Abstract
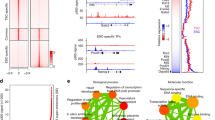
A fundamental challenge in biology is explaining the origin of novel phenotypic characters such as new cell types1,2,3,4; the molecular mechanisms that give rise to novelties are unclear5,6,7. We explored the gene regulatory landscape of mammalian endometrial cells using comparative RNA-Seq and found that 1,532 genes were recruited into endometrial expression in placental mammals, indicating that the evolution of pregnancy was associated with a large-scale rewiring of the gene regulatory network. About 13% of recruited genes are within 200 kb of a Eutherian-specific transposable element (MER20). These transposons have the epigenetic signatures of enhancers, insulators and repressors, directly bind transcription factors essential for pregnancy and coordinately regulate gene expression in response to progesterone and cAMP. We conclude that the transposable element, MER20, contributed to the origin of a novel gene regulatory network dedicated to pregnancy in placental mammals, particularly by recruiting the cAMP signaling pathway into endometrial stromal cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Darwin, C. On the Origin of Species. 6th edn. (Gramercy,1883).
Mayr, E. The emergence of evolutionary novelties. in Evolution after Darwin Vol. 1 (ed. Tax, S.) 349–380 (Harvard Univ. Press, 1960).
Mivart, S.G. On the Genesis of Species (D. Appleton, 1871).
Müller, G.B. & Wagner, G.P. Novelty in evolution: restructuring the concept. Annu. Rev. Ecol. Syst. 22, 229–256 (1991).
Prud'homme, B., Gompel, N. & Carroll, S.B. Emerging principles of regulatory evolution. Proc. Natl. Acad. Sci. USA 104, 8605–8612 (2007).
Carroll, S.B. Evo-devo and an expanding evolutionary synthesis: a genetic theory of morphological evolution. Cell 134, 25–36 (2008).
Wagner, G.P. & Lynch, V.J. Molecular evolution of evolutionary novelties: the vagina and uterus of therian mammals. J. Exp. Zool. B Mol. Dev. Evol. 304, 580–592 (2005).
Mess, A. & Carter, A.M. Evolutionary transformations of fetal membrane characters in Eutheria with special reference to Afrotheria. J. Exp. Zool. B Mol. Dev. Evol. 306, 140–163 (2006).
Wildman, D.E. et al. Evolution of the mammalian placenta revealed by phylogenetic analysis. Proc. Natl. Acad. Sci. USA 103, 3203–3208 (2006).
Gellersen, B. & Brosens, J. Cyclic AMP and progesterone receptor cross-talk in endometrium: a decidualizing affair. J. Endocrinol. 178, 357–372 (2003).
Gellersen, B., Brosens, I.M.D. & Brosens, J.M.D. Decidualization of the human endometrium: mechanisms, functions, and clinical perspectives. Semin. Reprod. Med. 25, 445–453 (2007).
Gerlo, S., Davis, J.R., Mager, D.L. & Kooijman, R. Prolactin in man: a tale of two promoters. Bioessays 28, 1051–1055 (2006).
Bourque, G. et al. Evolution of the mammalian transcription factor binding repertoire via transposable elements. Genome Res. 18, 1752–1762 (2008).
Sasaki, T. et al. Possible involvement of SINEs in mammalian-specific brain formation. Proc. Natl. Acad. Sci. USA 105, 4220–4225 (2008).
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nat. Genet. 42, 631–634 (2010).
Bejerano, G. et al. A distal enhancer and an ultraconserved exon are derived from a novel retroposon. Nature 441, 87–90 (2006).
Jordan, I.K., Rogozin, I.B., Glazko, G.V. & Koonin, E.V. Origin of a substantial fraction of human regulatory sequences from transposable elements. Trends Genet. 19, 68–72 (2003).
van de Lagemaat, L.N., Landry, J.-R., Mager, D.L. & Medstrand, P. Transposable elements in mammals promote regulatory variation and diversification of genes with specialized functions. Trends Genet. 19, 530–536 (2003).
Thornburg, B.G., Gotea, V. & Makalowski, W. Transposable elements as a significant source of transcription regulating signals. Gene 365, 104–110 (2006).
Christian, M. et al. Cyclic AMP-induced forkhead transcription factor, FKHR, cooperates with CCAAT/enhancer-binding protein beta in differentiating human endometrial stromal cells. J. Biol. Chem. 277, 20825–20832 (2002).
Mantena, S.R. et al. C/EEBP-beta is a critical mediator of steroid hormone-regulated cell proliferation and differentiation in the unterine epithelium and stroma. Proc. Natl. Acad. Sci. USA 103, 1870–1875 (2006).
