Abstract
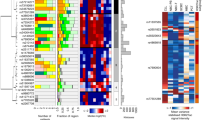
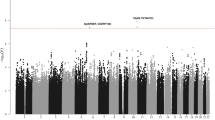
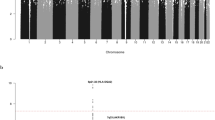
To identify new risk variants for chronic lymphocytic leukemia (CLL), we conducted a genome-wide association study of 299,983 tagging SNPs, with validation in four additional series totaling 2,503 cases and 5,789 controls. We identified four new risk loci for CLL at 2q37.3 (rs757978, FARP2; odds ratio (OR) = 1.39; P = 2.11 × 10−9), 8q24.21 (rs2456449; OR = 1.26; P = 7.84 × 10−10), 15q21.3 (rs7169431; OR = 1.36; P = 4.74 × 10−7) and 16q24.1 (rs305061; OR = 1.22; P = 3.60 × 10−7). We also found evidence for risk loci at 15q25.2 (rs783540, CPEB1; OR = 1.18; P = 3.67 × 10−6) and 18q21.1 (rs1036935; OR = 1.22; P = 2.28 × 10−6). These data provide further evidence for genetic susceptibility to this B-cell hematological malignancy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Goldin, L.R., Pfeiffer, R.M., Li, X. & Hemminki, K. Familial risk of lymphoproliferative tumors in families of patients with chronic lymphocytic leukemia: results from the Swedish Family-Cancer Database. Blood 104, 1850–1854 (2004).
Di Bernardo, M.C. et al. A genome-wide association study identifies six susceptibility loci for chronic lymphocytic leukemia. Nat. Genet. 40, 1204–1210 (2008).
Gunnarsson, R. et al. Screening for copy-number alterations and loss of heterozygosity in chronic lymphocytic leukemia—a comparative study of four differently designed, high resolution microarray platforms. Genes Chromosom. Cancer 47, 697–711 (2008).
Ferreira, B.I. et al. Comparative genome profiling across subtypes of low-grade B-cell lymphoma identifies type-specific and common aberrations that target genes with a role in B-cell neoplasia. Haematologica 93, 670–679 (2008).
Grubor, V. et al. Novel genomic alterations and clonal evolution in chronic lymphocytic leukemia revealed by representational oligonucleotide microarray analysis (ROMA). Blood 113, 1294–1303 (2009).
Pfeifer, D. et al. Genome-wide analysis of DNA copy number changes and LOH in CLL using high-density SNP arrays. Blood 109, 1202–1210 (2007).
Gudmundsson, J. et al. Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat. Genet. 39, 631–637 (2007).
Easton, D.F. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007).
Tomlinson, I. et al. A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nat. Genet. 39, 984–988 (2007).
Yeager, M. et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat. Genet. 39, 645–649 (2007).
Amundadottir, L.T. et al. A common variant associated with prostate cancer in European and African populations. Nat. Genet. 38, 652–658 (2006).
Kiemeney, L.A. et al. Sequence variant on 8q24 confers susceptibility to urinary bladder cancer. Nat. Genet. 40, 1307–1312 (2008).
Pomerantz, M.M. et al. The 8q24 cancer risk variant rs6983267 shows long-range interaction with MYC in colorectal cancer. Nat. Genet. 41, 882–884 (2009).
Tuupanen, S. et al. The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling. Nat. Genet. 41, 885–890 (2009).
Shaffer, A.L. et al. IRF4 addiction in multiple myeloma. Nature 454, 226–231 (2008).
Miyamoto, Y., Yamauchi, J. & Itoh, H. Src kinase regulates the activation of a novel FGD-1-related Cdc42 guanine nucleotide exchange factor in the signaling pathway from the endothelin A receptor to JNK. J. Biol. Chem. 278, 29890–29900 (2003).
Wang, H. & Morse, H.C. III. IRF8 regulates myeloid and B lymphoid lineage diversification. Immunol. Res. 43, 109–117 (2009).
De Jager, P.L. et al. Meta-analysis of genome scans and replication identify CD6, IRF8 and TNFRSF1A as new multiple sclerosis susceptibility loci. Nat. Genet. 41, 776–782 (2009).
