Abstract
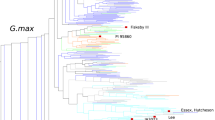
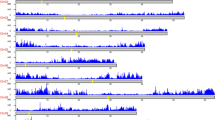
We report a large-scale analysis of the patterns of genome-wide genetic variation in soybeans. We re-sequenced a total of 17 wild and 14 cultivated soybean genomes to an average of approximately ×5 depth and >90% coverage using the Illumina Genome Analyzer II platform. We compared the patterns of genetic variation between wild and cultivated soybeans and identified higher allelic diversity in wild soybeans. We identified a high level of linkage disequilibrium in the soybean genome, suggesting that marker-assisted breeding of soybean will be less challenging than map-based cloning. We report linkage disequilibrium block location and distribution, and we identified a set of 205,614 tag SNPs that may be useful for QTL mapping and association studies. The data here provide a valuable resource for the analysis of wild soybeans and to facilitate future breeding and quantitative trait analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Hymowitz, T. On the domestication of soybean. Econ. Bot. 24, 408–421 (1970).
Hymowitz, T. & Harlan, J.R. Introduction of soybean to North America by Samuel Bowen in 1765. Econ. Bot. 37, 371–379 (1983).
Hyten, D.L. et al. Highly variable patterns of linkage disequilibrium in multiple soybean populations. Genetics 175, 1937–1944 (2007).
Hyten, D.L. et al. Impacts of genetic bottlenecks on soybean genome diversity. Proc. Natl. Acad. Sci. USA 103, 16666–16671 (2006).
Schmutz, J. et al. Genome sequence of the palaeopolyploid soybean. Nature 463, 178–183 (2010).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Li, R. et al. SNP detection for massively parallel whole-genome resequencing. Genome Res. 19, 1124–1132 (2009).
Wang, J. et al. The diploid genome sequence of an Asian individual. Nature 456, 60–65 (2008).
Xia, Q. et al. Complete resequencing of 40 genomes reveals domestication events and genes in silkworm (Bombyx). Science 326, 433–436 (2009).
Pritchard, J.K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Tajima, F. Evolutionary relationship of DNA sequences in finite populations. Genetics 105, 437–460 (1983).
Gutenkunst, R.N., Hernandez, R.D., Williamson, S.H. & Bustamante, C.D. Inferring the joint demographic history of multiple populations from multidimensional SNP frequency data. PLoS Genet. 5, e1000695 (2009).
Hernandez, R.D. et al. Demographic histories and patterns of linkage disequilibrium in Chinese and Indian Rhesus Macaques. Science 316, 240–243 (2007).
Caicedo, A.L. et al. Genome-wide patterns of nucleotide polymorphism in domesticated rice. PLoS Genet. 3, 1745–1756 (2007).
Gore, M.A. et al. A first-generation haplotype map of maize. Science 326, 1115–1117 (2009).
Barrett, J.C., Fry, B., Maller, J. & Daly, M.J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Kim, S. et al. Recombination and linkage disequilibrium in Arabidopsis thaliana. Nat. Genet. 39, 1151–1155 (2007).
Zhu, Q., Zheng, X., Luo, J., Gaut, B.S. & Ge, S. Multilocus analysis of nucleotide variation of Oryza sativa and its wild relatives: severe bottleneck during domestication of rice. Mol. Biol. Evol. 24, 875–888 (2007).
Flint-Garcia, S.A., Thornsberry, J.M. & Buckler, E.S. IV . Structure of linkage disequilibrium in plants. Annu. Rev. Plant Biol. 54, 357–374 (2003).
Gabriel, S.B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002).
Lindblad-Toh, K. et al. Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature 438, 803–819 (2005).
The Bovine HapMap Consortium. Genome-wide survey of SNP variation uncovers the genetic structure of cattle breeds. Science 324, 528–532 (2009).
Watterson, G.A. On the number of segregating sites in genetical models without recombination. Theor. Popul. Biol. 7, 256–276 (1975).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Liu, B. et al. QTL mapping of domestication-related traits in soybean (Glycine max). Ann. Bot. (Lond.) 100, 1027–1038 (2007).
Li, H. et al. Identification of QTL underlying vitamin E contents in soybean seed among multiple environments. Theor. Appl. Genet. 120, 1405–1413 (2010).
Huang, Z.-W., Zhao, T.-J., Yu, D.-Y., Chen, S.-Y. & Gai, J.-Y. Correlation and QTL mapping of biomass accumulation, apparent harvest index, and yield in soybean. Acta. Agron. Sin. 34, 944–951 (2008).
