Abstract
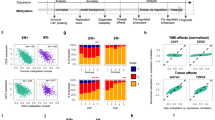
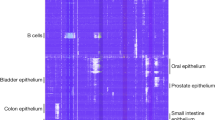
Tumor heterogeneity is a major barrier to effective cancer diagnosis and treatment. We recently identified cancer-specific differentially DNA-methylated regions (cDMRs) in colon cancer, which also distinguish normal tissue types from each other, suggesting that these cDMRs might be generalized across cancer types. Here we show stochastic methylation variation of the same cDMRs, distinguishing cancer from normal tissue, in colon, lung, breast, thyroid and Wilms' tumors, with intermediate variation in adenomas. Whole-genome bisulfite sequencing shows these variable cDMRs are related to loss of sharply delimited methylation boundaries at CpG islands. Furthermore, we find hypomethylation of discrete blocks encompassing half the genome, with extreme gene expression variability. Genes associated with the cDMRs and large blocks are involved in mitosis and matrix remodeling, respectively. We suggest a model for cancer involving loss of epigenetic stability of well-defined genomic domains that underlies increased methylation variability in cancer that may contribute to tumor heterogeneity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jones, P.A. & Baylin, S.B. The fundamental role of epigenetic events in cancer. Nat. Rev. Genet. 3, 415–428 (2002).
Feinberg, A.P. & Tycko, B. The history of cancer epigenetics. Nat. Rev. Cancer 4, 143–153 (2004).
Esteller, M. Epigenetics in cancer. N. Engl. J. Med. 358, 1148–1159 (2008).
Irizarry, R.A. et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nat. Genet. 41, 178–186 (2009).
Doi, A. et al. Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nat. Genet. 41, 1350–1353 (2009).
Irizarry, R.A. et al. Comprehensive high-throughput arrays for relative methylation (CHARM). Genome Res. 18, 780–790 (2008).
Bibikova, M. et al. High-throughput DNA methylation profiling using universal bead arrays. Genome Res. 16, 383–393 (2006).
Feinberg, A.P., Gehrke, C.W., Kuo, K.C. & Ehrlich, M. Reduced genomic 5-methylcytosine content in human colonic neoplasia. Cancer Res. 48, 1159–1161 (1988).
Ehrlich, M. DNA methylation in cancer: too much, but also too little. Oncogene 21, 5400–5413 (2002).
Ogino, S. et al. A cohort study of tumoral LINE-1 hypomethylation and prognosis in colon cancer. J. Natl. Cancer Inst. 100, 1734–1738 (2008).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Wen, B., Wu, H., Shinkai, Y., Irizarry, R.A. & Feinberg, A.P. Large histone H3 lysine 9 dimethylated chromatin blocks distinguish differentiated from embryonic stem cells. Nat. Genet. 41, 246–250 (2009).
Hesselberth, J.R. et al. Global mapping of protein-DNA interactions in vivo by digital genomic footprinting. Nat. Methods 6, 283–289 (2009).
Li, Y. et al. The DNA methylome of human peripheral blood mononuclear cells. PLoS Biol. 8, e1000533 (2010).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
Frigola, J. et al. Epigenetic remodeling in colorectal cancer results in coordinate gene suppression across an entire chromosome band. Nat. Genet. 38, 540–549 (2006).
Gal-Yam, E.N., Saito, Y., Egger, G. & Jones, P.A. Cancer epigenetics: modifications, screening, and therapy. Annu. Rev. Med. 59, 267–280 (2008).
Yu, A.E., Hewitt, R.E., Connor, E.W. & Stetler-Stevenson, W.G. Matrix metalloproteinases. Novel targets for directed cancer therapy. Drugs Aging 11, 229–244 (1997).
Aleman, M.J. et al. Inhibition of Single Minded 2 gene expression mediates tumor-selective apoptosis and differentiation in human colon cancer cells. Proc. Natl. Acad. Sci. USA 102, 12765–12770 (2005).
Yeung, H.Y. et al. Hypoxia-inducible factor-1-mediated activation of stanniocalcin-1 in human cancer cells. Endocrinology 146, 4951–4960 (2005).
Eurich, K., Segawa, M., Toei-Shimizu, S. & Mizoguchi, E. Potential role of chitinase 3-like-1 in inflammation-associated carcinogenic changes of epithelial cells. World J. Gastroenterol. 15, 5249–5259 (2009).
Fischer, H. et al. COL11A1 in FAP polyps and in sporadic colorectal tumors. BMC Cancer 1, 17 (2001).
Clark, S.J. Action at a distance: epigenetic silencing of large chromosomal regions in carcinogenesis. Hum. Mol. Genet. 16, R88–R95 (2007).
Feber, A. et al. Comparative methylome analysis of benign and malignant peripheral nerve sheath tumors. Genome Res. 21, 515–524 (2011).
Feinberg, A.P. & Irizarry, R. Stochastic epigenetic variation as a driving force of development, evolutionary adaptation, and disease. Proc. Natl. Acad. Sci. USA 107, 1757–1764 (2010).
Zilliox, M.J. & Irizarry, R.A. A gene expression bar code for microarray data. Nat. Methods 4, 911–913 (2007).
Bolstad, B.M., Irizarry, R.A., Astrand, M. & Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Leek, J.T. et al. Tackling the widespread and critical impact of batch effects in high-throughput data. Nat. Rev. Genet. 11, 733–739 (2010).
