Abstract
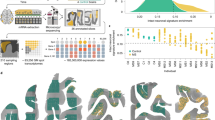
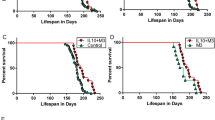
Amyotrophic lateral sclerosis (ALS) is a fatal neurodegenerative disease characterized by progressive loss of motor neurons. Using unbiased transcript profiling in an ALS mouse model, we identified a role for the co-stimulatory pathway, a key regulator of immune responses. Furthermore, we observed that this pathway is upregulated in the blood of 56% of human patients with ALS. A therapy using a monoclonal antibody to CD40L was developed that slows weight loss, delays paralysis and extends survival in an ALS mouse model. This work demonstrates that unbiased transcript profiling can identify cellular pathways responsive to therapeutic intervention in a preclinical model of human disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Kola, I. & Landis, J. Opinion: can the pharmaceutical industry reduce attrition rates? Nat. Rev. Drug Discov. 8, 711–716 (2004).
Hess, K.R. et al. Pharmacogenomic predictor of sensitivity to preoperative chemotherapy with paclitaxel and fluorouracil, doxorubicin, and cyclophosphamide in breast cancer. J. Clin. Oncol. 24, 4236–4244 (2006).
Bonnefoi, H. et al. Validation of gene signatures that predict the response of breast cancer to neoadjuvant chemotherapy: a substudy of the EORTC 10994/BIG 00–01 clinical trial. Lancet Oncol. 8, 1071–1078 (2007).
Cooper, C.S., Campbell, C. & Jhavar, S. Mechanisms of disease: biomarkers and molecular targets from microarray gene expression studies in prostate cancer. Nat. Clin. Pract. Urol. 4, 677–687 (2007).
Lacroix, L., Commo, F. & Soria, J.C. Gene expression profiling of non-small-cell lung cancer. Expert Rev. Mol. Diagn. 8, 167–178 (2008).
Kinter, J., Zeis, T. & Schaeren-Wiemers, N. RNA profiling of MS brain tissues. Int. MS J. 15, 51–58 (2008).
Bourquin, J.P. et al. Identification of distinct molecular phenotypes in acute megakaryoblastic leukemia by gene expression profiling. Proc. Natl. Acad. Sci. USA 103, 3339–3344 (2006).
Golub, T.R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999).
Decristofaro, M.F. & Daniels, K.K. Toxicogenomics in biomarker discovery. Methods Mol. Biol. 460, 185–194 (2008).
Merrick, B.A. & Bruno, M.E. Genomic and proteomic profiling for biomarkers and signature profiles of toxicity. Curr. Opin. Mol. Ther. 6, 600–607 (2004).
Ideker, T., Ozier, O., Schwikowski, B. & Siegel, A.F. Discovering regulatory and signalling circuits in molecular interaction networks. Bioinformatics 18 (Suppl. 1), 233–240 (2002).
Linsley, P.S., Clark, E.A. & Ledbetter, J.A. T-cell antigen CD28 mediates adhesion with B cells by interacting with activation antigen B7/BB-1. Proc. Natl. Acad. Sci. USA 87, 5031–5035 (1990).
Freeman, G.J. et al. Cloning of B7–2: a CTLA-4 counter-receptor that costimulates human T cell proliferation. Science 262, 909–911 (1993).
Noelle, R.J., Ledbetter, J.A. & Aruffo, A. CD40 and its ligand, an essential ligand-receptor pair for thymus-dependent B-cell activation. Immunol. Today 13, 431–433 (1992).
Roy, M., Waldschmidt, T., Aruffo, A., Ledbetter, J.A. & Noelle, R.J. The regulation of the expression of gp39, the CD40 ligand, on normal and cloned CD4+ T cells. J. Immunol. 151, 2497–2510 (1993).
Durie, F.H. et al. Prevention of collagen-induced arthritis with an antibody to gp39, the ligand for CD40. Science 261, 1328–1330 (1993).
Gerritse, K. et al. CD40–CD40 ligand interactions in experimental allergic encephalomyelitis and multiple sclerosis. Proc. Natl. Acad. Sci. USA 93, 2499–2504 (1996).