Soderberg, K.C., Jonsson, F., Winqvist, O., Hagmar, L. & Feychting, M. Autoimmune diseases, asthma and risk of haematological malignancies: a nationwide case-control study in Sweden. Eur. J. Cancer 42, 3028–3033 (2006).
Cartwright, R.A. et al. Chronic lymphocytic leukaemia: case control epidemiological study in Yorkshire. Br. J. Cancer 56, 79–82 (1987).
Ikeda, M. & Longnecker, R. The c-Cbl proto-oncoprotein downregulates EBV LMP2A signaling. Virology 385, 183–191 (2009).
Zhang, X.Y., Supakar, P.C., Wu, K.Z., Ehrlich, K.C. & Ehrlich, M. An MDBP site in the first intron of the human c-myc gene. Cancer Res. 50, 6865–6869 (1990).
Hamblin, T.J., Davis, Z., Gardiner, A., Oscier, D.G. & Stevenson, F.K. Unmutated Ig V(H) genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood 94, 1848–1854 (1999).
Damle, R.N. et al. Ig V gene mutation status and CD38 expression as novel prognostic indicators in chronic lymphocytic leukemia. Blood 94, 1840–1847 (1999).
Zenz, T., Dohner, H. & Stilgenbauer, S. Genetics and risk-stratified approach to therapy in chronic lymphocytic leukemia. Best Pract. Res. Clin. Haematol. 20, 439–453 (2007).
Goldin, L.R. et al. A genome scan of 18 families with chronic lymphocytic leukaemia. Br. J. Haematol. 121, 866–873 (2003).
Sellick, G.S. et al. A high-density SNP genome-wide linkage search of 206 families identifies susceptibility loci for chronic lymphocytic leukemia. Blood 110, 3326–3333 (2007).
Sellick, G.S. et al. A high-density SNP genomewide linkage scan for chronic lymphocytic leukemia-susceptibility loci. Am. J. Hum. Genet. 77, 420–429 (2005).
Weiss, N.S. Geographical variation in the incidence of the leukemias and lymphomas. Natl. Cancer Inst. Monogr. 53, 139–142 (1979).
Allan, J.M. et al. Polymorphism in glutathione S-transferase P1 is associated with susceptibility to chemotherapy-induced leukemia. Proc. Natl. Acad. Sci. USA 98, 11592–11597 (2001).
Smedby, K.E. et al. Ultraviolet radiation exposure and risk of malignant lymphomas. J. Natl. Cancer Inst. 97, 199–209 (2005).
Catovsky, D. et al. Assessment of fludarabine plus cyclophosphamide for patients with chronic lymphocytic leukaemia (the LRF CLL4 Trial): a randomised controlled trial. Lancet 370, 230–239 (2007).
Jaffe, E.S., Harris, N.L., Stein, H. & Vardiman, J.W. World Health Organization Classification of Tumours: Pathology and Genetics of Tumours of Haematopoietic and Lymphoid Tissues (IARC, Lyon, France, 2001).
van Dongen, J.J. et al. Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4–CT98–3936. Leukemia 17, 2257–2317 (2003).
van Krieken, J.H. et al. Improved reliability of lymphoma diagnostics via PCR-based clonality testing: report of the BIOMED-2 Concerted Action BHM4–CT98–3936. Leukemia 21, 201–206 (2007).
Brochet, X., Lefranc, M.P. & Giudicelli, V. IMGT/V-QUEST: the highly customized and integrated system for IG and TR standardized V-J and V-D-J sequence analysis. Nucleic Acids Res. 36, W503–W508 (2008).
Petitti, D.B. Meta-analysis Decision Analysis and Cost-Effectiveness Analysis (Oxford University Press, New York, 1994).
Higgins, J.P. & Thompson, S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558 (2002).
Houlston, R.S. & Ford, D. Genetics of coeliac disease. QJM 89, 737–743 (1996).
Stranger, B.E. et al. Genome-wide associations of gene expression variation in humans. PLoS Genet. 1, e78 (2005).
Stranger, B.E. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848–853 (2007).
Cuzick, J.A. Wilcoxon-type test for trend. Stat. Med. 4, 87–90 (1985).
