Abstract
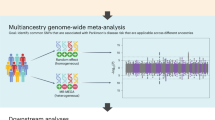
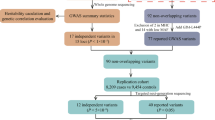
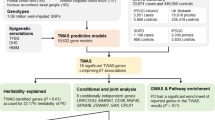
To identify susceptibility variants for Parkinson's disease (PD), we performed a genome-wide association study (GWAS) and two replication studies in a total of 2,011 cases and 18,381 controls from Japan. We identified a new susceptibility locus on 1q32 (P = 1.52 × 10−12) and designated this as PARK16, and we also identified BST1 on 4p15 as a second new risk locus (P = 3.94 × 10−9). We also detected strong associations at SNCA on 4q22 (P = 7.35 × 10−17) and LRRK2 on 12q12 (P = 2.72 × 10−8), both of which are implicated in autosomal dominant forms of parkinsonism. By comparing results of a GWAS performed on individuals of European ancestry, we identified PARK16, SNCA and LRRK2 as shared risk loci for PD and BST1 and MAPT as loci showing population differences. Our results identify two new PD susceptibility loci, show involvement of autosomal dominant parkinsonism loci in typical PD and suggest that population differences contribute to genetic heterogeneity in PD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
de Rijk, M.C. et al. Prevalence of parkinsonism and Parkinson's disease in Europe: the EUROPARKINSON collaborative study. J. Neurol. Neurosurg. Psychiatry 62, 10–15 (1997).
Polymeropoulos, M.H. et al. Mutation in the α-synuclein gene identified in families with Parkinson's disease. Science 276, 2045–2047 (1997).
Paisán-Ruíz, C. et al. Cloning of the gene containing mutations that cause PARK8-linked Parkinson's disease. Neuron 44, 595–600 (2004).
Zimprich, A. et al. Mutations in LRRK2 cause autosomal-dominant parkinsonism with pleomorphic pathology. Neuron 44, 601–607 (2004).
Farrer, M.J. Genetics of Parkinson disease: paradigm shifts and future prospects. Nat. Rev. Genet. 7, 306–318 (2006).
Thomas, B. & Beal, M.F. Parkinson's disease. Hum. Mol. Genet. 16 Spec No. 2, R183–R194 (2007).
Warner, T.T. & Schapira, A.H. Genetic and environmental factors in the cause of Parkinson's disease. Ann. Neurol. 53 (Suppl 3), S16–S23 (2003).
Pals, P. et al. α-Synuclein promoter confers susceptibility to Parkinson's disease. Ann. Neurol. 56, 591–595 (2004).
Mizuta, I. et al. Multiple candidate gene analysis identifies α-synuclein as a susceptibility gene for sporadic Parkinson's disease. Hum. Mol. Genet. 15, 1151–1158 (2006).
Müller, J.C. et al. Multiple regions of α-synuclein are associated with Parkinson's disease. Ann. Neurol. 57, 535–541 (2005).
Aharon-Peretz, J., Rosenbaum, H. & Gershoni-Baruch, R. Mutations in the glucocerebrosidase gene and Parkinson's disease in Ashkenazi Jews. N. Engl. J. Med. 351, 1972–1977 (2004).
Maraganore, D.M. et al. High-resolution whole-genome association study of Parkinson disease. Am. J. Hum. Genet. 77, 685–693 (2005).
Fung, H.C. et al. Genome-wide genotyping in Parkinson's disease and neurologically normal controls: first stage analysis and public release of data. Lancet Neurol. 5, 911–916 (2006).
Pankratz, N. et al. Genomewide association study for susceptibility genes contributing to familial Parkinson disease. Hum. Genet. 124, 593–605 (2009).
Welcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Gabriel, S.B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002).
Simón-Sánchez, J. et al. Genome-wide association study reveals genetic risk underlying Parkinson's disease. Nat. Genet. advance online publication, doi:10.1038/ng.487 (15 November 2009).
Kolisek, M. et al. SLC41A1 is a novel mammalian Mg2+ carrier. J. Biol. Chem. 283, 16235–16247 (2008).
Garruto, R.M. et al. Disappearance of high-incidence amyotrophic lateral sclerosis and parkinsonism-dementia on Guam. Neurology 35, 193–198 (1985).
Shimizu, F. et al. Cloning and chromosome assignment to 1q32 of a human cDNA (RAB7L1) encoding a small GTP-binding protein, a member of the RAS superfamily. Cytogenet. Cell Genet. 77, 261–263 (1997).
Ostvold, A.C. et al. Molecular cloning of a mammalian nuclear phosphoprotein NUCKS, which serves as a substrate for Cdk1 in vivo. Eur. J. Biochem. 268, 2430–2440 (2001).
Dixon, A.L. et al. A genome-wide association study of global gene expression. Nat. Genet. 39, 1202–1207 (2007).
Yamamoto-Katayama, S. et al. Crystallographic studies on human BST-1/CD157 with ADP-ribosyl cyclase and NAD glycohydrolase activities. J. Mol. Biol. 316, 711–723 (2002).
Lee, H.C. et al. Physiological functions of cyclic ADP-ribose and NAADP as calcium messengers. Annu. Rev. Pharmacol. Toxicol. 41, 317–345 (2001).