Mohan, C., Shi, Y., Laman, J.D. & Datta, S.K. Interaction between CD40 and its ligand gp39 in the development of murine lupus nephritis. J. Immunol. 154, 1470–1480 (1995).
Ma, J. et al. Autoimmune lpr/lpr mice deficient in CD40 ligand: spontaneous Ig class switching with dichotomy of autoantibody responses. J. Immunol. 157, 417–426 (1996).
Grewal, I.S. et al. Requirement for CD40 ligand in costimulation induction, T cell activation, and experimental allergic encephalomyelitis. Science 273, 1864–1867 (1996).
Kirk, A.D. et al. CTLA4-Ig and anti-CD40 ligand prevent renal allograft rejection in primates. Proc. Natl. Acad. Sci. USA 94, 8789–8794 (1997).
Larsen, C.P. et al. CD40-gp39 interactions play a critical role during allograft rejection. Suppression of allograft rejection by blockade of the CD40-gp39 pathway. Transplantation 61, 4–9 (1996).
Monk, N.J. et al. Fc-dependent depletion of activated T cells occurs through CD40L-specific antibody rather than costimulation blockade. Nat. Med. 9, 1275–1280 (2003).
Durie, F.H. et al. Prevention of collagen-induced arthritis with an antibody to gp39, the ligand for CD40. Science 261, 1328–1330 (1993).
Gallon, L. et al. Differential effects of B7–1 blockade in the rat experimental autoimmune encephalomyelitis model. J. Immunol. 159, 4212–4216 (1997).
Tan, J. et al. Role of CD40 ligand in amyloidosis in transgenic Alzheimer's mice. Nat. Neurosci. 5, 1288–1293 (2002).
Goeman, J.J., Van de Geer, S.A., De Kort, F. & Van Houwelingen, J.C. A global test for groups of genes: testing association with a clinical outcome. Bioinformatics 20, 93–99 (2004).
Beers, D.R., Henkel, J.S., Zhao, W., Wang, J. & Appel, S.H. CD4+ T cells support glial neuroprotection, slow disease progression, and modify glial morphology in an animal model of inherited ALS. Proc. Natl. Acad. Sci. USA 105, 15558–15563 (2008).
Beers, D.R. et al. Wild-type microglia extend survival in PU.1 knockout mice with familial amyotrophic lateral sclerosis. Proc. Natl. Acad. Sci. USA 103, 16021–16026 (2006).
Engelhardt, J.I., Tajti, J. & Appel, S.H. Lymphocytic infiltrates in the spinal cord in amyotrophic lateral sclerosis. Arch. Neurol. 50, 30–36 (1993).
Kawamata, T., Akiyama, H., Yamada, T. & McGeer, P.L. Immunologic reactions in amyotrophic lateral sclerosis brain and spinal cord tissue. Am. J. Pathol. 140, 691–707 (1992).
Troost, D., Van den Oord, J.J. & Vianney de Jong, J.M. Immunohistochemical characterization of the inflammatory infiltrate in amyotrophic lateral sclerosis. Neuropathol. Appl. Neurobiol. 16, 401–410 (1990).
Troost, D., van den Oord, J.J., de Jong, J.M. & Swaab, D.F. Lymphocytic infiltration in the spinal cord of patients with amyotrophic lateral sclerosis. Clin. Neuropathol. 8, 289–294 (1989).
Banerjee, R. et al. Adaptive immune neuroprotection in G93A–SOD1 amyotrophic lateral sclerosis mice. PLoS One 3, 2740 (2008).
Hughes, R., Atkinson, P., Coates, P., Hall, S. & Leibowitz, S. Sural nerve biopsies in Guillain-Barre syndrome: axonal degeneration and macrophage-associated demyelination and absence of cytomegalovirus genome. Muscle Nerve 15, 568–575 (1992).
Kiefer, R., Kieseier, B.C., Brück, W., Hartung, H.P. & Toyka, K.V. Macrophage differentiation antigens in acute and chronic autoimmune polyneuropathies. Brain 121, 469–479 (1998).
Honey, K., Cobbold, S.P. & Waldmann, H. CD40 ligand blockade induces CD4+ T cell tolerance and linked suppression. J. Immunol. 163, 4805–4810 (1999).
