Abstract
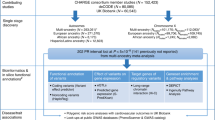
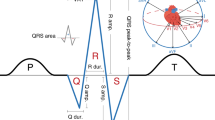
The QRS interval, from the beginning of the Q wave to the end of the S wave on an electrocardiogram, reflects ventricular depolarization and conduction time and is a risk factor for mortality, sudden death and heart failure. We performed a genome-wide association meta-analysis in 40,407 individuals of European descent from 14 studies, with further genotyping in 7,170 additional Europeans, and we identified 22 loci associated with QRS duration (P < 5 × 10−8). These loci map in or near genes in pathways with established roles in ventricular conduction such as sodium channels, transcription factors and calcium-handling proteins, but also point to previously unidentified biologic processes, such as kinase inhibitors and genes related to tumorigenesis. We demonstrate that SCN10A, a candidate gene at the most significantly associated locus in this study, is expressed in the mouse ventricular conduction system, and treatment with a selective SCN10A blocker prolongs QRS duration. These findings extend our current knowledge of ventricular depolarization and conduction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Desai, A.D. et al. Prognostic significance of quantitative QRS duration. Am. J. Med. 119, 600–606 (2006).
Elhendy, A., Hammill, S.C., Mahoney, D.W. & Pellikka, P.A. Relation of QRS duration on the surface 12-lead electrocardiogram with mortality in patients with known or suspected coronary artery disease. Am. J. Cardiol. 96, 1082–1088 (2005).
Oikarinen, L. et al. QRS duration and QT interval predict mortality in hypertensive patients with left ventricular hypertrophy: the Losartan Intervention for Endpoint Reduction in Hypertension Study. Hypertension 43, 1029–1034 (2004).
Dhingra, R. et al. Electrocardiographic QRS duration and the risk of congestive heart failure: the Framingham Heart Study. Hypertension 47, 861–867 (2006).
Busjahn, A. et al. QT interval is linked to 2 long-QT syndrome loci in normal subjects. Circulation 99, 3161–3164 (1999).
Hanson, B. et al. Genetic factors in the electrocardiogram and heart rate of twins reared apart and together. Am. J. Cardiol. 63, 606–609 (1989).
Bezzina, C.R. et al. Common sodium channel promoter haplotype in Asian subjects underlies variability in cardiac conduction. Circulation 113, 338–344 (2006).
Chambers, J.C. et al. Genetic variation in SCN10A influences cardiac conduction. Nat. Genet. 42, 149–152 (2010).
Holm, H. et al. Several common variants modulate heart rate, PR interval and QRS duration. Nat. Genet. 42, 117–122 (2010).
Dubois, P.C. et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 42, 295–302 (2010).
Newton-Cheh, C. et al. Common variants at ten loci influence QT interval duration in the QTGEN Study. Nat. Genet. 41, 399–406 (2009).
Pfeufer, A. et al. Common variants at ten loci modulate the QT interval duration in the QTSCD Study. Nat. Genet. 41, 407–414 (2009).
Pfeufer, A. et al. Genome-wide association study of PR interval. Nat. Genet. 42, 153–159 (2010).
Calvano, S.E. et al. A network-based analysis of systemic inflammation in humans. Nature 437, 1032–1037 (2005).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Zhang, B., Schmoyer, D., Kirov, S. & Snoddy, J. GOTree Machine (GOTM): a web-based platform for interpreting sets of interesting genes using Gene Ontology hierarchies. BMC Bioinformatics 5, 16 (2004).
Pallante, B.A. et al. Contactin-2 expression in the cardiac Purkinje fiber network. Circ. Arrhythm. Electrophysiol. 3, 186–194 (2010).
Jarvis, M.F. et al. A-803467, a potent and selective Nav1.8 sodium channel blocker, attenuates neuropathic and inflammatory pain in the rat. Proc. Natl. Acad. Sci. USA 104, 8520–8525 (2007).
Desplantez, T., Dupont, E., Severs, N.J. & Weingart, R. Gap junction channels and cardiac impulse propagation. J. Membr. Biol. 218, 13–28 (2007).
Abriel, H. Cardiac sodium channel Na(v)1.5 and interacting proteins: physiology and pathophysiology. J. Mol. Cell. Cardiol. 48, 2–11 (2010).
Remme, C.A., Wilde, A.A. & Bezzina, C.R. Cardiac sodium channel overlap syndromes: different faces of SCN5A mutations. Trends Cardiovasc. Med. 18, 78–87 (2008).
