Abstract
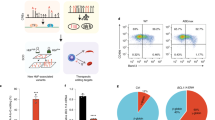
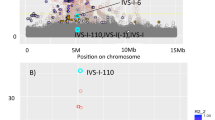
We used resequencing and genotyping in African Americans with sickle cell anemia (SCA) to characterize associations with fetal hemoglobin (HbF) levels at the BCL11A, HBS1L-MYB and β-globin loci. Fine-mapping of HbF association signals at these loci confirmed seven SNPs with independent effects and increased the explained heritable variation in HbF levels from 38.6% to 49.5%. We also identified rare missense variants that causally implicate MYB in HbF production.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Thein, S.L. et al. Proc. Natl. Acad. Sci. USA 104, 11346–11351 (2007).
Menzel, S. et al. Nat. Genet. 39, 1197–1199 (2007).
Uda, M. et al. Proc. Natl. Acad. Sci. USA 105, 1620–1625 (2008).
Lettre, G. et al. Proc. Natl. Acad. Sci. USA 105, 11869–11874 (2008).
Sankaran, V.G. et al. Science 322, 1839–1842 (2008).
Sankaran, V.G. et al. Nature 460, 1093–1097 (2009).
Manolio, T.A. et al. Nature 461, 747–753 (2009).
Nejentsev, S., Walker, N., Riches, D., Egholm, M. & Todd, J.A. Science 324, 387–389 (2009).
Dickson, S.P., Wang, K., Krantz, I., Hakonarson, H. & Goldstein, D.B. PLoS Biol. 8, e1000294 (2010).
Embury, S.H. et al. Sickle Cell Disease: Basic Principles and Clinical Practice (Lippincott Williams & Wilkins, Philadelphia, Pennsylvania, USA1994).
Labie, D. et al. Proc. Natl. Acad. Sci. USA 82, 2111–2114 (1985).
Solovieff, N. et al. Blood 115, 1815–1822 (2010).
Galanello, R. et al. Blood 114, 3935–3937 (2009).
Nuinoon, M. et al. Hum. Genet. 127, 303–314 (2009).
Pilia, G. et al. PLoS Genet. 2, e132 (2006).
Acknowledgements
We thank all the individuals who participated in this study, and T. Nguyen and M. Beaudoin for DNA genotyping support. We thank S. Raychaudhuri for critical reading of the manuscript, G. Boucher for statistical advice and the CARe Sickle Cell Disease working group for providing the Cooperative Study of Sickle Cell Disease (CSSCD) principal components. This work was funded by the Fondation de l'Institut de Cardiologie de Montréal (to G.L.) and was supported by an Innovations in Clinical Research Award grant from the Doris Duke Charitable Foundation (to G.L. and J.N.H.). Resequencing services were provided by the University of Washington, Department of Genome Sciences, under US Federal Government contract number N01-HV-48194 from the National Heart, Lung, and Blood Institute.
Author information
Authors and Affiliations
Contributions
V.G.S., S.H.O., J.N.H. and G.L. conceived and designed the experiment. G.G., C.D.P. and G.L. performed the experiments. G.G., C.D.P. and G.L. analyzed the data. G.G., C.D.P., V.G.S., S.H.O., J.N.H. and G.L. contributed reagents, materials and/or analysis tools. G.G. and G.L. wrote the paper with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–10 and Supplementary Figures 1 and 2 (PDF 579 kb)
Supplementary Table 2
DNA sequence variants identified at the BCL11A, HBS1L-MYB and A-globin loci by DNA re-sequencing in the HapMap Northern European (CEU) and West African (YRI) founders, and in 70 sickle cell anemia (SCA) patients from the Cooperative Study of Sickle Cell Disease (CSSCD) (XLS 255 kb)
Supplementary Table 3
Association results with HbF levels for all 95 common SNPs (minor allele frequency A1%) genotyped in 1,032 African-American sickle cell anemia (SCA) patients from the Cooperative Study of Sickle Cell Disease (CSSCD) (XLS 35 kb)
Supplementary Table 6
Association results to fetal hemoglobin (HbF) levels for imputed SNPs (XLS 103 kb)
Rights and permissions
About this article
Cite this article
Galarneau, G., Palmer, C., Sankaran, V. et al. Fine-mapping at three loci known to affect fetal hemoglobin levels explains additional genetic variation. Nat Genet 42, 1049–1051 (2010). https://doi.org/10.1038/ng.707
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.707
This article is cited by
-
Challenges and future directions for studying effects of host genetics on the gut microbiome
Nature Genetics (2022)
-
Targeting Genetic Modifiers of HBG Gene Expression in Sickle Cell Disease: The miRNA Option
Molecular Diagnosis & Therapy (2022)
-
Genome-based therapeutic interventions for β-type hemoglobinopathies
Human Genomics (2021)
-
High fetal hemoglobin level is associated with increased risk of cerebral vasculopathy in children with sickle cell disease in Mayotte
BMC Pediatrics (2020)
-
Beyond large-effect loci: large-scale GWAS reveals a mixed large-effect and polygenic architecture for age at maturity of Atlantic salmon
Genetics Selection Evolution (2020)