; ICCBD: C.H., M. Landén, P.L., P. Sullivan, M. Schalling, U.O., L.B., L.F., S.E.B., S.M.P., K.C., J.M., N.L., P. Sklar; INSERM U955: S.J., M. Leboyer, B.E., F.B.; Landspitali University: H.P., E. Sigurđsson; Max Planck Institute of Psychiatry, Munich: B.M.-M., S.L.; Psychiatric Center Nordbaden: M. Schwarz; Neuroscience Research Australia and University of New South Wales: P.R.S. and J.M.F.; Queensland Institute of Medical Research: N.M., G.W.M.; Centre National de Génotypage: M. Lathrop; Therapeia University Hospital, Reykjavík: H.Ó.; University of Bonn: S. Cichon, T.W.M., F.A.D., M. Mattheisen, J. Schumacher, W.M., M. Steffens, T.F.W., P.P., M.M.N.; University of Dresden: M. Bauer; University of New South Wales and Black Dog Institute: A.W., P.B. Mitchell; University of Tübingen: M. Hautzinger; University of Würzburg: A.R.
Corresponding author
Ethics declarations
Competing interests
The author declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–6 and Supplementary Tables 1–10. (PDF 3150 kb)
Rights and permissions
About this article
Cite this article
Psychiatric GWAS Consortium Bipolar Disorder Working Group. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat Genet 43, 977–983 (2011). https://doi.org/10.1038/ng.943
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.943
This article is cited by
-
Exploring functional dysconnectivity in schizophrenia: alterations in eigenvector centrality mapping and insights into related genes from transcriptional profiles
Schizophrenia (2024)
-
Blood epigenome-wide association studies of suicide attempt in adults with bipolar disorder
Translational Psychiatry (2024)
-
Progress and Implications from Genetic Studies of Bipolar Disorder
Neuroscience Bulletin (2024)
-
The association of genetic variation in CACNA1C with resting-state functional connectivity in youth bipolar disorder
International Journal of Bipolar Disorders (2023)
-
Brain microvascular endothelial cells and blood-brain barrier dysfunction in psychotic disorders
Molecular Psychiatry (2023)