Abstract
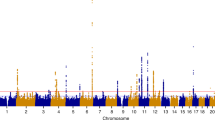
Endometriosis is a common gynecological disease associated with pelvic pain and subfertility. We conducted a genome-wide association study (GWAS) in 3,194 individuals with surgically confirmed endometriosis (cases) and 7,060 controls from Australia and the UK. Polygenic predictive modeling showed significantly increased genetic loading among 1,364 cases with moderate to severe endometriosis. The strongest association signal was on 7p15.2 (rs12700667) for 'all' endometriosis (P = 2.6 × 10−7, odds ratio (OR) = 1.22, 95% CI 1.13–1.32) and for moderate to severe disease (P = 1.5 × 10−9, OR = 1.38, 95% CI 1.24–1.53). We replicated rs12700667 in an independent cohort from the United States of 2,392 self-reported, surgically confirmed endometriosis cases and 2,271 controls (P = 1.2 × 10−3, OR = 1.17, 95% CI 1.06–1.28), resulting in a genome-wide significant P value of 1.4 × 10−9 (OR = 1.20, 95% CI 1.13–1.27) for 'all' endometriosis in our combined datasets of 5,586 cases and 9,331 controls. rs12700667 is located in an intergenic region upstream of the plausible candidate genes NFE2L3 and HOXA10.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Giudice, L.C. & Kao, L.C. Endometriosis. Lancet 364, 1789–1799 (2004).
Simoens, S., Hummelshoj, L. & D'Hooghe, T. Endometriosis: cost estimates and methodological perspective. Hum. Reprod. Update 13, 395–404 (2007).
Jones, G.L., Kennedy, S.H. & Jenkinson, C. Health-related quality of life measurement in women with common benign gynecologic conditions: a systematic review. Am. J. Obstet. Gynecol. 187, 501–511 (2002).
Kjerulff, K.H., Erickson, B.A. & Langenberg, P.W. Chronic gynecological conditions reported by US women: findings from the National Health Interview Survey, 1984 to 1992. Am. J. Public Health 86, 195–199 (1996).
Anonymous. Revised American Fertility Society classification of endometriosis: 1985. Fertil. Steril. 43, 351–352 (1985).
Koninckx, P.R., Oosterlynck, D., D'Hooghe, T. & Meuleman, C. Deeply infiltrating tendometriosis is a disease whereas mild endometriosis could be considered a non-disease. Ann. NY Acad. Sci. 734, 333–341 (1994).
Hadfield, R.M., Mardon, H.J., Barlow, D.H. & Kennedy, S.H. Endometriosis in monozygotic twins. Fertil. Steril. 68, 941–942 (1997).
Kennedy, S. The genetics of endometriosis. J. Reprod. Med. 43, 263–268 (1998).
Treloar, S.A., O'Connor, D.T., O'Connor, V.M. & Martin, N.G. Genetic influences on endometriosis in an Australian twin sample. Fertil. Steril. 71, 701–710 (1999).
Zondervan, K.T., Cardon, L.R. & Kennedy, S.H. The genetic basis of endometriosis. Curr. Opin. Obstet. Gynecol. 13, 309–314 (2001).
Simpson, J.L. & Bischoff, F.Z. Heritability and molecular genetic studies of endometriosis. Ann. NY Acad. Sci. 955, 239–251 (2002).
Stefansson, H. et al. Genetic factors contribute to the risk of developing endometriosis. Hum. Reprod. 17, 555–559 (2002).
Zondervan, K.T. et al. Familial aggregation of endometriosis in a large pedigree of rhesus macaques. Hum. Reprod. 19, 448–455 (2004).
Montgomery, G.W.M. et al. The search for genes contributing to endometriosis risk. Hum. Reprod. Update 14, 447–457 (2008).
Treloar, S. et al. The International Endogene Study: a collection of families for genetic research in endometriosis. Fertil. Steril. 78, 679–685 (2002).
Sturm, R.A. et al. A single SNP in an evolutionary conserved region within intron 86 of the HERC2 gene determines human blue-brown eye color. Am. J. Hum. Genet. 82, 424–431 (2008).
