Abstract
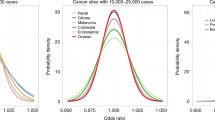
We report a set of tools to estimate the number of susceptibility loci and the distribution of their effect sizes for a trait on the basis of discoveries from existing genome-wide association studies (GWASs). We propose statistical power calculations for future GWASs using estimated distributions of effect sizes. Using reported GWAS findings for height, Crohn's disease and breast, prostate and colorectal (BPC) cancers, we determine that each of these traits is likely to harbor additional loci within the spectrum of low-penetrance common variants. These loci, which can be identified from sufficiently powerful GWASs, together could explain at least 15–20% of the known heritability of these traits. However, for BPC cancers, which have modest familial aggregation, our analysis suggests that risk models based on common variants alone will have modest discriminatory power (63.5% area under curve), even with new discoveries.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Manolio, T.A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Hirschhorn, J.N. Genomewide association studies–illuminating biologic pathways. N. Engl. J. Med. 360, 1699–1701 (2009).
Goldstein, D.B. Common genetic variation and human traits. N. Engl. J. Med. 360, 1696–1698 (2009).
Kraft, P. et al. Beyond odds ratios–communicating disease risk based on genetic profiles. Nat. Rev. Genet. 10, 264–269 (2009).
Pharoah, P.D. et al. Polygenic susceptibility to breast cancer and implications for prevention. Nat. Genet. 31, 33–36 (2002).
Gail, M.H. Value of adding single-nucleotide polymorphism genotypes to a breast cancer risk model. J. Natl. Cancer Inst. 101, 959–963 (2009).
Gail, M.H. Discriminatory accuracy from single-nucleotide polymorphisms in models to predict breast cancer risk. J. Natl. Cancer Inst. 100, 1037–1041 (2008).
Xu, J. et al. Estimation of absolute risk for prostate cancer using genetic markers and family history. Prostate 69, 1565–1572 (2009).
Meigs, J.B. et al. Genotype score in addition to common risk factors for prediction of type 2 diabetes. N. Engl. J. Med. 359, 2208–2219 (2008).
Wacholder, S. et al. Performance of common genetic variants in breast-cancer risk models. N. Engl. J. Med. 362, 986–993 (2010).
Kraft, P. & Hunter, D.J. Genetic risk prediction–are we there yet? N. Engl. J. Med. 360, 1701–1703 (2009).
Visscher, P.M. Sizing up human height variation. Nat. Genet. 40, 489–490 (2008).
Gudbjartsson, D.F. et al. Many sequence variants affecting diversity of adult human height. Nat. Genet. 40, 609–615 (2008).
Lettre, G. et al. Identification of ten loci associated with height highlights new biological pathways in human growth. Nat. Genet. 40, 584–591 (2008).
Weedon, M.N. et al. Genome-wide association analysis identifies 20 loci that influence adult height. Nat. Genet. 40, 575–583 (2008).
Weedon, M.N. & Frayling, T.M. Reaching new heights: insights into the genetics of human stature. Trends Genet. 24, 595–603 (2008).
Barrett, J.C. et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn's disease. Nat. Genet. 40, 955–962 (2008).
Lichtenstein, P. et al. Environmental and heritable factors in the causation of cancer–analyses of cohorts of twins from Sweden, Denmark, and Finland. N. Engl. J. Med. 343, 78–85 (2000).
Easton, D.F. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007).
Eeles, R.A. et al. Multiple newly identified loci associated with prostate cancer susceptibility. Nat. Genet. 40, 316–321 (2008).
Houlston, R.S. et al. Meta-analysis of genome-wide association data identifies four new susceptibility loci for colorectal cancer. Nat. Genet. 40, 1426–1435 (2008).
Thomas, G. et al. A multistage genome-wide association study in breast cancer identifies two new risk alleles at 1p11.2 and 14q24.1 (RAD51L1). Nat. Genet. 41, 579–584 (2009).