Akopian, A.N. et al. The tetrodotoxin-resistant sodium channel SNS has a specialized function in pain pathways. Nat. Neurosci. 2, 541–548 (1999).
Saimi, Y. & Kung, C. Calmodulin as an ion channel subunit. Annu. Rev. Physiol. 64, 289–311 (2002).
Potet, F. et al. Functional interactions between distinct sodium channel cytoplasmic domains through the action of calmodulin. J. Biol. Chem. 284, 8846–8854 (2009).
Wolf, C.M. & Berul, C.I. Inherited conduction system abnormalities—one group of diseases, many genes. J. Cardiovasc. Electrophysiol. 17, 446–455 (2006).
Zhu, Y. et al. Tbx5-dependent pathway regulating diastolic function in congenital heart disease. Proc. Natl. Acad. Sci. USA 105, 5519–5524 (2008).
Lebrec, J.J., Stijnen, T. & van Houwelingen, H.C. Dealing with heterogeneity between cohorts in genomewide SNP association studies. Stat. Appl. Genet. Mol. Biol. 9 article 8 (2010).
Wei, L., Hanna, A.D., Beard, N.A. & Dulhunty, A.F. Unique isoform-specific properties of calsequestrin in the heart and skeletal muscle. Cell Calcium 45, 474–484 (2009).
Terentyev, D. et al. Abnormal interactions of calsequestrin with the ryanodine receptor calcium release channel complex linked to exercise-induced sudden cardiac death. Circ. Res. 98, 1151–1158 (2006).
Priori, S.G. et al. Clinical and molecular characterization of patients with catecholaminergic polymorphic ventricular tachycardia. Circulation 106, 69–74 (2002).
Postma, A.V. et al. Absence of calsequestrin 2 causes severe forms of catecholaminergic polymorphic ventricular tachycardia. Circ. Res. 91, e21–e26 (2002).
Wang, Y. & Goldhaber, J.I. Return of calcium: manipulating intracellular calcium to prevent cardiac pathologies. Proc. Natl. Acad. Sci. USA 101, 5697–5698 (2004).
Vasan, R.S. et al. Genetic variants associated with cardiac structure and function: a meta-analysis and replication of genome-wide association data. J. Am. Med. Assoc. 302, 168–178 (2009).
Eijgelsheim, M. et al. Genome-wide association analysis identifies multiple loci related with resting heart rate. Hum. Mol. Genet. 19, 3885–3895 (2010).
Braz, J.C. et al. PKC-alpha regulates cardiac contractility and propensity toward heart failure. Nat. Med. 10, 248–254 (2004).
Baillat, G. et al. Molecular cloning and characterization of phocein, a protein found from the Golgi complex to dendritic spines. Mol. Biol. Cell 12, 663–673 (2001).
Meurs, K.M. et al. Genome-wide association identifies a deletion in the 3′ untranslated region of Striatin in a canine model of arrhythmogenic right ventricular cardiomyopathy. Hum. Genet. 128, 315–324. (2010).
Boogerd, K.J. et al. Msx1 and Msx2 are functional interacting partners of T-box factors in the regulation of Connexin43. Cardiovasc. Res. 78, 485–493 (2008).
Hoogaars, W.M. et al. The transcriptional repressor Tbx3 delineates the developing central conduction system of the heart. Cardiovasc. Res. 62, 489–499 (2004).
Singh, R. et al. Tbx20 interacts with smads to confine tbx2 expression to the atrioventricular canal. Circ. Res. 105, 442–452 (2009).
Posch, M.G. et al. A gain-of-function TBX20 mutation causes congenital atrial septal defects, patent foramen ovale and cardiac valve defects. J. Med. Genet. 47, 230–235 (2009).
Bakker, M.L. et al. Transcription factor Tbx3 is required for the specification of the atrioventricular conduction system. Circ. Res. 102, 1340–1349 (2008).
Riley, P., Anson-Cartwright, L. & Cross, J.C. The Hand1 bHLH transcription factor is essential for placentation and cardiac morphogenesis. Nat. Genet. 18, 271–275 (1998).
Reamon-Buettner, S.M. et al. A functional genetic study identifies HAND1 mutations in septation defects of the human heart. Hum. Mol. Genet. 18, 3567–3578 (2009).
Breckenridge, R.A. et al. Overexpression of the transcription factor Hand1 causes predisposition towards arrhythmia in mice. J. Mol. Cell. Cardiol. 47, 133–141 (2009).
Rentschler, S. et al. Neuregulin-1 promotes formation of the murine cardiac conduction system. Proc. Natl. Acad. Sci. USA 99, 10464–10469 (2002).