Ferreira, M.A. et al. Quantitative trait loci for CD4:CD8 lymphocyte ratio are associated with risk of type 1 diabetes and HIV-1 immune control. Am. J. Hum. Genet. 86, 88–92 (2010).
Yang, J. et al. Common SNPs explain a large proportion of heritability for human height. Nat. Genet. 42, 565–569 (2010).
The International Schizophrenia Consortium et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Hunter, D.J. et al. A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer. Nat. Genet. 39, 870–874 (2007).
Missmer, S.A. et al. Incidence of laparoscopically confirmed endometriosis by demographic, anthropometric, and lifestyle factors. Am. J. Epidemiol. 160, 784–796 (2004).
Kraft, P. Curses--winner?s and otherwise--in genetic epidemiology. Epidemiology 19, 649–651 (2008).
Zondervan, K.T., Cardon, L.R. & Kennedy, S.H. What makes a good case-control study? Design issues for complex traits such as endometriosis. Hum. Reprod. 17, 1415–1423 (2002).
Taylor, H.S., Bagot, C., Kardana, A., Olive, D. & Arici, A. HOX gene expression is altered in the endometrium of women with endometriosis. Hum. Reprod. 14, 1328–1331 (1999).
Wu, Y. et al. Aberrant methylation at HOXA10 may be responsible for its aberrant expression in the endometrium of patients with endometriosis. Am. J. Obstet. Gynecol. 193, 371–380 (2005).
Uno, S. et al. A genome-wide association study identifies genetic variants in the CDKN2BAS locus associated with endometriosis in Japanese. Nat. Genet. 42, 707–710 (2010).
Vainio, S., Heikkilä, M., Kispert, A., Chin, N. & McMahon, A.P. Female development in mammals is regulated by Wnt-4 signalling. Nature 397, 405–409 (1999).
Naillat, F. et al. Wnt4/5a signalling coordinates cell adhesion and entry into meiosis during presumptive ovarian follicle development. Hum. Mol. Genet. 19, 1539–1550 (2010).
Boyer, A. et al. WNT4 is required for normal ovarian follicle development and female fertility. FASEB J. 24, 3010–3025 (2010).
Treloar, S.A. et al. Genomewide linkage study in 1,176 affected sister pair families identifies a significant susceptibility locus for endometriosis on chromosome 10q26. Am. J. Hum. Genet. 77, 365–376 (2005).
McGregor, B. et al. Genetic and environmental contributions to size, color, shape and other characteristics of melanocytic naevi in a sample of adolescent twins. Genet. Epidemiol. 16, 40–53 (1999).
Zhu, G. et al. A major quantitative-trait locus for mole density is linked to the familial melanoma gene CDKN2A: a maximum-likelihood combined linkage and association analysis in twins and their sibs. Am. J. Hum. Genet. 65, 483–492 (1999).
Medland, S.E. et al. Common variants in the trichohyalin gene are associated with straight hair in Europeans. Am. J. Hum. Genet. 85, 750–755 (2009).
Teo, Y.Y. et al. A genotype calling algorithm for the Illumina BeadArray platform. Bioinformatics 23, 2741–2746 (2007).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Hayes, B.J., Visscher, P.M. & Goddard, M.E. Increased accuracy of artificial selection by using the realized relationship matrix. Genet. Res. 91, 47–60 (2009).
Oliehoek, P.A., Windig, J.J., van Arendonk, J.A. & Bijma, P. Estimating relatedness between individuals in general populations with a focus on their use in conservation programs. Genetics 173, 483–496 (2006).
Patterson, H.D. & Thompson, R. Recovery of interblock information when block sizes are unequal. Biometrika 58, 545–554 (1971).
Gilmour, A.R., Gogel, B.J., Cullis, B.R. & Thompson, R. ASReml User Guide Release 2.0. (VSN International, Hemel Hempstead, UK, 2006).
