Abstract
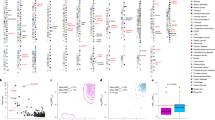
We undertook a meta-analysis of six Crohn's disease genome-wide association studies (GWAS) comprising 6,333 affected individuals (cases) and 15,056 controls and followed up the top association signals in 15,694 cases, 14,026 controls and 414 parent-offspring trios. We identified 30 new susceptibility loci meeting genome-wide significance (P < 5 × 10−8). A series of in silico analyses highlighted particular genes within these loci and, together with manual curation, implicated functionally interesting candidate genes including SMAD3, ERAP2, IL10, IL2RA, TYK2, FUT2, DNMT3A, DENND1B, BACH2 and TAGAP. Combined with previously confirmed loci, these results identify 71 distinct loci with genome-wide significant evidence for association with Crohn's disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Barrett, J.C. et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn's disease. Nat. Genet. 40, 955–962 (2008).
Duerr, R.H. et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science 314, 1461–1463 (2006).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Libioulle, C. et al. Novel Crohn disease locus identified by genome-wide association maps to a gene desert on 5p13.1 and modulates expression of PTGER4. PLoS Genet. 3, e58 (2007).
Imielinski, M. et al. Common variants at five new loci associated with early-onset inflammatory bowel disease. Nat. Genet. 41, 1335–1340 (2009).
McGovern, D.P. et al. Fucosyltransferase 2 (FUT2) non-secretor status is associated with Crohn's disease. Hum. Mol. Genet. 19, 3468–3476 (2010).
Zhernakova, A. et al. Genetic analysis of innate immunity in Crohn's disease and ulcerative colitis identifies two susceptibility loci harboring CARD9 and IL18RAP. Am. J. Hum. Genet. 82, 1202–1210 (2008).
Dixon, A.L. et al. A genome-wide association study of global gene expression. Nat. Genet. 39, 1202–1207 (2007).
Raychaudhuri, S. et al. Identifying relationships among genomic disease regions: predicting genes at pathogenic SNP associations and rare deletions. PLoS Genet. 5, e1000534 (2009).
Rueda, B. et al. The IL23R Arg381Gln non-synonymous polymorphism confers susceptibility to ankylosing spondylitis. Ann. Rheum. Dis. 67, 1451–1454 (2008).
Yamazaki, K. et al. Single nucleotide polymorphisms in TNFSF15 confer susceptibility to Crohn′s disease. Hum. Mol. Genet. 14, 3499–3506 (2005).
Zinovieva, E. et al. Comprehensive linkage and association analyses identify haplotype, near to the TNFSF15 gene, significantly associated with spondyloarthritis. PLoS Genet. 5, e1000528 (2009).
Franke, A. et al. Sequence variants in IL10, ARPC2 and multiple other loci contribute to ulcerative colitis susceptibility. Nat. Genet. 40, 1319–1323 (2008).
Carlsson, B. et al. The G428A nonsense mutation in FUT2 provides strong but not absolute protection against symptomatic GII.4 Norovirus infection. PLoS ONE 4, e5593 (2009).
Gamper, C.J., Agoston, A.T., Nelson, W.G. & Powell, J.D. Identification of DNA methyltransferase 3a as a T cell receptor-induced regulator of Th1 and Th2 differentiation. J. Immunol. 183, 2267–2276 (2009).
El Gazzar, M. et al. G9a and HP1 couple histone and DNA methylation to TNFα transcription silencing during endotoxin tolerance. J. Biol. Chem. 283, 32198–32208 (2008).
Park, J.H. et al. Estimation of effect size distribution from genome-wide association studies and implications for future discoveries. Nat. Genet. 42, 570–575 (2010).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010).
Murray, R.Z., Kay, J.G., Sangermani, D.G. & Stow, J.L. A role for the phagosome in cytokine secretion. Science 310, 1492–1495 (2005).
Luftman, K., Hasan, N., Day, P., Hardee, D. & Hu, C. Silencing of VAMP3 inhibits cell migration and integrin-mediated adhesion. Biochem. Biophys. Res. Commun. 380, 65–70 (2009).
Campbell, B.J., Yu, L.G. & Rhodes, J.M. Altered glycosylation in inflammatory bowel disease: a possible role in cancer development. Glycoconj. J. 18, 851–858 (2001).