Hofer, A. et al. C-erbB2/neu transfection induces gap junctional communication incompetence in glial cells. J. Neurosci. 16, 4311–4321 (1996).
Besson, A. & Yong, V.W. Involvement of p21(Waf1/Cip1) in protein kinase C alpha-induced cell cycle progression. Mol. Cell. Biol. 20, 4580–4590 (2000).
Wilkinson, L. et al. CRIM1 regulates the rate of processing and delivery of bone morphogenetic proteins to the cell surface. J. Biol. Chem. 278, 34181–34188 (2003).
Kolle, G., Georgas, K., Holmes, G.P., Little, M.H. & Yamada, T. CRIM1, a novel gene encoding a cysteine-rich repeat protein, is developmentally regulated and implicated in vertebrate CNS development and organogenesis. Mech. Dev. 90, 181–193 (2000).
Pardali, K., Kowanetz, M., Heldin, C.H. & Moustakas, A. Smad pathway-specific transcriptional regulation of the cell cycle inhibitor p21(WAF1/Cip1). J. Cell. Physiol. 204, 260–272 (2005).
Laederich, M.B. et al. The leucine-rich repeat protein LRIG1 is a negative regulator of ErbB family receptor tyrosine kinases. J. Biol. Chem. 279, 47050–47056 (2004).
Minakuchi, M. et al. Identification and characterization of SEB, a novel protein that binds to the acute undifferentiated leukemia-associated protein SET. Eur. J. Biochem. 268, 1340–1351 (2001).
Zhao, J. & Zhong, C.J. A review on research progress of transketolase. Neurosci. Bull. 25, 94–99 (2009).
Fedi, P. et al. Isolation and biochemical characterization of the human Dkk-1 homologue, a novel inhibitor of mammalian Wnt signaling. J. Biol. Chem. 274, 19465–19472 (1999).
Ai, Z., Fischer, A., Spray, D.C., Brown, A.M. & Fishman, G.I. Wnt-1 regulation of connexin43 in cardiac myocytes. J. Clin. Invest. 105, 161–171 (2000).
Korol, O., Gupta, R.W. & Mercola, M. A novel activity of the Dickkopf-1 amino terminal domain promotes axial and heart development independently of canonical Wnt inhibition. Dev. Biol. 324, 131–138 (2008).
Tsai, I.C. et al. A Wnt-CKIvarepsilon-Rap1 pathway regulates gastrulation by modulating SIPA1L1, a Rap GTPase activating protein. Dev. Cell 12, 335–347 (2007).
Chen, W.M. & Abecasis, G.R. Family-based association tests for genomewide association scans. Am. J. Hum. Genet. 81, 913–926 (2007).
Devlin, B., Roeder, K. & Wasserman, L. Genomic control, a new approach to genetic-based association studies. Theor. Popul. Biol. 60, 155–166 (2001).
de Bakker, P.I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
Pe'er, I., Yelensky, R., Altshuler, D. & Daly, M.J. Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet. Epidemiol. 32, 381–385 (2008).
Johnson, A.D. et al. SNAP: a web-based tool for identification and annotation of proxy SNPs using HapMap. Bioinformatics 24, 2938–2939 (2008).
Sreejit, P., Kumar, S. & Verma, R.S. An improved protocol for primary culture of cardiomyocyte from neonatal mice. In Vitro Cell. Dev. Biol. Anim. 44, 45–50 (2008).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408 (2001).
Lee, P. et al. Conditional lineage ablation to model human diseases. Proc. Natl. Acad. Sci. USA 95, 11371–11376 (1998).
Acknowledgements
Acknowledgments are available in the Supplementary Note.
Author information
Authors and Affiliations
Contributions
Study concept and design: N.S., A.A., D.E.A., P.I.W.d.B., E.B., H.C., A.C., C.M.v.D., M.E., S.B.F., G.I.F., A.R.F., J.F., V.G., P.v.d.H., S.R.H., A.A.H., A.H., A.I., S.K., H.K.K., C.N.-C., B.A.O., A. Pfeufer, P.P.P., B.M.P., J.I.R., I.R., H.S., E.Z.S., B.H.C.S., A.G.U., A.V.S., U.V., H.V., T.J.W., J.F.W., A.F.W., N.J.S., Y.J.