Wellcome Trust Case Control Consortium et al. Association scan of 14,500 nonsynonymous SNPs in four diseases identifies autoimmunity variants. Nat. Genet. 39, 1329–1337 (2007).
Wellcome Trust Case Control Consortium et al. Genome-wide association study of CNVs in 16,000 cases of eight common diseases and 3,000 shared controls. Nature 464, 713–720 (2010).
Vitonis, A.F., Baer, H.J., Hankinson, S.E., Laufer, M.R. & Missmer, S.A. A prospective study of body size during childhood and early adulthood and the incidence of endometriosis. Hum. Reprod. 25, 1325–1334 (2010).
Curhan, G.C. & Taylor, E.N. 24-h uric acid excretion and the risk of kidney stones. Kidney Int. 73, 489–496 (2008).
Zhao, Z.Z. et al. Genetic variation in tumour necrosis factor and lymphotoxin is not associated with endometriosis in an Australian sample. Hum. Reprod. 22, 2389–2397 (2007).
Zhao, Z.Z. et al. Common variation in the fibroblast growth factor receptor 2 gene is not associated with endometriosis risk. Hum. Reprod. 23, 1661–1668 (2008).
Mägi, R. & Morris, A.P. GWAMA: software for genome-wide association meta-analysis. BMC Bioinformatics 11, 288 (2010).
Purcell, S., Cherny, S.S. & Sham, P.C. Genetic Power Calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics 19, 149–150 (2003).
Acknowledgements
We acknowledge with appreciation all the women who participated in the QIMR, OXEGENE and NHS studies. We thank Endometriosis Associations for supporting the study recruitment. We also thank the many hospital directors and staff, gynecologists, general practitioners and pathology services in Australia, the UK and the United States who provided assistance with confirmation of diagnoses. We thank S. Nicolaides and the Queensland Medical Laboratory for pro bono collection and delivery of blood samples and other pathology services for assistance with blood collection.
The QIMR Study was supported by grants from the National Health and Medical Research Council (NHMRC) of Australia (241944, 339462, 389927, 389875, 389891, 389892, 389938, 443036, 442915, 442981, 496610, 496739, 552485 and 552498), the Cooperative Research Centre for Discovery of Genes for Common Human Diseases (CRC), Cerylid Biosciences (Melbourne) and donations from N. Hawkins and S. Hawkins. D.R.N. was supported by the NHMRC Fellowship (339462 and 613674) and the ARC Future Fellowship (FT0991022) schemes. S.M. was supported by NHMRC Career Development Awards (496674, 613705). P.M.V. (442915) and G.W.M. (339446, 619667) were supported by the NHMRC Fellowships Scheme. We thank B. Haddon, D. Smyth, H. Beeby, O. Zheng, B. Chapman and S. Medland for project and database management, sample processing, genotyping and imputation. We thank Brisbane gynecologist D.T. O'Connor for his important role in initiating the early stages of the project and for confirmation of diagnosis and staging of disease from clinical records of many cases, including 251 in these analyses. We are grateful to the many research assistants and interviewers for assistance with the studies contributing to the QIMR collection.
The work presented here was supported by a grant from the Wellcome Trust (WT084766/Z/08/Z) and makes use of WTCCC2 control data generated by the Wellcome Trust Case-Control Consortium. A full list of the investigators who contributed to the generation of these data is available from http://www.wtccc.org.uk. Funding for the WTCCC project was provided by the Wellcome Trust under awards 076113 and 085475. Imputation analyses were conducted using computational resources at the Oxford Supercomputing Centre (OSC). C.A.A. was funded by the Wellcome Trust (WT91745/Z/10/Z). A.P.M. was supported by a Wellcome Trust Senior Research Fellowship. S.H.K. is supported by the Oxford Partnership Comprehensive Biomedical Research Centre with funding from the Department of Health NIHR Biomedical Research Centres funding scheme. K.T.Z. is supported by a Wellcome Trust Research Career Development Fellowship (WT085235/Z/08/Z). We thank L. Cotton, L. Pope, G. Chalk and G. Farmer (University of Oxford). We also thank P. Koninckx (Leuven, Belgium), M. Sillem (Heidelberg, Germany), C. O'Herlihy and M. Wingfield (Dublin, Ireland), M. Moen (Trondheim, Norway), L. Adamyan (Moscow, Russia), E. McVeigh (Oxford, UK), C. Sutton (Guildford, UK), D. Adamson (Palo Alto, California, USA) and R. Batt (Buffalo, New York, USA) for providing diagnostic confirmation.