Buzzio, O.L., Lu, Z., Miller, C.D., Unterman, T.G. & Kim, J.J. FOXO1A differentially regulates genes of decidualization. Endocrinology 147, 3870–3876 (2006).
Lynch, V.J. et al. Adaptive changes in the transcription factor HoxA-11 are essential for the evolution of pregnancy in mammals. Proc. Natl. Acad. Sci. USA 105, 14928–14933 (2008).
Hsieh-Li, H.M. et al. Hoxa 11 structure, extensive antisense transcription, and function in male and female fertility. Development 121, 1373–1385 (1995).
Ravasi, T. et al. An atlas of combinatorial transcriptional regulation in mouse and man. Cell 140, 744–752 (2010).
Wei, W. & Brennan, M.D. The gypsy insulator can act as a promoter-specific transcriptional stimulator. Mol. Cell. Biol. 21, 7714–7720 (2001).
Abhyankar, M.M., Urekar, C. & Reddi, P.P. A novel CpG-free vertebrate insulator ilences the testis-specific SP-10 gene in somatic tissues. J. Biol. Chem. 282, 36143–36154 (2007).
Kim, J., Kollhoff, A., Bergmann, A. & Stubbs, L. Methylation-sensitive binding of transcription factor YY1 to an insulator sequence within the paternally expressed imprinted gene, Peg3. Hum. Mol. Genet. 12, 233–245 (2003).
Carroll, S.B. Evolution at two levels: on genes and form. PLoS Biol. 3, e245 (2005).
McClintock, B. Components of action of the regulators Spm and Ac. Year B. Carnegie Inst. Wash. 64, 527–536 (1965).
Britten, R.J. & Davidson, E.H. Gene regulation for higher cells: a theory. Science 165, 349–357 (1969).
Feschotte, C. Transposable elements and the evolution of regulatory networks. Nat. Rev. Genet. 9, 397–405 (2008).
Adamska, M. et al. The evolutionary origin of hedgehog proteins. Curr. Biol. 17, R836–R837 (2007).
Wagner, G.P. & Lynch, V.J. Evolutionary novelties. Curr. Biol. 20, R48–R52 (2010).
Oliver, K.R. & Greene, W.K. Transposable elements: powerful facilitators of evolution. Bioessays 31, 703–714 (2009).
Harti, D. Essential Genetics: A Genomics Perspective (Jones and Bartlett Publishers, 2010).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Wang, Z. et al. Combinatorial patterns of histone acetylations and methylations in the human genome. Nat. Genet. 40, 897–903 (2008).
Acknowledgements
The authors would like to thank A. Pyle and the three anonymous reviewers for comments on an earlier version of this manuscript. We would also like to thank R.W. Truman (National Hansen's Disease Program/US National Institutes of Allergy and Infectious Diseases IAA-2646) and K. Smith for the generous gifts of pregnant armadillo and opossum uterus and R. Bjornson and N. Carriero for assistance with RNA-Seq read mapping. This work was funded by a grant from the John Templeton Foundation, no. 12793, Genetics and the Origin of Organismal Complexity; results presented here do not necessarily reflect the views of the John Templeton Foundation. The funders had no role in study design, data collection and analysis, decision to publish or manuscript preparation.
Author information
Authors and Affiliations
Contributions
V.J.L. and G.P.W. designed experiments and wrote the manuscript. V.J.L. and G.M. performed experiments and analyzed data, and R.D.L. designed and performed bioinformatics analyses.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1 and 2 and Supplementary Figures 1–3 (PDF 599 kb)
Rights and permissions
About this article
Cite this article
Lynch, V., Leclerc, R., May, G. et al. Transposon-mediated rewiring of gene regulatory networks contributed to the evolution of pregnancy in mammals. Nat Genet 43, 1154–1159 (2011). https://doi.org/10.1038/ng.917
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.917
This article is cited by
-
A burst of genomic innovation at the origin of placental mammals mediated embryo implantation
Communications Biology (2023)
-
Comparative analysis of bats and rodents’ genomes suggests a relation between non-LTR retrotransposons, cancer incidence, and ageing
Scientific Reports (2023)
-
Orphan gene expressed in flame cone cells uniquely found in seahorse epithelium
Cell and Tissue Research (2023)
-
Expansion and collapse of VEGF diversity in major clades of the animal kingdom
Angiogenesis (2023)
-
Systematic identification and characterization of repeat sequences in African swine fever virus genomes
Veterinary Research (2022)