Abstract
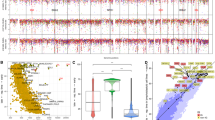
Diffuse large B-cell lymphoma (DLBCL) is the most common form of human lymphoma. Although a number of structural alterations have been associated with the pathogenesis of this malignancy, the full spectrum of genetic lesions that are present in the DLBCL genome, and therefore the identity of dysregulated cellular pathways, remains unknown. By combining next-generation sequencing and copy number analysis, we show that the DLBCL coding genome contains, on average, more than 30 clonally represented gene alterations per case. This analysis also revealed mutations in genes not previously implicated in DLBCL pathogenesis, including those regulating chromatin methylation (MLL2; 24% of samples) and immune recognition by T cells. These results provide initial data on the complexity of the DLBCL coding genome and identify novel dysregulated pathways underlying its pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
19 August 2011
In the version of this supplementary file originally posted online, Supplementary Figures 2, 4 and 6 and Supplementary Tables 12, 14, 18 and 19 contained errors. The errors have been corrected in this file as of 19 August 2011.
References
Abramson, J.S. & Shipp, M.A. Advances in the biology and therapy of diffuse large B-cell lymphoma: moving toward a molecularly targeted approach. Blood 106, 1164–1174 (2005).
Swerdlow, S.H. et al. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues (International Agency for Research on Cancer (IARC), Lyon, France, 2008).
Alizadeh, A.A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
Rosenwald, A. et al. Molecular diagnosis of primary mediastinal B cell lymphoma identifies a clinically favorable subgroup of diffuse large B cell lymphoma related to Hodgkin lymphoma. J. Exp. Med. 198, 851–862 (2003).
Savage, K.J. et al. The molecular signature of mediastinal large B-cell lymphoma differs from that of other diffuse large B-cell lymphomas and shares features with classical Hodgkin lymphoma. Blood 102, 3871–3879 (2003).
Lenz, G. et al. Molecular subtypes of diffuse large B-cell lymphoma arise by distinct genetic pathways. Proc. Natl. Acad. Sci. USA 105, 13520–13525 (2008).
Rosenwald, A. et al. The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N. Engl. J. Med. 346, 1937–1947 (2002).
Morin, R.D. et al. Somatic mutations altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas of germinal-center origin. Nat. Genet. 42, 181–185 (2010).
Savage, K.J. et al. MYC gene rearrangements are associated with a poor prognosis in diffuse large B-cell lymphoma patients treated with R-CHOP chemotherapy. Blood 114, 3533–3537 (2009).
Mandelbaum, J. et al. BLIMP1 is a tumor suppressor gene frequently disrupted in activated B cell-like diffuse large B cell lymphoma. Cancer Cell 18, 568–579 (2010).
Pasqualucci, L. et al. Inactivation of the PRDM1/BLIMP1 gene in diffuse large B cell lymphoma. J. Exp. Med. 203, 311–317 (2006).
Tam, W. et al. Mutational analysis of PRDM1 indicates a tumor-suppressor role in diffuse large B-cell lymphomas. Blood 107, 4090–4100 (2006).
Compagno, M. et al. Mutations of multiple genes cause deregulation of NF-κB in diffuse large B-cell lymphoma. Nature 459, 717–721 (2009).
Davis, R.E. et al. Chronic active B-cell-receptor signalling in diffuse large B-cell lymphoma. Nature 463, 88–92 (2010).
Lenz, G. et al. Oncogenic CARD11 mutations in human diffuse large B cell lymphoma. Science 319, 1676–1679 (2008).
Ngo, V.N. et al. Oncogenically active MYD88 mutations in human lymphoma. Nature 470, 115–119 (2011).
Green, M.R. et al. Integrative analysis reveals selective 9p24.1 amplification, increased PD-1 ligand expression, and further induction via JAK2 in nodular sclerosing Hodgkin lymphoma and primary mediastinal large B-cell lymphoma. Blood 116, 3268–3277 (2010).
Rui, L. et al. Cooperative epigenetic modulation by cancer amplicon genes. Cancer Cell 18, 590–605 (2010).
Steidl, C. et al. MHC class II transactivator CIITA is a recurrent gene fusion partner in lymphoid cancers. Nature 471, 377–381 (2011).
Iqbal, J. et al. Distinctive patterns of BCL6 molecular alterations and their functional consequences in different subgroups of diffuse large B-cell lymphoma. Leukemia 21, 2332–2343 (2007).
Pasqualucci, L. et al. Inactivating mutations of acetyltransferase genes in B-cell lymphoma. Nature 471, 189–195 (2011).
Annunziata, C.M. et al. Frequent engagement of the classical and alternative NF-κB pathways by diverse genetic abnormalities in multiple myeloma. Cancer Cell 12, 115–130 (2007).
Fabbri, G. et al. Analysis of the chronic lymphocytic leukemia coding genome: role of NOTCH1 mutational activation. J. Exp. Med. 208, 1389–1401 (2011).
Keats, J.J. et al. Promiscuous mutations activate the noncanonical NF-κB pathway in multiple myeloma. Cancer Cell 12, 131–144 (2007).