The Nurse' Health Studies I and II were supported by grants from the National Institutes of Health (NIH) of the United States, NHS1 cohort (primary investigator: S. Hankinson)-P01 CA087969, NHS1 blood cohort (primary investigator, S. Hankinson)-R01 CA049449, NHS1 Breast Cancer GWAS (primary investigator, D. Hunter)-UO1 CA098233, NHS1/NHS2 Kidney Stones GWAS (primary investigator, G. Curhan)-P01 DK070756, NHS2 cohort (primary investigator, W. Willett)-R01 CA050385, NHS2 blood cohort (primary investigator, S. Hankinson)-R01 CA067262, NHS2 endometriosis (primary investigator, S. Missmer)-R01 HD052473 and R01 HD057210. We thank L. Marshall, D. Hunter and R. Barbieri for their contributions to the endometriosis case validation study and B. Egan and L. Ward for surgical records procurement.
Author information
Authors and Affiliations
Contributions
The International Endogene Consortium
Manuscript preparation: J.N.P., C.A.A., D.R.N., S.M., S.H.L., P.M.V., P.K., N.G.M., A.P.M., S.A.T., S.H.K., S.A.M., G.W.M., K.T.Z.
Study conception and design: J.N.P., C.A.A., D.R.N., P.M.V., N.G.M., S.M., A.P.M., S.A.T., S.H.K., S.A.M., G.W.M., K.T.Z.
GWAS data collection, sample preparation and clinical phenotyping: J.N.P., J.L., A.L., F.R., L.W., A.K.H., N.G.M., S.A.T., S.H.K., G.W.M., K.T.Z.
Replication datasets collection and clinical phenotyping: Q.G., P.K., S.A.M.
Replication genotyping: Z.Z.Z., A.K.H., G.W.M.
Data analysis: GWAS analysis subgroup: J.N.P., C.A.A., D.R.N., S.D.G., A.P.M., K.T.Z.; proportion of variance subgroup: S.H.L., P.M.V.; polygenic prediction analysis subgroup: S.M., P.M.V.; replication and meta-analysis subgroup: J.N.P., D.R.N., Q.G., P.K. S.A.M., G.W.M.; imputation: D.R.N., A.P.M.; bioinformatic analysis subgroup: J.N.P., G.W.M., K.T.Z.
Obtaining study funding: S.M., N.G.M., S.A.T., S.H.K., S.A.M., G.W.M., K.T.Z.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplemementary Note, Supplementary Tables 1–4 and Supplementary Figures 1–3 (PDF 941 kb)
Rights and permissions
About this article
Cite this article
Painter, J., Anderson, C., Nyholt, D. et al. Genome-wide association study identifies a locus at 7p15.2 associated with endometriosis. Nat Genet 43, 51–54 (2011). https://doi.org/10.1038/ng.731
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.731
This article is cited by
-
Whole-exome sequencing reveals candidate high-risk susceptibility genes for endometriosis
Human Genomics (2023)
-
Genomic characterisation of the overlap of endometriosis with 76 comorbidities identifies pleiotropic and causal mechanisms underlying disease risk
Human Genetics (2023)
-
The genetic basis of endometriosis and comorbidity with other pain and inflammatory conditions
Nature Genetics (2023)
-
Genetic polymorphisms of Vascular Endothelial Growth Factor (VEGF) associated with endometriosis in Nigerian women
Human Genomics (2021)
-
Integration of genome-wide association study and expression quantitative trait locus mapping for identification of endometriosis-associated genes
Scientific Reports (2021)