Acknowledgements
We thank L. Padyukov (Karolinska Institutet) and the EIRA group for providing control samples from the Swedish population for the Swedish replication study. We also acknowledge the work of S. Richards and R. Wade (CTSU, Oxford) in furbishing survival data from CLL-4. We are grateful to all clinicians for their involvement in patient ascertainment. Finally, we are grateful to all the patients and other individuals for their participation. This study makes use of data generated by the Wellcome Trust Case-Control Consortium 1 and 2. A full list of the investigators who contributed to the generation of the data is available from http://www.wtccc.org.uk/. Leukaemia Research Fund provided principal funding for the study (LRF05001 and 06002). Additional funding was provided by CLL Global Research Foundation, Cancer Research UK (C1298/A8362 supported by the Bobby Moore Fund) and the Arbib Fund. D.C.-S. was supported by the Institute of Cancer Research. Funding for the project was provided by the Wellcome Trust under awards 076113 and 085475. In the UK, National Health Service funding for the Royal Marsden Biomedical Research Centre is acknowledged. E.C. and A.E. acknowledge Fundación Genoma España and CICYT funding (SAF 08/3630) and the Red Ternática de Investigación Cooperativa en Cancer from the Insituto de Salud Carlos III. A.C. and M.T. acknowledge Xunta de Galicia funding for Spanish control population (PGIDIT 08CSA005208PR) and the support of the Spanish National Genotyping Center (USC Node).
Author information
Authors and Affiliations
Contributions
R.S.H. designed the study, obtained funding, directed the study and oversaw analyses. R.S.H. drafted the manuscript, with help from D.C.-S., P.B., M.C.D.B. and S.E.D. M.C.D.B. performed statistical analyses. D.C.-S., M.C.D.B. and S.E.D. performed bioinformatics analyses. R.S.H. and D.C. established the parent study. R.S.H., D.C. and D.C.-S. developed patient recruitment and sample acquisition and performed sample collection of cases. For the GWA and UK replication series 1, D.C.-S. and P.B. supervised laboratory management and oversaw genotyping of cases. D.C.-S. conducted sequencing. J.M.A. and D.J.A. conceived of the Newcastle-based CLL study. J.M.A. established the study, supervised laboratory management and oversaw genotyping of cases and controls. N.J.S. performed sample management of cases and controls. A.G.H. developed the Newcastle Haematology Biobank, incorporating the Newcastle-based CLL study. T.M.-F., G.H.J., G.S., A.P., M.J.S.D., E. Matutes, C.D., R.J.H., A.R.P., D.O., D.J.A., J.R.B., G.P., C.P. and C.F. developed patient recruitment and sample acquisition and performed sample collection of cases. Coordination of the Spain replication series was conducted by E.C. and A.E. E.C. and A.E. provided CLL samples and compiled detailed phenotypic information from cases; A.C. provided controls and compiled detailed phenotypic information from controls. C.R.S. and E. Montserrat performed sample acquisition. Genotyping was performed by M.T. and A.C., and M.T. supervised laboratory management and quality control. For the Swedish case-control study, M.M. performed sample collection and prepared DNA. R.R. performed sample collection for all cases, while J.J., G.J. and K.E.S. performed sample collection of cases in the SCALE study. P.B. and D.C.-S. performed genotyping of cases and controls. All authors contributed to the final paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Tables 1–5 (PDF 2597 kb)
Rights and permissions
About this article
Cite this article
Crowther-Swanepoel, D., Broderick, P., Di Bernardo, M. et al. Common variants at 2q37.3, 8q24.21, 15q21.3 and 16q24.1 influence chronic lymphocytic leukemia risk. Nat Genet 42, 132–136 (2010). https://doi.org/10.1038/ng.510
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.510
This article is cited by
-
Distinct germline genetic susceptibility profiles identified for common non-Hodgkin lymphoma subtypes
Leukemia (2022)
-
Differential Levels of mRNAs in Normal B Lymphocytes, Monoclonal B Lymphocytosis and Chronic Lymphocytic Leukemia Cells from the Same Family Identify Susceptibility Genes
Oncology and Therapy (2021)
-
Genome-wide association study identifies risk loci for progressive chronic lymphocytic leukemia
Nature Communications (2021)
-
Identification of leukemia stem cell expression signatures through Monte Carlo feature selection strategy and support vector machine
Cancer Gene Therapy (2020)
-
Genetic predisposition for multiple myeloma
Leukemia (2020)