Acquisition of data: A.A., D.E.A., L.H.v.d.B., R.A.d.B., E.B., M.J.C., A.C., J.M.C., A.F.D., M.D., C.M.v.D., R.S.N.F., A.R.F., L.F., S.G., H.J.M.G., T.B.H., P.v.d.H., C.H., G.v.H., A.I., W.H.L.K., N.K., J.A.K., A.K., L.L., M.L., F.-Y.L., I.M.L., G.t.M., P.B.M., G.N., C.N.-C., B.A.O., R.A.O., S. Perz, A. Pfeufer, A. Petersmann, O.P., B.M.P., J.Q., F.R., J.I.R., I.R., N.J.S., C.S., M.P.S.S., M.F.S., E.Z.S., B.H.C.S., A.T., A.G.U., D.J.v.V., C.B.V., R.K.W., C.W., J.F.W., J.C.M.W., D.L., T.D.S.
Statistical analysis and interpretation of data: A.A., D.E.A., T.A., P.I.W.d.B., N.S., E.B., A.C., L.A.C., M.E., K.E., G.I.F., A.R.F., L.F., J.F., C.F., S.A.G., W.H.v.G., S.G., V.G., P.v.d.H., C.H., S.R.H., A.I., T.J., W.H.L.K., X.L., K.D.M., I.M.L., M.M., I.M.N., S. Padmanabhan, A. Pfeufer, O.P., B.M.P., K.R., H.S., A.T., A.V.S., S.H.W., Y.A.W., N.J.S.
Drafting of the manuscript: N.S., A.A., D.E.A, F.W.A., P.I.W.d.B., M.D., C.M.v.D,. M.E., G.I.F., J.F., S.A.G., V.G., C.H., A.I., Y.J., S.K., J.W.M., I.M.N., O.P., N.J.S., H.S., C.N.-C., P.v.d.H.
Critical revision of the manuscript: A.A., D.E.A., T.A., F.W.A., J.C.B., R.A.d.B., E.B., H.C., M.J.C., A.C., J.M.C., L.A.C., A.F.D., M.D., C.M.v.D., M.E., K.E., S.B.F., G.I.F., A.R.F., J.F., W.H.v.G., V.G., T.B.H., P.v.d.H., C.H., S.R.H., G.v.H., A.A.H., A.H., A.I., Y.J., T.J., S.K., W.H.L.K., N.K., J.A.K., A.K., H.K.K., L.L., D.L., M.L., J.W.M., I.M.L., T.M., M.M., P.B.M., G.N., C.N.-C., I.M.N., C.J.O., B.A.O., S. Padmanabhan, S. Perz, A. Pfeufer, A. Petersmann, O.P., B.M.P., F.R., J.I.R., I.R., M.P.S.S., M.F.S., D.S.S., H.S., B.H.C.S., E.Z.S., A.T., A.G.U., D.J.v.V., U.V., H.V., T.J.W., H.-E.W., A.V.S., S.H.W., J.F.W., J.C.M.W., A.F.W.
Obtained funding: L.H.v.d.B., E.B., H.C., M.J.C., A.C., J.M.C., A.F.D., C.M.v.D., S.B.F., G.I.F., W.H.v.G., H.J.M.G., V.G., P.v.d.H., A.H., Y.J., S.K., H.K.K., L.L., P.B.M., G.N., C.N.-C., C.J.O., B.A.O., R.A.O., P.P.P., B.M.P., J.I.R., I.R., N.J.S., N.S., T.D.S., A.G.U., D.J.v.V., U.V., H.V., T.J.W., R.K.W., H.-E.W., C.W., J.F.W., A.F.W., D.L.
Corresponding authors
Ethics declarations
Competing interests
A.C. is a paid member of the Scientific Advisory Board of Affymetrix, a role that is managed by the Committee on Conflict of Interest of the Johns Hopkins University School of Medicine.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1–5 and Supplementary Note (PDF 2282 kb)
Rights and permissions
About this article
Cite this article
Sotoodehnia, N., Isaacs, A., de Bakker, P. et al. Common variants in 22 loci are associated with QRS duration and cardiac ventricular conduction. Nat Genet 42, 1068–1076 (2010). https://doi.org/10.1038/ng.716
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.716
This article is cited by
-
Phenotypic and Genetic Factors Associated with Absence of Cardiomyopathy Symptoms in PLN:c.40_42delAGA Carriers
Journal of Cardiovascular Translational Research (2023)
-
Genome-wide association analyses identify new Brugada syndrome risk loci and highlight a new mechanism of sodium channel regulation in disease susceptibility
Nature Genetics (2022)
-
Genetics and genomics of arrhythmic risk: current and future strategies to prevent sudden cardiac death
Nature Reviews Cardiology (2021)
-
Association of T66A polymorphism in CASQ2 with PR interval in a Chinese population
Herz (2021)
-
Inhibition of NaV1.8 prevents atrial arrhythmogenesis in human and mice
Basic Research in Cardiology (2020)