Nagelkerken, L. et al. FcR interactions do not play a major role in inhibition of experimental autoimmune encephalomyelitis by anti-CD154 monoclonal antibodies. J. Immunol. 173, 993–999 (2004).
Ruderman, E & Pope, R. Co-stimulatory pathways in the therapy of rheumatoid arthritis. in New Therapeutic Targets in Rheumatoid Arthritis (ed. Tak, P.-P.) 27–43 (Birkhäuser, Basel, 2009).
Noelle, R.J. et al. A 39-kDa protein on activated helper T cells binds CD40 and transduces the signal for cognate activation of B cells. Proc. Natl. Acad. Sci. USA 89, 6550–6554 (1992).
Graca, L., Honey, K., Adams, E., Cobbold, S.P. & Waldmann, H. Cutting edge: anti-CD154 therapeutic antibodies induce infectious transplantation tolerance. J. Immunol. 165, 4783–4786 (2000).
Kalled, S.L., Cutler, A.H. & Ferrant, J.L. Long-term anti-CD154 dosing in nephritic mice is required to maintain survival and inhibit mediators of renal fibrosis. Lupus 10, 9–22 (2001).
Gill, A., Kidd, J., Vieira, F., Thompson, K. & Perrin, S. No benefit from chronic lithium dosing in a sibling-matched, gender balanced, investigator-blinded trial using a standard mouse model of familial ALS. PLoS One 4, e6489 (2009).
Scott, S. et al. Design, power, and interpretation of studies in the standard murine model of ALS. Amyotroph. Lateral Scler. 9, 4–15 (2008).
Harraz, M.M. et al. SOD1 mutations disrupt redox-sensitive Rac regulation of NADPH oxidase in a familial ALS model. J. Clin. Invest. 118, 659–670 (2008).
Acknowledgements
We thank S. Appel, J. McCoy, S. Hesterlee and R. Goldstein for their thoughtful scientific discussions and contributions, R. Puchalski and the staff of the Allen Institute for Brain Science for the contribution of the in situ hybridization data, S.F. Scott, who lost his battle with ALS in 2009, for establishing the rigorous parameters required to test therapeutics in the SOD1G93A model, J. Heywood and his family for establishing ALS TDI and continuing to support our efforts and A. Nieto and L. Nieto for their dedication and financial support for the development of therapeutics for ALS. This work was supported by the Muscular Dystrophy Association/Augie's Quest, the US Department of Defense, the RGK Foundation and all our patients with ALS and their families.
Author information
Authors and Affiliations
Contributions
J.M.L., F.G.V., A.G. and S.P. designed the experiments. M.Z.W. performed the immunohistochemistry and FACS experiments. R.S. and I.J.C. performed the motor neuron histology. B.A.L. oversaw and consulted on the human blood sample study. K.T., J.K. and A.M. performed all the animal studies. G.S.D.Z. consulted on the interpretation of the results. B.M.A. and S.M.S. wrote the simulation and LIMS software. A.G. performed all pharmacological statistical analysis. S.P., J.M.L., A.G. and F.G.V. wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Table 4 (PDF 3784 kb)
Supplementary Table 1
Calculated Q scores of significantly different pathways in SOD1G93A mice compared to non-transgenic littermates. (XLS 27 kb)
Supplementary Table 2
RMA normalized gene expression data of all genes in five significantly changing pathways in SOD1G93A mice compared to non-transgenic littermates. (XLS 735 kb)
Supplementary Table 3
Calculated fold-change data of all genes in five significantly changing pathways in SOD1G93A mice compared to non-transgenic littermates derived from RMA normalized data. (XLS 107 kb)
Supplementary Table 5
Relative normalized transcript expression of costimulatory genes in human clinical blood samples. (XLS 142 kb)
Supplementary Note
Clinical annotation of human clinical blood samples (XLS 69 kb)
Rights and permissions
About this article
Cite this article
Lincecum, J., Vieira, F., Wang, M. et al. From transcriptome analysis to therapeutic anti-CD40L treatment in the SOD1 model of amyotrophic lateral sclerosis. Nat Genet 42, 392–399 (2010). https://doi.org/10.1038/ng.557
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.557