McAuley, J.L. et al. MUC1 cell surface mucin is a critical element of the mucosal barrier to infection. J. Clin. Invest. 117, 2313–2324 (2007).
Aoh, Q.L., Castle, A.M., Hubbard, C.H., Katsumata, O. & Castle, J.D. SCAMP3 negatively regulates epidermal growth factor receptor degradation and promotes receptor recycling. Mol. Biol. Cell 20, 1816–1832 (2009).
Mota, L.J., Ramsden, A.E., Liu, M., Castle, J.D. & Holden, D.W. SCAMP3 is a component of the Salmonella-induced tubular network and reveals an interaction between bacterial effectors and post-Golgi trafficking. Cell. Microbiol. 11, 1236–1253 (2009).
Sleiman, P.M. et al. Variants of DENND1B associated with asthma in children. N. Engl. J. Med. 362, 36–44 (2010).
Glocker, E.O. et al. Inflammatory bowel disease and mutations affecting the interleukin-10 receptor. N. Engl. J. Med. 361, 2033–2045 (2009).
Danik, J.S. et al. Novel loci, including those related to Crohn disease, psoriasis, and inflammation, identified in a genome-wide association study of fibrinogen in 17,686 women: the Women's Genome Health Study. Circ. Cardiovasc. Genet. 2, 134–141 (2009).
Rippe, V. et al. Identification of a gene rearranged by 2p21 aberrations in thyroid adenomas. Oncogene 22, 6111–6114 (2003).
Saveanu, L. et al. Concerted peptide trimming by human ERAP1 and ERAP2 aminopeptidase complexes in the endoplasmic reticulum. Nat. Immunol. 6, 689–697 (2005).
Burton, P.R. et al. Association scan of 14,500 nonsynonymous SNPs in four diseases identifies autoimmunity variants. Nat. Genet. 39, 1329–1337 (2007).
Harvey, K.F., Shearwin-Whyatt, L.M., Fotia, A., Parton, R.G. & Kumar, S. N4WBP5, a potential target for ubiquitination by the Nedd4 family of proteins, is a novel Golgi-associated protein. J. Biol. Chem. 277, 9307–9317 (2002).
Putz, U. et al. Nedd4 family-interacting protein 1 (Ndfip1) is required for the exosomal secretion of Nedd4 family proteins. J. Biol. Chem. 283, 32621–32627 (2008).
Xi, H., Schwartz, R., Engel, I., Murre, C. & Kersh, G.J. Interplay between RORγt, Egr3, and E proteins controls proliferation in response to pre-TCR signals. Immunity 24, 813–826 (2006).
Mao, M. et al. T lymphocyte activation gene identification by coregulated expression on DNA microarrays. Genomics 83, 989–999 (2004).
Dendrou, C.A. et al. Cell-specific protein phenotypes for the autoimmune locus IL2RA using a genotype-selectable human bioresource. Nat. Genet. 41, 1011–1015 (2009).
Burchill, M.A., Yang, J., Vang, K.B. & Farrar, M.A. Interleukin-2 receptor signaling in regulatory T cell development and homeostasis. Immunol. Lett. 114, 1–8 (2007).
Stroud, C.K. et al. Disruption of FADS2 gene in mice impairs male reproduction and causes dermal and intestinal ulceration. J. Lipid Res. 50, 1870–1880 (2009).
Loser, K. et al. Epidermal RANKL controls regulatory T-cell numbers via activation of dendritic cells. Nat. Med. 12, 1372–1379 (2006).
Moschen, A.R. et al. The RANKL/OPG system is activated in inflammatory bowel disease and relates to the state of bone loss. Gut 54, 479–487 (2005).
Lu, L. et al. Role of SMAD and non-SMAD signals in the development of Th17 and regulatory T cells. J. Immunol. 184, 4295–4306 (2010).
Ghoreschi, K., Laurence, A. & O'Shea, J.J. Janus kinases in immune cell signaling. Immunol. Rev. 228, 273–287 (2009).
Hazra, A. et al. Common variants of FUT2 are associated with plasma vitamin B12 levels. Nat. Genet. 40, 1160–1162 (2008).
Hindorff, L.A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 106, 9362–9367 (2009).