Mendez-Lago, M. et al. Mutations in MLL2 and MEF2B genes in follicular lymphoma and diffuse large B-cell lymphoma. Blood (ASH Annual Meeting Abstracts) 116–473 (2010).
Parsons, D.W. et al. The genetic landscape of the childhood cancer medulloblastoma. Science 331, 435–439 (2011).
Puente, X.S. et al. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature 475, 101–105 (2011).
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007).
Cigudosa, J.C. et al. Cytogenetic analysis of 363 consecutively ascertained diffuse large B-cell lymphomas. Genes Chromosom. Cancer 25, 123–133 (1999).
Beroukhim, R. et al. Assessing the significance of chromosomal aberrations in cancer: methodology and application to glioma. Proc. Natl. Acad. Sci. USA 104, 20007–20012 (2007).
Prasad, R. et al. Structure and expression pattern of human ALR, a novel gene with strong homology to ALL-1 involved in acute leukemia and to Drosophila trithorax. Oncogene 15, 549–560 (1997).
Sneeringer, C.J. et al. Coordinated activities of wild-type plus mutant EZH2 drive tumor-associated hypertrimethylation of lysine 27 on histone H3 (H3K27) in human B-cell lymphomas. Proc. Natl. Acad. Sci. USA 107, 20980–20985 (2010).
Yap, D.B. et al. Somatic mutations at EZH2 Y641 act dominantly through a mechanism of selectively altered PRC2 catalytic activity, to increase H3K27 trimethylation. Blood 117, 2451–2459 (2011).
Dou, Y. et al. Physical association and coordinate function of the H3 K4 methyltransferase MLL1 and the H4 K16 acetyltransferase MOF. Cell 121, 873–885 (2005).
Ernst, P., Wang, J., Huang, M., Goodman, R.H. & Korsmeyer, S.J. MLL and CREB bind cooperatively to the nuclear coactivator CREB-binding protein. Mol. Cell Biol. 21, 2249–2258 (2001).
Flanagan, J.F. et al. Double chromodomains cooperate to recognize the methylated histone H3 tail. Nature 438, 1181–1185 (2005).
Li, H. et al. Molecular basis for site-specific read-out of histone H3K4me3 by the BPTF PHD finger of NURF. Nature 442, 91–95 (2006).
Wysocka, J. et al. A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442, 86–90 (2006).
Cresswell, P., Ackerman, A.L., Giodini, A., Peaper, D.R. & Wearsch, P.A. Mechanisms of MHC class I-restricted antigen processing and cross-presentation. Immunol. Rev. 207, 145–157 (2005).
Moingeon, P. et al. CD2-mediated adhesion facilitates T lymphocyte antigen recognition function. Nature 339, 312–314 (1989).
Wang, C., Lin, G.H., McPherson, A.J. & Watts, T.H. Immune regulation by 4–1BB and 4–1BBL: complexities and challenges. Immunol. Rev. 229, 192–215 (2009).
Middendorp, S. et al. Mice deficient for CD137 ligand are predisposed to develop germinal center-derived B-cell lymphoma. Blood 114, 2280–2289 (2009).
Lenz, G. & Staudt, L.M. Mechanisms of disease: aggressive lymphomas. N. Engl. J. Med. 362, 1417–1429 (2010).
Davis, R.E., Brown, K.D., Siebenlist, U. & Staudt, L.M. Constitutive nuclear factor κB activity is required for survival of activated B cell-like diffuse large B cell lymphoma cells. J. Exp. Med. 194, 1861–1874 (2001).
Saito, M. et al. A signaling pathway mediating downregulation of BCL6 in germinal center B cells is blocked by BCL6 gene alterations in B cell lymphoma. Cancer Cell 12, 280–292 (2007).
Phan, R.T. & Dalla-Favera, R. The BCL6 proto-oncogene suppresses p53 expression in germinal-centre B cells. Nature 432, 635–639 (2004).
Phan, R.T., Saito, M., Basso, K., Niu, H. & Dalla-Favera, R. BCL6 interacts with the transcription factor Miz-1 to suppress the cyclin-dependent kinase inhibitor p21 and cell cycle arrest in germinal center B cells. Nat. Immunol. 6, 1054–1060 (2005).
Potthoff, M.J. & Olson, E.N. MEF2: a central regulator of diverse developmental programs. Development 134, 4131–4140 (2007).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Pleasance, E.D. et al. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature 463, 184–190 (2010).
Chapman, M.A. et al. Initial genome sequencing and analysis of multiple myeloma. Nature 471, 467–472 (2011).
Ley, T.J. et al. DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 456, 66–72 (2008).
Radtke, I. et al. Genomic analysis reveals few genetic alterations in pediatric acute myeloid leukemia. Proc. Natl. Acad. Sci. USA 106, 12944–12949 (2009).
Walter, M.J. et al. Acquired copy number alterations in adult acute myeloid leukemia genomes. Proc. Natl. Acad. Sci. USA 106, 12950–12955 (2009).
Pasqualucci, L. et al. Hypermutation of multiple proto-oncogenes in B-cell diffuse large-cell lymphomas. Nature 412, 341–346 (2001).