McNally, K.L. et al. Genomewide SNP variation reveals relationships among landraces and modern varieties of rice. Proc. Natl. Acad. Sci. USA 106, 12273–12278 (2009).
Clark, R.M. et al. Common sequence polymorphisms shaping genetic diversity in Arabidopsis thaliana. Science 317, 338–342 (2007).
Jordan, I.K., Rogozin, I.B., Wolf, Y.I. & Koonin, E.V. Essential genes are more evolutionarily conserved than are nonessential genes in bacteria. Genome Res. 12, 962–968 (2002).
Dangl, J.L. & Jones, J.D.G. Plant pathogens and integrated defence responses to infection. Nature 411, 826–833 (2001).
Blanc, G. & Wolfe, K.H. Functional divergence of duplicated genes formed by polyploidy during Arabidopsis evolution. Plant Cell 16, 1679–1691 (2004).
Maere, S. et al. Modeling gene and genome duplications in eukaryotes. Proc. Natl. Acad. Sci. USA 102, 5454–5459 (2005).
Lynch, M. & Conery, J.S. The evolutionary fate and consequences of duplicate genes. Science 290, 1151–1155 (2000).
Li, R. et al. Building the sequence map of the human pan-genome. Nat. Biotechnol. 28, 57–63 (2010).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004).
Stanke, M., Schöffmann, O., Morgenstern, B. & Waack, S. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC Bioinformatics 7, 62 (2006).
Lu, J. et al. The accumulation of deleterious mutations in rice genomes: a hypothesis on the cost of domestication. TIG 22, 126–131 (2006).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Doyle, J.J. & Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19, 11–15 (1987).
Schwartz, S. et al. Human-mouse alignments with BLASTZ. Genome Res. 13, 103–107 (2003).
Tamura, K., Dudley, J., Nei, M. & Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596–1599 (2007).
Hudson, R.R. Generating samples under a Wright-Fisher neutral model of genetic variation. Bioinformatics 18, 337–338 (2002).
Akey, J.M., Zhang, G., Zhang, K., Jin, L. & Shriver, M.D. Interrogating a high-density SNP map for signatures of natural selection. Genome Res. 12, 1805–1814 (2002).
McDonald, J.H. & Kreitman, M. Adaptive protein evolution at the Adh locus in Drosophila. Nature 351, 652–654 (1991).
Kent, W.J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Acknowledgements
T. Han, X. Yan, H. Liao, B. Zhuang and Y.-K. Lau provided valuable advice, information and other aid. This work was partially supported by the Hong Kong RGC General Research Fund 468610 (to H.-M.L.), the Hong Kong UGC AoE Center for Plant and Agricultural Biotechnology Project AoE-B-07/09 and a special fund from the Resource Allocation Committee, The Chinese University of Hong Kong (to H.-M.L. and S.S.-M.S.). We also acknowledge the funding support from the National Natural Science Foundation of China (30725008), the Chinese 973 program (2007CB815703; 2007CB815705), Chinese Ministry of Agriculture (948 program), the Shenzhen Municipal Government of China and grants from Shenzhen Bureau of Science Technology & Information, China (ZYC200903240077A; CXB200903110066A). We thank L. Goodman for assistance in editing the manuscript.
Author information
Authors and Affiliations
Contributions
H.-M.L., G.Z., S.S.-M.S. and Jun Wang managed the project. H.-M.L., X.X., X.L, N.Q. and G.Y. designed the experiments and led the data analysis. W.H., B.W., J.L., W.C., M.J. and Jian Wang contributed to DNA sequencing and bioinformatics. F.-L.W., M.-W.L. and G.S. prepared samples and contributed to data analysis. H.-M.L., X.X. and X.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 and Supplementary Tables 1–7 (PDF 845 kb)
Rights and permissions
About this article
Cite this article
Lam, HM., Xu, X., Liu, X. et al. Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat Genet 42, 1053–1059 (2010). https://doi.org/10.1038/ng.715
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.715
This article is cited by
-
Genomic variation and candidate genes dissect quality and yield traits in Boehmeria nivea (L.) Gaudich
Cellulose (2024)
-
Genome-wide association study on resistance of cultivated soybean to Fusarium oxysporum root rot in Northeast China
BMC Plant Biology (2023)
-
Association mapping in bambara groundnut [Vigna subterranea (L.) Verdc.] reveals loci associated with agro-morphological traits
BMC Genomics (2023)
-
Polymorphism analysis of the chloroplast and mitochondrial genomes in soybean
BMC Plant Biology (2023)
-
Resequencing of Rosa rugosa accessions revealed the history of population dynamics, breed origin, and domestication pathways
BMC Plant Biology (2023)