Abstract
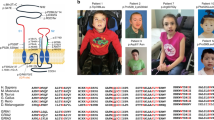
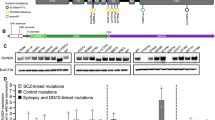
N-methyl-D-aspartate (NMDA) receptors mediate excitatory neurotransmission in the mammalian brain. Two glycine-binding NR1 subunits and two glutamate-binding NR2 subunits each form highly Ca2+-permeable cation channels which are blocked by extracellular Mg2+ in a voltage-dependent manner1. Either GRIN2B or GRIN2A, encoding the NMDA receptor subunits NR2B and NR2A, was found to be disrupted by chromosome translocation breakpoints in individuals with mental retardation and/or epilepsy. Sequencing of GRIN2B in 468 individuals with mental retardation revealed four de novo mutations: a frameshift, a missense and two splice-site mutations. In another cohort of 127 individuals with idiopathic epilepsy and/or mental retardation, we discovered a GRIN2A nonsense mutation in a three-generation family. In a girl with early-onset epileptic encephalopathy, we identified the de novo GRIN2A mutation c.1845C>A predicting the amino acid substitution p.N615K. Analysis of NR1-NR2AN615K (NR2A subunit with the p.N615K alteration) receptor currents revealed a loss of the Mg2+ block and a decrease in Ca2+ permeability. Our findings suggest that disturbances in the neuronal electrophysiological balance during development result in variable neurological phenotypes depending on which NR2 subunit of NMDA receptors is affected.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Accessions
GenBank/EMBL/DDBJ
Protein Data Bank
References
Cull-Candy, S., Brickley, S. & Farrant, M. NMDA receptor subunits: diversity, development and disease. Curr. Opin. Neurobiol. 11, 327–335 (2001).
Cull-Candy, S.G. & Leszkiewicz, D.N. Role of distinct NMDA receptor subtypes at central synapses. Sci. STKE 2004, re16 (2004).
Lau, C.G. & Zukin, R.S. NMDA receptor trafficking in synaptic plasticity and neuropsychiatric disorders. Nat. Rev. Neurosci. 8, 413–426 (2007).
Kurotaki, N. et al. Haploinsufficiency of NSD1 causes Sotos syndrome. Nat. Genet. 30, 365–366 (2002).
Najm, J. et al. Mutations of CASK cause an X-linked brain malformation phenotype with microcephaly and hypoplasia of the brainstem and cerebellum. Nat. Genet. 40, 1065–1067 (2008).
Ray, P.N. et al. Cloning of the breakpoint of an X;21 translocation associated with Duchenne muscular dystrophy. Nature 318, 672–675 (1985).
Tonkin, E.T. et al. NIPBL, encoding a homolog of fungal Scc2-type sister chromatid cohesion proteins and fly Nipped-B, is mutated in Cornelia de Lange syndrome. Nat. Genet. 36, 636–641 (2004).
Akashi, K. et al. NMDA receptor GluN2B (GluR epsilon 2/NR2B) subunit is crucial for channel function, postsynaptic macromolecular organization, and actin cytoskeleton at hippocampal CA3 synapses. J. Neurosci. 29, 10869–10882 (2009).
Tang, Y.P. et al. Genetic enhancement of learning and memory in mice. Nature 401, 63–69 (1999).
Wang, D. et al. Genetic enhancement of memory and long-term potentiation but not CA1 long-term depression in NR2B transgenic rats. PLoS ONE 4, e7486 (2009).
Reutlinger, C. et al. Deletions in 16p13 including GRIN2A in patients with intellectual disability, various dysmorphic features and seizure disorder of the rolandic region. Epilepsia published online, doi:10.1111/j.1528-1167.2010.02555.x (2 April 2010).
Sakimura, K. et al. Reduced hippocampal LTP and spatial learning in mice lacking NMDA receptor epsilon 1 subunit. Nature 373, 151–155 (1995).