Yu, W., Clyne, M., Khoury, M.J. & Gwinn, M. Phenopedia and Genopedia: disease-centered and gene-centered views of the evolving knowledge of human genetic associations. Bioinformatics 26, 145–146 (2010).
Rioux, J.D. et al. Genome-wide association study identifies new susceptibility loci for Crohn disease and implicates autophagy in disease pathogenesis. Nat. Genet. 39, 596–604 (2007).
Parkes, M. et al. Sequence variants in the autophagy gene IRGM and multiple other replicating loci contribute to Crohn's disease susceptibility. Nat. Genet. 39, 830–832 (2007).
Fried, L.P. et al. The Cardiovascular Health Study: design and rationale. Ann. Epidemiol. 1, 263–276 (1991).
Browning, B.L. & Browning, S.R. A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am. J. Hum. Genet. 84, 210–223 (2009).
Browning, S.R. & Browning, B.L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Morris, J.A., Randall, J.C., Maller, J.B. & Barrett, J.C. Evoker: a visualization tool for genotype intensity data. Bioinformatics 26, 1786–1787 (2010).
Risch, N.J. Searching for genetic determinants in the new millennium. Nature 405, 847–856 (2000).
Ahmad, T., Satsangi, J., McGovern, D., Bunce, M. & Jewell, D.P. The genetics of inflammatory bowel disease. Aliment. Pharmacol. Ther. 15, 731–748 (2001).
Acknowledgements
We thank all subjects who contributed samples and the physicians and nursing staff who helped with recruitment globally. This study was supported by the German Ministry of Education and Research through the National Genome Research Network and infrastructure support through the Deutsche Forschungsgemeinschaft cluster of excellence 'Inflammation at Interfaces' and by the Italian Ministry for Health GR-2008-1144485, with case collections supported by the Italian Group for IBD and the Italian Society for Paediatric Gastroenterology, Hepatology and Nutrition. We acknowledge funding provided by the Royal Brisbane and Women's Hospital Foundation; the University of Queensland (Ferguson Fellowship); the National Health and Medical Research Council, Australia, and by the European Community (Fifth Framework Program for Research and Development Technology) and by the European Crohn's and Colitis Organization. UK case collections were supported by the National Association for Colitis and Crohn's disease, Action Medical Research, Wellcome Trust, Medical Research Council UK, the University of Edinburgh and the Peninsular College of Medicine and Dentistry, Exeter. We also acknowledge the National Institute of Health Research (NIHR) Biomedical Research Centre awards to Guy's & St. Thomas' National Health Service Trust/King's College London and to Addenbrooke's Hospital/University of Cambridge School of Clinical Medicine. The NIDDK IBD Genetics Consortium is funded by the following grants: DK062431 (S.R.B.), DK062422 (J.H.C.), DK062420 (R.H.D.), DK062432 and DK064869 (J.D.R.), DK062423 (M.S.S.), DK062413 (D.P.B.M.), DK76984 (M.J.D.) and DK084554 (M.J.D. and D.P.B.M.) and DK062429 (J.H.C.). J.H.C. is also funded by the Crohn's and Colitis Foundation of America, and S.L.G. is funded by DK069513 and the Primary Children's Medical Center Foundation. Cedars Sinai was supported by National Center for Research Resources (NCRR) grant M01-RR00425; US National Institutes of Health/NIDDK grant P01-DK046763; DK 063491; and Cedars-Sinai Medical Center Inflammatory Bowel Disease Research Funds. R.K.W. is supported by a clinical fellow grant (90700281) from The Netherlands Organization for Scientific Research. E.L., D.F. and S.V. are senior clinical investigators for the Funds for Scientific Research (FWO/FNRS) Belgium. S.B. was supported by the Deutsche Forschungsgemeinschaft (DFG; BR 1912/5-1). J.C. Barrett is supported by Wellcome Trust grant WT089120/Z/09/Z. Replication genotyping was supported by unrestricted grants from Abbott Laboratories Ltd. and Giuliani SpA. We acknowledge the Wellcome Trust Case Control Consortium. We thank the 1958 British Birth Cohort and Banco Nacional de ADN, Salamanca, Spain, who supplied control DNA samples. The Cardiovascular Health Studies research reported in this article was supported by contract numbers N01-HC-85079 through N01-HC-85086, N01-HC-35129, N01 HC-15103, N01 HC-55222, N01-HC-75150, N01-HC-45133, grant numbers U01 HL080295 and R01 HL087652 from the National Heart, Lung, and Blood Institute, with additional contribution from the National Institute of Neurological Disorders and Stroke. A full list of principal CHS investigators and institutions can be found at http://www.chs-nhlbi.org/pi.htm. Other significant contributors were K. Hanigan, Z.-Z. Zhao, N. Huang, P. Webb, N. Hayward, A. Rutherford, R. Gwilliam, J. Ghori, D. Strachan, W. McCardle, W. Ouwehand, M. Newsky, S. Ehlers, I. Pauselius, K. Holm, C. Sina, L. Baidoo, A. Andriulli and M.C. Renda.