Aryee, M.J. et al. Accurate genome-scale percentage DNA methylation estimates from microarray data. Biostatistics 12, 197–210 (2011).
Bormann Chung, C.A. et al. Whole methylome analysis by ultra-deep sequencing using two-base encoding. PLoS ONE 5, e9320 (2010).
Deng, J. et al. Targeted bisulfite sequencing reveals changes in DNA methylation associated with nuclear reprogramming. Nat. Biotechnol. 27, 353–360 (2009).
Xi, Y. & Li, W. BSMAP: whole genome bisulfite sequence MAPping program. BMC Bioinformatics 10, 232 (2009).
Loader, C. Local Regression and Likelihood (Springer Verlag, New York, New York, USA, 1999).
Eckhardt, F. et al. DNA methylation profiling of human chromosomes 6, 20 and 22. Nat. Genet. 38, 1378–1385 (2006).
Jurka, J. Repbase update: a database and an electronic journal of repetitive elements. Trends Genet. 16, 418–420 (2000).
Olshen, A.B., Venkatraman, E.S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004).
Sabates-Bellver, J. et al. Transcriptome profile of human colorectal adenomas. Mol. Cancer Res. 5, 1263–1275 (2007).
Gyorffy, B., Molnar, B., Lage, H., Szallasi, Z. & Eklund, A.C. Evaluation of microarray preprocessing algorithms based on concordance with RT-PCR in clinical samples. PLoS ONE 4, e5645 (2009).
Galamb, O. et al. Reversal of gene expression changes in the colorectal normal-adenoma pathway by NS398 selective COX2 inhibitor. Br. J. Cancer 102, 765–773 (2010).
Smith, J.C., Boone, B.E., Opalenik, S.R., Williams, S.M. & Russell, S.B. Gene profiling of keloid fibroblasts shows altered expression in multiple fibrosis-associated pathways. J. Invest. Dermatol. 128, 1298–1310 (2008).
Chen, Y. et al. Developing and applying a gene functional association network for anti-angiogenic kinase inhibitor activity assessment in an angiogenesis co-culture model. BMC Genomics 9, 264 (2008).
Duarte, T.L., Cooke, M.S. & Jones, G.D. Gene expression profiling reveals new protective roles for vitamin C in human skin cells. Free Radic. Biol. Med. 46, 78–87 (2009).
Acknowledgements
We thank C. Adams and Applied Biosystems, Inc. for supplying reagents for the sequencing experiments, B. Vogelstein, F. Giardiello and M. Zeiger for tumor samples and M. Newhouse for computer assistance. This work was supported by US National Institutes of Health grants R37CA054358, R01HG005220, 5P50HG003233, F32CA138111, 5R01GM083084 and R01DA025779 (K.Z.).
Author information
Authors and Affiliations
Contributions
K.D.H. and R.A.I. wrote the DMR finder and smoothing algorithms. W.T. performed and analyzed the arrays with H.C.B., who wrote new software for this purpose. S.S. made the libraries and performed validation. B.L. wrote new methylation sequence alignment software. O.G.M. performed the histopathologic analysis. B.W. and H.W. performed LOCK experiments. Y.L. performed copy number experiments. D.D. and K.Z. performed bisulfite capture. E.B. performed the sequencing. R.A.I. and A.P.F. conceived and led the experiments and wrote the paper with the predominant assistance of K.D.H., W.T., H.C.B. and B.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1, 2, 4–6, 8–10 and 13–19 and Supplementary Figures 1–24. (PDF 15293 kb)
Supplementary Table 3
List of block locations (XLS 2912 kb)
Supplementary Table 7
List of small DMRs (XLS 1969 kb)
Supplementary Table 11
List of genes showing statistically significant over-expression in cancer compared to normal samples and are within 2,000 bp from an outward methylation boundary shift. (XLS 263 kb)
Supplementary Table 12
Genes with higher gene expression variability in cancer compared to normal. (XLS 170 kb)
Supplementary Table 20
As Supplementary Table 3, but for sample-specific blocks. (XLS 8811 kb)
Supplementary Table 21
As Supplementary Table 7, but for sample-specific small DMRs. (XLS 5053 kb)
Supplementary Table 22
Primers used for bisulfite pyrosequencing (XLS 35 kb)
Supplementary Table 23
List of microarrays used to identify tissue-specific genes (XLS 87 kb)
Rights and permissions
About this article
Cite this article
Hansen, K., Timp, W., Bravo, H. et al. Increased methylation variation in epigenetic domains across cancer types. Nat Genet 43, 768–775 (2011). https://doi.org/10.1038/ng.865
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.865
This article is cited by
-
DNMT3B PWWP mutations cause hypermethylation of heterochromatin
EMBO Reports (2024)
-
Comprehensive analyses of partially methylated domains and differentially methylated regions in esophageal cancer reveal both cell-type- and cancer-specific epigenetic regulation
Genome Biology (2023)
-
Extensive intratumor regional epigenetic heterogeneity in clear cell renal cell carcinoma targets kidney enhancers and is associated with poor outcome
Clinical Epigenetics (2023)
-
Gene expression variability in long-term survivors of childhood cancer and cancer-free controls in response to ionizing irradiation
Molecular Medicine (2023)
-
Diverse silent chromatin states modulate genome compartmentalization and loop extrusion barriers
Nature Structural & Molecular Biology (2023)