Kiyama, Y. et al. Increased thresholds for long-term potentiation and contextual learning in mice lacking the NMDA-type glutamate receptor epsilon1 subunit. J. Neurosci. 18, 6704–6712 (1998).
Kutsuwada, T. et al. Impairment of suckling response, trigeminal neuronal pattern formation, and hippocampal LTD in NMDA receptor epsilon 2 subunit mutant mice. Neuron 16, 333–344 (1996).
Wang, G.S. & Cooper, T.A. Splicing in disease: disruption of the splicing code and the decoding machinery. Nat. Rev. Genet. 8, 749–761 (2007).
Laube, B. et al. Molecular determinants of agonist discrimination by NMDA receptor subunits: analysis of the glutamate binding site on the NR2B subunit. Neuron 18, 493–503 (1997).
Laube, B., Schemm, R. & Betz, H. Molecular determinants of ligand discrimination in the glutamate-binding pocket of the NMDA receptor. Neuropharmacology 47, 994–1007 (2004).
Mayer, M.L. Glutamate receptors at atomic resolution. Nature 440, 456–462 (2006).
Wollmuth, L.P., Kuner, T. & Sakmann, B. Adjacent asparagines in the NR2-subunit of the NMDA receptor channel control the voltage-dependent block by extracellular Mg2+. J. Physiol. (Lond.) 506, 13–32 (1998).
Yashiro, K. & Philpot, B.D. Regulation of NMDA receptor subunit expression and its implications for LTD, LTP, and metaplasticity. Neuropharmacology 55, 1081–1094 (2008).
Ding, Y.X. et al. A possible association of responsiveness to adrenocorticotropic hormone with specific GRIN1 haplotypes in infantile spasms. Dev. Med. Child Neurol. published online, doi:10.1111/j.1469-8749.2010.03746.x (16 August 2010).
Barrett, M.T. et al. Comparative genomic hybridization using oligonucleotide microarrays and total genomic DNA. Proc. Natl. Acad. Sci. USA 101, 17765–17770 (2004).
Spitz, R. et al. Oligonucleotide array-based comparative genomic hybridization (aCGH) of 90 neuroblastomas reveals aberration patterns closely associated with relapse pattern and outcome. Genes Chromosom. Cancer 45, 1130–1142 (2006).
Chen, W. et al. CGHPRO–a comprehensive data analysis tool for array CGH. BMC Bioinformatics 6, 85 (2005).
Desmet, F.O. et al. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 37, e67 (2009).
Reese, M.G., Eeckman, F.H., Kulp, D. & Haussler, D. Improved splice site detection in Genie. J. Comput. Biol. 4, 311–323 (1997).
Hebsgaard, S.M. et al. Splice site prediction in Arabidopsis thaliana pre-mRNA by combining local and global sequence information. Nucleic Acids Res. 24, 3439–3452 (1996).
Kumar, P., Henikoff, S. & Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Ramensky, V., Bork, P. & Sunyaev, S. Human non-synonymous SNPs: server and survey. Nucleic Acids Res. 30, 3894–3900 (2002).
Bromberg, Y. & Rost, B. SNAP: predict effect of non-synonymous polymorphisms on function. Nucleic Acids Res. 35, 3823–3835 (2007).
Thomas, P.D. et al. PANTHER: a library of protein families and subfamilies indexed by function. Genome Res. 13, 2129–2141 (2003).
Sobolevsky, A.I., Rosconi, M.P. & Gouaux, E. X-ray structure, symmetry and mechanism of an AMPA-subtype glutamate receptor. Nature 462, 745–756 (2009).
Furukawa, H., Singh, S.K., Mancusso, R. & Gouaux, E. Subunit arrangement and function in NMDA receptors. Nature 438, 185–192 (2005).
Fiser, A. & Sali, A. Modeller: generation and refinement of homology-based protein structure models. Methods Enzymol. 374, 461–491 (2003).
Kleywegt, G.J. Use of non-crystallographic symmetry in protein structure refinement. Acta Crystallogr. D Biol. Crystallogr. 52, 842–857 (1996).