Author information
Authors and Affiliations
Contributions
A.F., D.P.B.M., G.L.R.-S., T.A., J.L., R. Roberts, J.C. Bis, T.H., A. Latiano, C.G.M., N.J.P., J.I.R., P.S., Y.S., L.A.S., K.D.T., D. Whiteman, C.W., G.K.-U., J.D.R., M.D.'A., R.K.W., S.V., R.H.D., J. Satsangi, S.S., V.A., H.H. and M.P. were involved in establishing DNA collections and/or assembling phenotypic data. A.F., D.E., J.C. Barrett, K.W., T.G., S.R., C.A.A., L.J. and M.J.D. performed statistical analyses. D.P.B.M., G.L.R.-S., C.W.L., E.M.F., R.N.B., M.B., T.M.B., S. Brand, C.B., A.C., J.-F.C., M.C., L.S., T.D., M.D.V., R.D.'I., M.D., C.E., T.F., D.F., A.M.G., R.G., J.G., A.V.G., S.L.G., J.H., H.W.V., J.-P.H., A.K., D.L., I.L., M.L., A. Levine, C.L., E.L., C.M., W.N., J.P., A.P., D.D.P., M.R., P.R., R. Russell, J. Satsangi, M.S.S., M.S., F.S., A.H.S., P.C.F.S., S.R.T., L.T., T.W., S.R.B., R.K.W., S.K., A.M.G., J.C.M., S.V., D. Wilson, R.H.D., M.S., J. Sanderson, S.S., J.H.C., V.A. and M.P. recruited patients. A.F., D.P.B.M., T.B., S. Bumpstead, J.I.R., M.G. and G.W.M. supervised laboratory work. A.F., D.P.B.M., J.C. Barrett, K.W., S. Brand, R.H.D., J. Satsangi, S.S., J.H.C., M.J.D. and M.P. contributed to writing the manuscript. All authors read and approved the final manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–6, Supplementary Figures 1–4 and Supplementary Note. (PDF 1562 kb)
Supplementary Table 3
Odds ratios (OR) and risk allele frequencies (RAF) for the 71 SNPs listed in Tables 1 and 2 (XLS 75 kb)
Supplementary Table 4
Raw allele counts and empirical variance for the 71 SNPs listed in Tables 1 and 2 (XLS 170 kb)
Supplementary Table 6
Positional candidate genes mapping within regions of confirmed association with Crohn's disease (XLS 65 kb)
Rights and permissions
About this article
Cite this article
Franke, A., McGovern, D., Barrett, J. et al. Genome-wide meta-analysis increases to 71 the number of confirmed Crohn's disease susceptibility loci. Nat Genet 42, 1118–1125 (2010). https://doi.org/10.1038/ng.717
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.717
This article is cited by
-
Shared genetic architecture between autoimmune disorders and B-cell acute lymphoblastic leukemia: insights from large-scale genome-wide cross-trait analysis
BMC Medicine (2024)
-
Increased CpG methylation at the CDH1 locus in inflamed ileal mucosa of patients with Crohn disease
Clinical Epigenetics (2024)
-
Monogene Varianten in „laccase domain-containing 1“ (LACC1) als Ursache einer juvenilen Arthritis
Zeitschrift für Rheumatologie (2024)
-
Role of pH-sensing receptors in colitis
Pflügers Archiv - European Journal of Physiology (2024)
-
LRRK2 is involved in the chemotaxis of neutrophils and differentiated HL-60 cells, and the inhibition of LRRK2 kinase activity increases fMLP-induced chemotactic activity
Cell Communication and Signaling (2023)