Thomas, G. et al. Multiple loci identified in a genome-wide association study of prostate cancer. Nat. Genet. 40, 310–315 (2008).
Eeles, R.A. et al. Identification of seven new prostate cancer susceptibility loci through a genome-wide association study. Nat. Genet. 41, 1116–1121 (2009).
Orr, H.A. The population genetics of adaptation: The distribution of factors fixed during adaptive evolution. Evolution 52, 935–949 (1998).
Eberle, M.A. et al. Power to detect risk alleles using genome-wide tag SNP panels. PLoS Genet. 3, 1827–1837 (2007).
Schork, N.J. Power calculations for genetic association studies using estimated probability distributions. Am. J. Hum. Genet. 70, 1480–1489 (2002).
Ambrosius, W.T., Lange, E.M. & Langefeld, C.D. Power for genetic association studies with random allele frequencies and genotype distributions. Am. J. Hum. Genet. 74, 683–693 (2004).
Spencer, C.C., Su, Z., Donnelly, P. & Marchini, J. Designing genome-wide association studies: sample size, power, imputation, and the choice of genotyping chip. PLoS Genet. 5, e1000477 (2009).
Dickson, S.P., Wang, K., Krantz, I., Hakonarson, H. & Goldstein, D.B. Rare variants create synthetic genome-wide associations. PLoS Biol. 8, e1000294 (2010).
Yu, K. et al. Flexible design for following up positive findings. Am. J. Hum. Genet. 81, 540–551 (2007).
Ghosh, A., Zou, F. & Wright, F.A. Estimating odds ratios in genome scans: an approximate conditional likelihood approach. Am. J. Hum. Genet. 82, 1064–1074 (2008).
Li, B. & Leal, S.M. Discovery of rare variants via sequencing: implications for the design of complex trait association studies. PLoS Genet. 5, e1000481 (2009).
Li, B. & Leal, S.M. Methods for detecting associations with rare variants for common diseases: application to analysis of sequence data. Am. J. Hum. Genet. 83, 311–321 (2008).
Zhong, H. & Prentice, R.L. Bias-reduced estimators and confidence intervals for odds ratios in genome-wide association studies. Biostatistics 9, 621–634 (2008).
Zhong, H. & Prentice, R.L. Correcting “winner's curse” in odds ratios from genomewide association findings for major complex human diseases. Genet. Epidemiol. 34, 78–91 (2009).
Acknowledgements
This work was supported by the intramural program of the National Cancer Institute, US National Institutes of Health. The research of N.C. and J.-H.P. was also partially funded by the Gene-Environment Initiative of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
J.-H.P. and N.C. developed the statistical methods and designed the analyses. J.-H.P. implemented the methods and carried out all analyses. N.C. and S.J.C. drafted the manuscript. S.W., M.H.G., K.B.J. and U.P. made important suggestions for presentation and interpretation of the results. All the authors participated in critically reviewing the paper and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–7 and Supplementary Note. (PDF 544 kb)
Rights and permissions
About this article
Cite this article
Park, JH., Wacholder, S., Gail, M. et al. Estimation of effect size distribution from genome-wide association studies and implications for future discoveries. Nat Genet 42, 570–575 (2010). https://doi.org/10.1038/ng.610
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.610
This article is cited by
-
The APOE locus is linked to decline in general cognitive function: 20-years follow-up in the Doetinchem Cohort Study
Translational Psychiatry (2022)
-
Different responses to risperidone treatment in Schizophrenia: a multicenter genome-wide association and whole exome sequencing joint study
Translational Psychiatry (2022)
-
Reconstructing SNP allele and genotype frequencies from GWAS summary statistics
Scientific Reports (2022)
-
A genome-wide association analysis for body weight at 35 days measured on 137,343 broiler chickens
Genetics Selection Evolution (2021)
-
Competition for priority harms the reliability of science, but reforms can help
Nature Human Behaviour (2021)