Schüler, T. et al. Formation of NR1/NR2 and NR1/NR3 heterodimers constitutes the initial step in N-methyl-D-aspartate receptor assembly. J. Biol. Chem. 283, 37–46 (2008).
Haeger, S. et al. An intramembrane aromatic network determines pentameric assembly of Cys-loop receptors. Nat. Struct. Mol. Biol. 17, 90–98 (2010).
Madry, C., Betz, H., Geiger, J.R. & Laube, B. Potentiation of glycine-gated NR1/NR3A NMDA receptors relieves Ca-dependent outward rectification. Front. Mol. Neurosci. 3, 6 (2010).
Acknowledgements
We thank all subjects and healthy (control) individuals for their participation in this study; P. De Jonghe for providing DNA samples of individuals with idiopathic epilepsy; A. Gal and B. Horsthemke for continuous support; H.-H. Richardt and H. Petri for clinical evaluation of subjects 3 and 1; A. Tzschach and M. Hoeltzenbein for initial help with collecting clinical data of subject 2; S. Fuchs for chromosome analysis; B. Lübker and C. Menzel for FISH analysis; R. Ullmann and A. Ahmed for array comparative genomic hybridization (CGH) on subject 2, O. Riess and M. Bonin for copy number variation analysis on subjects 8 and 9, and I. Göhring and A.B. Ekici for microarray analysis of the Erlanger cohort; S. Berkel, J. Hoyer and K. Cremer for support with data maintenance; and A. Diem, S. Freier, I. Jantke, S. Meien and K. Ziegler for skillfull technical assistance. This work was part of the German Mental Retardation Network (MRNET) funded through a grant from the German Ministry of Research and Education to A. Rauch and A. Reis (01GS08160), G. Rappold (01GS08168-9), H.-H.R. (01GS08161) and D.W. (01GS08164). M.M. was supported by the Fondation Française pour la Recherche sur l'Epilepsie, B.L. was supported by the Gemeinnützige Hertie-Stiftung, and G. Rosenberger and K.K. were supported by a grant from the Deutsche Forschungsgemeinschaft (FOR 885/IRP5).
Author information
Authors and Affiliations
Contributions
Mutation analysis: S.E., B.P. and G. Rosenberger. Transcript analysis: K.K. and G. Rosenberger. Functional analysis of NMDA receptors: K.G. and B.L. NMDA receptor modeling: C.T. and B.L. Subject ascertainment: Y.H., L.V.M., M.M., U.M., G. Rappold, A. Rauch, S.v.S., I.S., N.V., L.V., D.W., B.Z. and M.Z. FISH analysis and breakpoint mapping: A.F., V.M.K., F.K., F.K.P. and H.-H.R. Array CGH analysis: J.K. and H.T. Manuscript writing: K.K., V.M.K., B.L., L.V.M., A. Reis, G. Rosenberger and D.W. Study design: K.K., L.V.M., A. Reis, G. Rosenberger and D.W. All authors contributed to the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–8 and Supplementary Note (PDF 520 kb)
Rights and permissions
About this article
Cite this article
Endele, S., Rosenberger, G., Geider, K. et al. Mutations in GRIN2A and GRIN2B encoding regulatory subunits of NMDA receptors cause variable neurodevelopmental phenotypes. Nat Genet 42, 1021–1026 (2010). https://doi.org/10.1038/ng.677
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.677
This article is cited by
-
Genetik und genetische Diagnostik fokaler Epilepsien des Kindesalters – Was? Wann? Warum?
Clinical Epileptology (2024)
-
Association of polygenic score and the involvement of cholinergic and glutamatergic pathways with lithium treatment response in patients with bipolar disorder
Molecular Psychiatry (2023)
-
Gene variations of glutamate metabolism pathway and epilepsy
Acta Epileptologica (2022)
-
Common synaptic phenotypes arising from diverse mutations in the human NMDA receptor subunit GluN2A
Communications Biology (2022)
-
Complex functional phenotypes of NMDA receptor disease variants
Molecular Psychiatry (2022)