Ng, S.B. et al. Exome sequencing identifies MLL2 mutations as a cause of Kabuki syndrome. Nat. Genet. 42, 790–793 (2010).
Krivtsov, A.V. & Armstrong, S.A. MLL translocations, histone modifications and leukaemia stem-cell development. Nat. Rev. Cancer 7, 823–833 (2007).
Dalgliesh, G.L. et al. Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 463, 360–363 (2010).
Jordanova, E.S., Riemersma, S.A., Philippo, K., Schuuring, E. & Kluin, P.M. Beta2-microglobulin aberrations in diffuse large B-cell lymphoma of the testis and the central nervous system. Int. J. Cancer 103, 393–398 (2003).
Riemersma, S.A. et al. Extensive genetic alterations of the HLA region, including homozygous deletions of HLA class II genes in B-cell lymphomas arising in immune-privileged sites. Blood 96, 3569–3577 (2000).
Tchernitchko, D., Goossens, M. & Wajcman, H. In silico prediction of the deleterious effect of a mutation: proceed with caution in clinical genetics. Clin. Chem. 50, 1974–1978 (2004).
Lin, M. et al. dChipSNP: significance curve and clustering of SNP-array-based loss-of-heterozygosity data. Bioinformatics 20, 1233–1240 (2004).
Mullighan, C.G. et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 446, 758–764 (2007).
Pounds, S. et al. Reference alignment of SNP microarray signals for copy number analysis of tumors. Bioinformatics 25, 315–321 (2009).
Venkatraman, E.S. & Olshen, A.B. A faster circular binary segmentation algorithm for the analysis of array CGH data. Bioinformatics 23, 657–663 (2007).
Acknowledgements
We would like to thank T. Palomero, E. Tzilianos, V. Miljkovic and the Genomics Technologies Shared Resource of the Herbert Irving Comprehensive Cancer Center at Columbia University; D. Burgess at Roche NimbleGen (Madison, Wisconsin, USA) and B. Boese at 454 Life Sciences (Branford, Connecticut, USA) for assistance with the whole-exome capture and sequencing procedure; K. Basso and A. Holmes for help with the manual curation and analysis of the copy number data; M. Malladi and Y.K. Lieu for help with the analysis of B2M and CD58; and J. Zhang for filtering the list of candidate mutations against the database of germline variants discovered from the St. Jude Children's Research Hospital-Washington University Pediatric Cancer Genome Project. The Affymetrix SNP6.0 array experiments were processed in part at the Affymetrix Research Services Laboratory. Automated DNA sequencing was performed at Genewiz, Inc. This work was supported by US National Institutes of Health (NIH) Grants PO1-CA092625 and RO1-CA37295 (to R.D.-F.), a Specialized Center of Research grant from the Leukemia & Lymphoma Society (to R.D.-F.), NIH Grant CA121852-05, the Northeast Biodefense Center (U54-AI057158) and the National Library of Medicine (1R01LM010140-01) (to R.R.), and the Associazione Italiana per la Ricerca sul Cancro (AIRC) Special Program Molecular Clinical Oncology-5 per mille (Contract No. 10007, Milan, Italy). A. Chiarenza was on leave from the Division of Hematology, Ospedale Ferrarotto, University of Catania, Catania, Italy.
Author information
Authors and Affiliations
Contributions
L.P. and R.D.-F. designed the study and wrote the manuscript. L.P. conducted experiments, analyzed data and supervised the study. G.F., A. Chiarenza, A.G., V.A.W. and M.M. performed PCR amplification and sequencing analysis. C.G.M. and J.M. developed methods for analysis of high-density SNP array data, which was conducted by C.G.M., J.M., L.P. and G.F. Pathologically characterized subject samples were provided by D.R., G.G., G.B. and A. Chadburn. V.T. and R.R. analyzed high-throughput sequencing data and developed the ComFocal algorithm for analysis of copy number data with the help of O.E. and J.C. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–6 and Supplementary Tables 1–19. (PDF 1768 kb)
Rights and permissions
About this article
Cite this article
Pasqualucci, L., Trifonov, V., Fabbri, G. et al. Analysis of the coding genome of diffuse large B-cell lymphoma. Nat Genet 43, 830–837 (2011). https://doi.org/10.1038/ng.892
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.892
This article is cited by
-
Identification of FAT4 as a positive prognostic biomarker in DLBCL by comprehensive genomic analysis
Clinical and Experimental Medicine (2023)
-
Genomic landscape of mature B-cell non-Hodgkin lymphomas — an appraisal from lymphomagenesis to drug resistance
Journal of the Egyptian National Cancer Institute (2022)
-
APR-246 triggers ferritinophagy and ferroptosis of diffuse large B-cell lymphoma cells with distinct TP53 mutations
Leukemia (2022)
-
The genomic landscape of canine diffuse large B-cell lymphoma identifies distinct subtypes with clinical and therapeutic implications
Lab Animal (2022)
-
Complex genetic and histopathological study of 15 patient-derived xenografts of aggressive lymphomas
Laboratory Investigation (2022)