Wilson, C.J. & Callaway, J.C. Coupled oscillator model of the dopaminergic neuron of the substantia nigra. J. Neurophysiol. 83, 3084–3100 (2000).
Chan, C.S. et al. 'Rejuvenation' protects neurons in mouse models of Parkinson's disease. Nature 447, 1081–1086 (2007).
Surmeier, D.J. Calcium, ageing, and neuronal vulnerability in Parkinson's disease. Lancet Neurol. 6, 933–938 (2007).
Singleton, A.B. et al. α-Synuclein locus triplication causes Parkinson's disease. Science 302, 841 (2003).
Uldry, M. et al. Identification of a mammalian H(+)-myo-inositol symporter expressed predominantly in the brain. EMBO J. 20, 4467–4477 (2001).
Skipper, L. et al. Comprehensive evaluation of common genetic variation within LRRK2 reveals evidence for association with sporadic Parkinson's disease. Hum. Mol. Genet. 14, 3549–3556 (2005).
Biskup, S. et al. Common variants of LRRK2 are not associated with sporadic Parkinson's disease. Ann. Neurol. 58, 905–908 (2005).
Smith, W.W. et al. Kinase activity of mutant LRRK2 mediates neuronal toxicity. Nat. Neurosci. 9, 1231–1233 (2006).
West, A.B. et al. Parkinson's disease-associated mutations in leucine-rich repeat kinase 2 augment kinase activity. Proc. Natl. Acad. Sci. USA 102, 16842–16847 (2005).
Gilks, W.P. et al. A common LRRK2 mutation in idiopathic Parkinson's disease. Lancet 365, 415–416 (2005).
Hutton, M. et al. Association of missense and 5′-splice-site mutations in tau with the inherited dementia FTDP-17. Nature 393, 702–705 (1998).
Pittman, A.M. et al. Linkage disequilibrium fine mapping and haplotype association analysis of the tau gene in progressive supranuclear palsy and corticobasal degeneration. J. Med. Genet. 42, 837–846 (2005).
Webb, A. et al. Role of the tau gene region chromosome inversion in progressive supranuclear palsy, corticobasal degeneration, and related disorders. Arch. Neurol. 65, 1473–1478 (2008).
Healy, D.G. et al. Tau gene and Parkinson's disease: a case-control study and meta-analysis. J. Neurol. Neurosurg. Psychiatry 75, 962–965 (2004).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Skol, A.D., Scott, L.J., Abecasis, G.R. & Boehnke, M. Joint analysis is more efficient than replication-based analysis for two-stage genome-wide association studies. Nat. Genet. 38, 209–213 (2006).
Barrett, J.C., Fry, B., Maller, J. & Daly, M. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Acknowledgements
We are grateful to the individuals with PD who participated in this study. We also thank T. Takeshima and E. Ohta for PD samples, H. Inoko and K. Tokunaga for control samples, M. Kanagawa and K. Ura for editing, K. Yasuno and R. Ashida for analyses, and K. Yamada for genotyping. We appreciate all the volunteers and participating institutions of BioBank Japan for samples. This work was supported by a grant from Core Research for Evolutional Science and Technology (CREST), Japan Science and Technology Agency (JST); by the Global COE program and KAKENHI (17019044 and 19590990), both from the Ministry of Education, Culture, Sports, Science and Technology of Japan; and by the Grant-in-Aid for 'the Research Committee for the Neurodegenerative Diseases' of the Research on Measures for Intractable Diseases and Research Grant (H19-Genome-Ippan-001), all from the Ministry of Health, Labor and Welfare of Japan.
Author information
Authors and Affiliations
Contributions
T. Toda conceived the study. W.S., I.M. and T. Toda designed the study. W.S., Y.N., C.I., M.K. and T.Y. performed genotyping. W.S. and T. Toda wrote the manuscript. W.S., T.K. and T. Tsunoda performed data analysis. W.S., I.M., Y.H., M.W., A.T., H.T., K.N., K.H., F.O., H.K., S.S., M.Y., N.H., M.M. and T. Toda managed Parkinson clinical information and DNA samples. M.K. and Y.N. managed DNA samples belonging to BioBank Japan. T. Toda obtained funding for the study.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–3 and Supplementary Figures 1–4. (PDF 576 kb)
Rights and permissions
About this article
Cite this article
Satake, W., Nakabayashi, Y., Mizuta, I. et al. Genome-wide association study identifies common variants at four loci as genetic risk factors for Parkinson's disease. Nat Genet 41, 1303–1307 (2009). https://doi.org/10.1038/ng.485
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.485
This article is cited by
-
Cell-to-cell transmitted alpha-synuclein recapitulates experimental Parkinson’s disease
npj Parkinson's Disease (2024)
-
Transcriptomic analysis reveals associations of blood-based A-to-I editing with Parkinson’s disease
Journal of Neurology (2024)
-
Clinical biomarkers for Lewy body diseases
Cell & Bioscience (2023)
-
Research progress on the role of extracellular vesicles in neurodegenerative diseases
Translational Neurodegeneration (2023)
-
Quantitative proteomics and phosphoproteomics of urinary extracellular vesicles define putative diagnostic biosignatures for Parkinson’s disease
Communications Medicine (2023)